Unlocking the potential of phytochemicals in inhibiting SARS-CoV-2 M Pro protein - An in-silico and cell-based approach
et al., Research Square, doi:10.21203/rs.3.rs-3888947/v1, Jan 2024
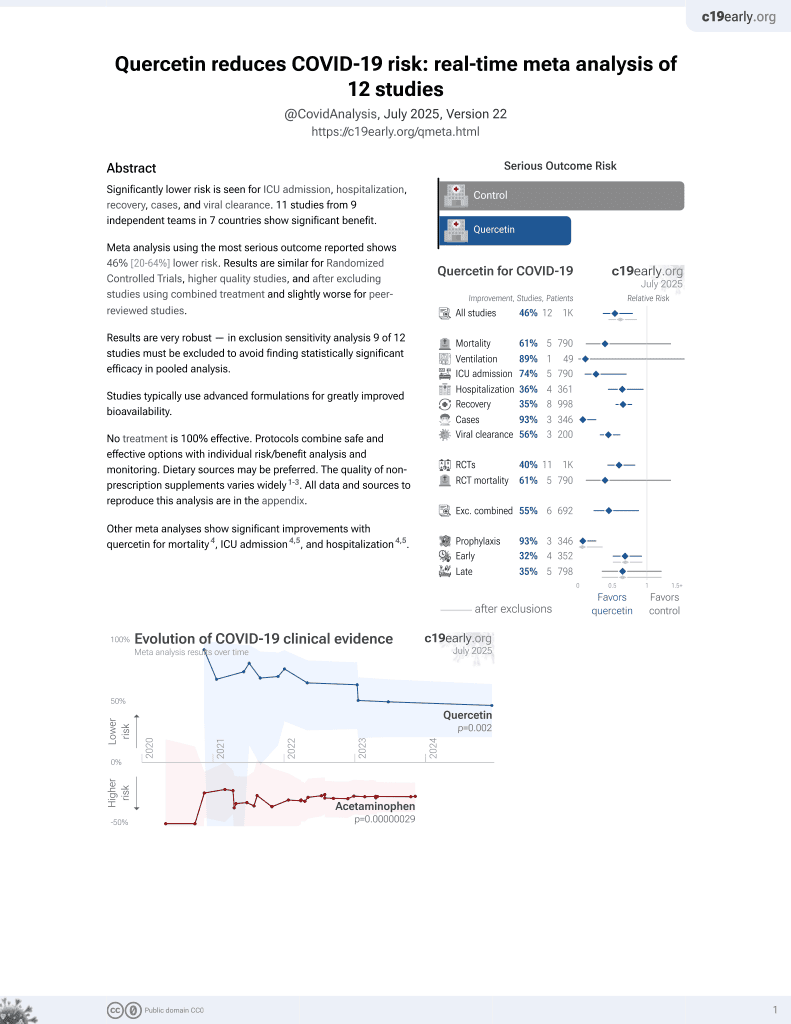
Quercetin for COVID-19
36th treatment shown to reduce risk in
January 2022, now with p = 0.0018 from 9 studies.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
In silico and in vitro study including quercetin and curcumin derivatives as potential SARS-CoV-2 main protease (Mpro) inhibitors. Molecular dynamics simulations and virtual screening identified quercetin and curcumin derivatives demethoxycurcumin and hexahydrocurcumin as potential binders of Mpro. Demethoxycurcumin was tested in vitro, showing significant inhibitory activity against SARS-CoV-2, with no cytotoxicity observed.
92 preclinical studies support the efficacy of quercetin for COVID-19:
In silico studies predict inhibition of SARS-CoV-2, or minimization of side effects, with quercetin or metabolites via binding to the spikeA,12,13,19,20,33,35,36,38,41,49,50,52,53,76 (and specifically the receptor binding domainB,9), MproC,8,9,12,13,17,19,21,23,25,27,29,31,34,35,38,41,45,47-49,53-56,73 , RNA-dependent RNA polymeraseD,9,11-13,19,43 , PLproE,13,48,56 , ACE2F,28,33,34,38,39,48,52 , TMPRSS2G,33, nucleocapsidH,13, helicaseI,13,40,45 , endoribonucleaseJ,50, NSP16/10K,16, cathepsin LL,37, Wnt-3M,33, FZDN,33, LRP6O,33, ezrinP,51, ADRPQ,49, NRP1R,52, EP300S,26, PTGS2T,34, HSP90AA1U,26,34 , matrix metalloproteinase 9V,42, IL-6W,32,46 , IL-10X,32, VEGFAY,46, and RELAZ,46 proteins, and inhibition of spike-ACE2 interactionAA,10.
In vitro studies demonstrate inhibition of the MproC,25,59,64,72 protein, and inhibition of spike-ACE2 interactionAA,60.
In vitro studies demonstrate efficacy in Calu-3AB,63, A549AC,32, HEK293-ACE2+AD,71, Huh-7AE,36, Caco-2AF,62, Vero E6AG,30,53,62 , mTECAH,65, RAW264.7AI,65, and HLMECAJ,10 cells.
Animal studies demonstrate efficacy in K18-hACE2 miceAK,68, db/db miceAL,65,75 , BALB/c miceAM,74, and rats30.
Quercetin reduced proinflammatory cytokines and protected lung and kidney tissue against LPS-induced damage in mice74, inhibits LPS-induced cytokine storm by modulating key inflammatory and antioxidant pathways in macrophages15, may block ACE2-spike interaction and NLRP3 inflammasome, limiting viral entry and inflammation6, upregulates the SIRT1/AMPK axis to inhibit oxidative injury and accelerate viral clearance77, inhibits SARS-CoV-2 ORF3a ion channel activity, which contributes to viral pathogenicity and cytotoxicity67, may alleviate COVID-19 ARDS via inhibition of EGFR and JAK2 inflammatory targets2, may destabilize the Spike protein, IL-6R, and integrins via conserved residues, blocking viral entry, hyperinflammation, and platelet aggregation78, and may reduce COVID-19 neuroinflammation and cognitive dysfunction through anti-inflammatory mechanisms and neuroprotective effects1.
Study covers curcumin and quercetin.
1.
Xie et al., Identification of key genes modules linking brain aging signatures and COVID-19-associated cognitive impairment, Virus Research, doi:10.1016/j.virusres.2026.199707.
2.
Gupta et al., Harnessing phytoconstituents to treat COVID-19 triggered acute respiratory distress syndrome: Insights from network pharmacology, and molecular modeling, Phytochemistry Letters, doi:10.1016/j.phytol.2025.104105.
3.
Sun et al., Feasibility of the inhibitor development for SARS-CoV-2: a systematic approach for drug design, Journal of Molecular Modeling, doi:10.1007/s00894-025-06541-2.
4.
Torabfam et al., Improving quercetin solubility via structural modification enhances dual-target coronavirus entry: an integrated in-vitro and in-silico study, Scientific Reports, doi:10.1038/s41598-025-27374-2.
5.
Abdelhameed et al., Phytochemical and antiviral investigation of Cynanchum acutum L. extract and derived semi-synthetic analogs targeting SARS-CoV-2 main protease, Future Journal of Pharmaceutical Sciences, doi:10.1186/s43094-025-00907-2.
6.
Manikyam et al., INP-Guided Network Pharmacology Discloses Multi-Target Therapeutic Strategy Against Cytokine and IgE Storms in the SARS-CoV-2 NB.1.8.1 Variant, Research Square, doi:10.21203/rs.3.rs-6819274/v1.
7.
Makoana et al., Integration of metabolomics and chemometrics with in-silico and in-vitro approaches to unravel SARS-Cov-2 inhibitors from South African plants, PLOS ONE, doi:10.1371/journal.pone.0320415.
8.
Bano et al., Biochemical Screening of Phytochemicals and Identification of Scopoletin as a Potential Inhibitor of SARS-CoV-2 Mpro, Revealing Its Biophysical Impact on Structural Stability, Viruses, doi:10.3390/v17030402.
9.
Rajamanickam et al., Exploring the Potential of Siddha Formulation MilagaiKudineer-Derived Phytotherapeutics Against SARS-CoV-2: An In-Silico Investigation for Antiviral Intervention, Journal of Pharmacy and Pharmacology Research, doi:10.26502/fjppr.0105.
10.
Moharram et al., Secondary metabolites of Alternaria alternate appraisal of their SARS-CoV-2 inhibitory and anti-inflammatory potentials, PLOS ONE, doi:10.1371/journal.pone.0313616.
11.
Metwaly et al., Integrated study of Quercetin as a potent SARS-CoV-2 RdRp inhibitor: Binding interactions, MD simulations, and In vitro assays, PLOS ONE, doi:10.1371/journal.pone.0312866.
12.
Al balawi et al., Assessing multi-target antiviral and antioxidant activities of natural compounds against SARS-CoV-2: an integrated in vitro and in silico study, Bioresources and Bioprocessing, doi:10.1186/s40643-024-00822-z.
13.
Haque et al., Exploring potential therapeutic candidates against COVID-19: a molecular docking study, Discover Molecules, doi:10.1007/s44345-024-00005-5.
14.
Pan et al., Decoding the mechanism of Qingjie formula in the prevention of COVID-19 based on network pharmacology and molecular docking, Heliyon, doi:10.1016/j.heliyon.2024.e39167.
15.
Xu et al., Quercetin inhibited LPS-induced cytokine storm by interacting with the AKT1-FoxO1 and Keap1-Nrf2 signaling pathway in macrophages, Scientific Reports, doi:10.1038/s41598-024-71569-y.
16.
Tamil Selvan et al., Computational Investigations to Identify Potent Natural Flavonoid Inhibitors of the Nonstructural Protein (NSP) 16/10 Complex Against Coronavirus, Cureus, doi:10.7759/cureus.68098.
17.
Sunita et al., Characterization of Phytochemical Inhibitors of the COVID-19 Primary Protease Using Molecular Modelling Approach, Asian Journal of Microbiology and Biotechnology, doi:10.56557/ajmab/2024/v9i28800.
18.
Wu et al., Biomarkers Prediction and Immune Landscape in Covid-19 and “Brain Fog”, Elsevier BV, doi:10.2139/ssrn.4897774.
19.
Raman et al., Phytoconstituents of Citrus limon (Lemon) as Potential Inhibitors Against Multi Targets of SARS‐CoV‐2 by Use of Molecular Modelling and In Vitro Determination Approaches, ChemistryOpen, doi:10.1002/open.202300198.
20.
Asad et al., Exploring the antiviral activity of Adhatoda beddomei bioactive compounds in interaction with coronavirus spike protein, Archives of Medical Reports, 1:1, archmedrep.com/index.php/amr/article/view/3.
21.
Irfan et al., Phytoconstituents of Artemisia Annua as potential inhibitors of SARS CoV2 main protease: an in silico study, BMC Infectious Diseases, doi:10.1186/s12879-024-09387-w.
22.
Yuan et al., Network pharmacology and molecular docking reveal the mechanisms of action of Panax notoginseng against post-COVID-19 thromboembolism, Review of Clinical Pharmacology and Pharmacokinetics - International Edition, doi:10.61873/DTFA3974.
23.
Nalban et al., Targeting COVID-19 (SARS-CoV-2) main protease through phytochemicals of Albizia lebbeck: molecular docking, molecular dynamics simulation, MM–PBSA free energy calculations, and DFT analysis, Journal of Proteins and Proteomics, doi:10.1007/s42485-024-00136-w.
24.
Zhou et al., Bioinformatics and system biology approaches to determine the connection of SARS-CoV-2 infection and intrahepatic cholangiocarcinoma, PLOS ONE, doi:10.1371/journal.pone.0300441.
25.
Waqas et al., Discovery of Novel Natural Inhibitors Against SARS-CoV-2 Main Protease: A Rational Approach to Antiviral Therapeutics, Current Medicinal Chemistry, doi:10.2174/0109298673292839240329081008.
26.
Hasanah et al., Decoding the therapeutic potential of empon-empon: a bioinformatics expedition unraveling mechanisms against COVID-19 and atherosclerosis, International Journal of Applied Pharmaceutics, doi:10.22159/ijap.2024v16i2.50128.
27.
Shaik et al., Computational identification of selected bioactive compounds from Cedrus deodara as inhibitors against SARS-CoV-2 main protease: a pharmacoinformatics study, Indian Drugs, doi:10.53879/id.61.02.13859.
28.
Wang et al., Investigating the Mechanism of Qu Du Qiang Fei 1 Hao Fang Formula against Coronavirus Disease 2019 Based on Network Pharmacology Method, World Journal of Traditional Chinese Medicine, doi:10.4103/2311-8571.395061.
29.
Singh et al., Unlocking the potential of phytochemicals in inhibiting SARS-CoV-2 M Pro protein - An in-silico and cell-based approach, Research Square, doi:10.21203/rs.3.rs-3888947/v1.
30.
El-Megharbel et al., Chemical and spectroscopic characterization of (Artemisinin/Quercetin/ Zinc) novel mixed ligand complex with assessment of its potent high antiviral activity against SARS-CoV-2 and antioxidant capacity against toxicity induced by acrylamide in male rats, PeerJ, doi:10.7717/peerj.15638.
31.
Akinwumi et al., Evaluation of therapeutic potentials of some bioactive compounds in selected African plants targeting main protease (Mpro) in SARS-CoV-2: a molecular docking study, Egyptian Journal of Medical Human Genetics, doi:10.1186/s43042-023-00456-4.
32.
Yang et al., Active ingredient and mechanistic analysis of traditional Chinese medicine formulas for the prevention and treatment of COVID-19: Insights from bioinformatics and in vitro experiments, Medicine, doi:10.1097/MD.0000000000036238.
33.
Chandran et al., Molecular docking analysis of quercetin with known CoVid-19 targets, Bioinformation, doi:10.6026/973206300191081.
34.
Qin et al., Exploring the bioactive compounds of Feiduqing formula for the prevention and management of COVID-19 through network pharmacology and molecular docking, Medical Data Mining, doi:10.53388/MDM202407003.
35.
Moschovou et al., Exploring the Binding Effects of Natural Products and Antihypertensive Drugs on SARS-CoV-2: An In Silico Investigation of Main Protease and Spike Protein, International Journal of Molecular Sciences, doi:10.3390/ijms242115894.
36.
Pan (B) et al., Quercetin: A promising drug candidate against the potential SARS-CoV-2-Spike mutants with high viral infectivity, Computational and Structural Biotechnology Journal, doi:10.1016/j.csbj.2023.10.029.
37.
Ahmed et al., Evaluation of the Effect of Zinc, Quercetin, Bromelain and Vitamin C on COVID-19 Patients, International Journal of Diabetes Management, doi:10.61797/ijdm.v2i2.259.
38.
Thapa et al., In-silico Approach for Predicting the Inhibitory Effect of Home Remedies on Severe Acute Respiratory Syndrome Coronavirus-2, Makara Journal of Science, doi:10.7454/mss.v27i3.1609.
39.
Alkafaas et al., A study on the effect of natural products against the transmission of B.1.1.529 Omicron, Virology Journal, doi:10.1186/s12985-023-02160-6.
40.
Singh (B) et al., Flavonoids as Potent Inhibitor of SARS-CoV-2 Nsp13 Helicase: Grid Based Docking Approach, Middle East Research Journal of Pharmaceutical Sciences, doi:10.36348/merjps.2023.v03i04.001.
41.
Mandal et al., In silico anti-viral assessment of phytoconstituents in a traditional (Siddha Medicine) polyherbal formulation – Targeting Mpro and pan-coronavirus post-fusion Spike protein, Journal of Traditional and Complementary Medicine, doi:10.1016/j.jtcme.2023.07.004.
42.
Sai Ramesh et al., Computational analysis of the phytocompounds of Mimusops elengi against spike protein of SARS CoV2 – An Insilico model, International Journal of Biological Macromolecules, doi:10.1016/j.ijbiomac.2023.125553.
43.
Corbo et al., Inhibitory potential of phytochemicals on five SARS-CoV-2 proteins: in silico evaluation of endemic plants of Bosnia and Herzegovina, Biotechnology & Biotechnological Equipment, doi:10.1080/13102818.2023.2222196.
44.
Azmi et al., Utilization of quercetin flavonoid compounds in onion (Allium cepa L.) as an inhibitor of SARS-CoV-2 spike protein against ACE2 receptors, 11th International Seminar on New Paradigm and Innovation on Natural Sciences and its Application, doi:10.1063/5.0140285.
45.
Alanzi et al., Structure-based virtual identification of natural inhibitors of SARS-CoV-2 and its Delta and Omicron variant proteins, Future Virology, doi:10.2217/fvl-2022-0184.
46.
Yang (B) et al., In silico evidence implicating novel mechanisms of Prunella vulgaris L. as a potential botanical drug against COVID-19-associated acute kidney injury, Frontiers in Pharmacology, doi:10.3389/fphar.2023.1188086.
47.
Wang (B) et al., Computational Analysis of Lianhua Qingwen as an Adjuvant Treatment in Patients with COVID-19, Society of Toxicology Conference, 2023, www.researchgate.net/publication/370491709_Y_Wang_A_E_Tan_O_Chew_A_Hsueh_and_D_E_Johnson_2023_Computational_Analysis_of_Lianhua_Qingwen_as_an_Adjuvant_Treatment_in_Patients_with_COVID-19_Toxicologist_1921_507.
48.
Ibeh et al., Computational studies of potential antiviral compounds from some selected Nigerian medicinal plants against SARS-CoV-2 proteins, Informatics in Medicine Unlocked, doi:10.1016/j.imu.2023.101230.
49.
Nguyen et al., The Potential of Ameliorating COVID-19 and Sequelae From Andrographis paniculata via Bioinformatics, Bioinformatics and Biology Insights, doi:10.1177/11779322221149622.
50.
Alavi et al., Interaction of Epigallocatechin Gallate and Quercetin with Spike Glycoprotein (S-Glycoprotein) of SARS-CoV-2: In Silico Study, Biomedicines, doi:10.3390/biomedicines10123074.
51.
Chellasamy et al., Docking and molecular dynamics studies of human ezrin protein with a modelled SARS-CoV-2 endodomain and their interaction with potential invasion inhibitors, Journal of King Saud University - Science, doi:10.1016/j.jksus.2022.102277.
52.
Şimşek et al., In silico identification of SARS-CoV-2 cell entry inhibitors from selected natural antivirals, Journal of Molecular Graphics and Modelling, doi:10.1016/j.jmgm.2021.108038.
53.
Kandeil et al., Bioactive Polyphenolic Compounds Showing Strong Antiviral Activities against Severe Acute Respiratory Syndrome Coronavirus 2, Pathogens, doi:10.3390/pathogens10060758.
54.
Rehman et al., Natural Compounds as Inhibitors of SARS-CoV-2 Main Protease (3CLpro): A Molecular Docking and Simulation Approach to Combat COVID-19, Current Pharmaceutical Design, doi:10.2174/1381612826999201116195851.
55.
Sekiou et al., In-Silico Identification of Potent Inhibitors of COVID-19 Main Protease (Mpro) and Angiotensin Converting Enzyme 2 (ACE2) from Natural Products: Quercetin, Hispidulin, and Cirsimaritin Exhibited Better Potential Inhibition than Hydroxy-Chloroquine Against COVID-19 Main Protease Active Site and ACE2, ChemRxiv, doi:10.26434/chemrxiv.12181404.v1.
56.
Zhang et al., In silico screening of Chinese herbal medicines with the potential to directly inhibit 2019 novel coronavirus, Journal of Integrative Medicine, doi:10.1016/j.joim.2020.02.005.
57.
Sisti et al., Evaluation of respiratory virus transmissibility and resilience from fomites: the case of 11 SARS-CoV-2 clinical isolates, Applied and Environmental Microbiology, doi:10.1128/aem.00774-25.
58.
Spinelli et al., Amphibian‐Derived Peptides as Natural Inhibitors of SARS‐CoV‐2 Main Protease (Mpro): A Combined In Vitro and In Silico Approach, Chemistry & Biodiversity, doi:10.1002/cbdv.202403202.
59.
Aguilera-Rodriguez et al., Inhibition of SARS-CoV-2 3CLpro by chemically modified tyrosinase from Agaricus bisporus, RSC Medicinal Chemistry, doi:10.1039/D4MD00289J.
60.
Emam et al., Establishment of in-house assay for screening of anti-SARS-CoV-2 protein inhibitors, AMB Express, doi:10.1186/s13568-024-01739-8.
61.
Fang et al., Development of nanoparticles incorporated with quercetin and ACE2-membrane as a novel therapy for COVID-19, Journal of Nanobiotechnology, doi:10.1186/s12951-024-02435-2.
62.
Roy et al., Quercetin inhibits SARS-CoV-2 infection and prevents syncytium formation by cells co-expressing the viral spike protein and human ACE2, Virology Journal, doi:10.1186/s12985-024-02299-w.
63.
DiGuilio et al., Quercetin improves and protects Calu-3 airway epithelial barrier function, Frontiers in Cell and Developmental Biology, doi:10.3389/fcell.2023.1271201.
64.
Zhang (B) et al., Discovery of the covalent SARS‐CoV‐2 Mpro inhibitors from antiviral herbs via integrating target‐based high‐throughput screening and chemoproteomic approaches, Journal of Medical Virology, doi:10.1002/jmv.29208.
65.
Wu (B) et al., SARS-CoV-2 N protein induced acute kidney injury in diabetic db/db mice is associated with a Mincle-dependent M1 macrophage activation, Frontiers in Immunology, doi:10.3389/fimmu.2023.1264447.
66.
Xu (B) et al., Bioactive compounds from Huashi Baidu decoction possess both antiviral and anti-inflammatory effects against COVID-19, Proceedings of the National Academy of Sciences, doi:10.1073/pnas.2301775120.
67.
Fam et al., Channel activity of SARS-CoV-2 viroporin ORF3a inhibited by adamantanes and phenolic plant metabolites, Scientific Reports, doi:10.1038/s41598-023-31764-9.
68.
Aguado et al., Senolytic therapy alleviates physiological human brain aging and COVID-19 neuropathology, bioRxiv, doi:10.1101/2023.01.17.524329.
69.
Goc et al., Inhibitory effects of specific combination of natural compounds against SARS-CoV-2 and its Alpha, Beta, Gamma, Delta, Kappa, and Mu variants, European Journal of Microbiology and Immunology, doi:10.1556/1886.2021.00022.
70.
Munafò et al., Quercetin and Luteolin Are Single-digit Micromolar Inhibitors of the SARS-CoV-2 RNA-dependent RNA Polymerase, Research Square, doi:10.21203/rs.3.rs-1149846/v1.
71.
Singh (C) et al., The spike protein of SARS-CoV-2 virus induces heme oxygenase-1: Pathophysiologic implications, Biochimica et Biophysica Acta (BBA) - Molecular Basis of Disease, doi:10.1016/j.bbadis.2021.166322.
72.
Bahun et al., Inhibition of the SARS-CoV-2 3CLpro main protease by plant polyphenols, Food Chemistry, doi:10.1016/j.foodchem.2021.131594.
73.
Abian et al., Structural stability of SARS-CoV-2 3CLpro and identification of quercetin as an inhibitor by experimental screening, International Journal of Biological Macromolecules, doi:10.1016/j.ijbiomac.2020.07.235.
74.
Shaker et al., Anti-cytokine Storm Activity of Fraxin, Quercetin, and their Combination on Lipopolysaccharide-Induced Cytokine Storm in Mice: Implications in COVID-19, Iranian Journal of Medical Sciences, doi:10.30476/ijms.2023.98947.3102.
75.
Wu (C) et al., Treatment with Quercetin inhibits SARS-CoV-2 N protein-induced acute kidney injury by blocking Smad3-dependent G1 cell cycle arrest, Molecular Therapy, doi:10.1016/j.ymthe.2022.12.002.
76.
Azmi (B) et al., The role of vitamin D receptor and IL‐6 in COVID‐19, Molecular Genetics & Genomic Medicine, doi:10.1002/mgg3.2172.
a.
The trimeric spike (S) protein is a glycoprotein that mediates viral entry by binding to the host ACE2 receptor, is critical for SARS-CoV-2's ability to infect host cells, and is a target of neutralizing antibodies. Inhibition of the spike protein prevents viral attachment, halting infection at the earliest stage.
b.
The receptor binding domain is a specific region of the spike protein that binds ACE2 and is a major target of neutralizing antibodies. Focusing on the precise binding site allows highly specific disruption of viral attachment with reduced potential for off-target effects.
c.
The main protease or Mpro, also known as 3CLpro or nsp5, is a cysteine protease that cleaves viral polyproteins into functional units needed for replication. Inhibiting Mpro disrupts the SARS-CoV-2 lifecycle within the host cell, preventing the creation of new copies.
d.
RNA-dependent RNA polymerase (RdRp), also called nsp12, is the core enzyme of the viral replicase-transcriptase complex that copies the positive-sense viral RNA genome into negative-sense templates for progeny RNA synthesis. Inhibiting RdRp blocks viral genome replication and transcription.
e.
The papain-like protease (PLpro) has multiple functions including cleaving viral polyproteins and suppressing the host immune response by deubiquitination and deISGylation of host proteins. Inhibiting PLpro may block viral replication and help restore normal immune responses.
f.
The angiotensin converting enzyme 2 (ACE2) protein is a host cell transmembrane protein that serves as the cellular receptor for the SARS-CoV-2 spike protein. ACE2 is expressed on many cell types, including epithelial cells in the lungs, and allows the virus to enter and infect host cells. Inhibition may affect ACE2's physiological function in blood pressure control.
g.
Transmembrane protease serine 2 (TMPRSS2) is a host cell protease that primes the spike protein, facilitating cellular entry. TMPRSS2 activity helps enable cleavage of the spike protein required for membrane fusion and virus entry. Inhibition may especially protect respiratory epithelial cells, buy may have physiological effects.
h.
The nucleocapsid (N) protein binds and encapsulates the viral genome by coating the viral RNA. N enables formation and release of infectious virions and plays additional roles in viral replication and pathogenesis. N is also an immunodominant antigen used in diagnostic assays.
i.
The helicase, or nsp13, protein unwinds the double-stranded viral RNA, a crucial step in replication and transcription. Inhibition may prevent viral genome replication and the creation of new virus components.
j.
The endoribonuclease, also known as NendoU or nsp15, cleaves specific sequences in viral RNA which may help the virus evade detection by the host immune system. Inhibition may hinder the virus's ability to mask itself from the immune system, facilitating a stronger immune response.
k.
The NSP16/10 complex consists of non-structural proteins 16 and 10, forming a 2'-O-methyltransferase that modifies the viral RNA cap structure. This modification helps the virus evade host immune detection by mimicking host mRNA, making NSP16/10 a promising antiviral target.
l.
Cathepsin L is a host lysosomal cysteine protease that can prime the spike protein through an alternative pathway when TMPRSS2 is unavailable. Dual targeting of cathepsin L and TMPRSS2 may maximize disruption of alternative pathways for virus entry.
m.
Wingless-related integration site (Wnt) ligand 3 is a host signaling molecule that activates the Wnt signaling pathway, which is important in development, cell growth, and tissue repair. Some studies suggest that SARS-CoV-2 infection may interfere with the Wnt signaling pathway, and that Wnt3a is involved in SARS-CoV-2 entry.
n.
The frizzled (FZD) receptor is a host transmembrane receptor that binds Wnt ligands, initiating the Wnt signaling cascade. FZD serves as a co-receptor, along with ACE2, in some proposed mechanisms of SARS-CoV-2 infection. The virus may take advantage of this pathway as an alternative entry route.
o.
Low-density lipoprotein receptor-related protein 6 is a cell surface co-receptor essential for Wnt signaling. LRP6 acts in tandem with FZD for signal transduction and has been discussed as a potential co-receptor for SARS-CoV-2 entry.
p.
The ezrin protein links the cell membrane to the cytoskeleton (the cell's internal support structure) and plays a role in cell shape, movement, adhesion, and signaling. Drugs that occupy the same spot on ezrin where the viral spike protein would bind may hindering viral attachment, and drug binding could further stabilize ezrin, strengthening its potential natural capacity to impede viral fusion and entry.
q.
The Adipocyte Differentiation-Related Protein (ADRP, also known as Perilipin 2 or PLIN2) is a lipid droplet protein regulating the storage and breakdown of fats in cells. SARS-CoV-2 may hijack the lipid handling machinery of host cells and ADRP may play a role in this process. Disrupting ADRP's interaction with the virus may hinder the virus's ability to use lipids for replication and assembly.
r.
Neuropilin-1 (NRP1) is a cell surface receptor with roles in blood vessel development, nerve cell guidance, and immune responses. NRP1 may function as a co-receptor for SARS-CoV-2, facilitating viral entry into cells. Blocking NRP1 may disrupt an alternative route of viral entry.
s.
EP300 (E1A Binding Protein P300) is a transcriptional coactivator involved in several cellular processes, including growth, differentiation, and apoptosis, through its acetyltransferase activity that modifies histones and non-histone proteins. EP300 facilitates viral entry into cells and upregulates inflammatory cytokine production.
t.
Prostaglandin G/H synthase 2 (PTGS2, also known as COX-2) is an enzyme crucial for the production of inflammatory molecules called prostaglandins. PTGS2 plays a role in the inflammatory response that can become severe in COVID-19 and inhibitors (like some NSAIDs) may have benefits in dampening harmful inflammation, but note that prostaglandins have diverse physiological functions.
u.
Heat Shock Protein 90 Alpha Family Class A Member 1 (HSP90AA1) is a chaperone protein that helps other proteins fold correctly and maintains their stability. HSP90AA1 plays roles in cell signaling, survival, and immune responses. HSP90AA1 may interact with numerous viral proteins, but note that it has diverse physiological functions.
v.
Matrix metalloproteinase 9 (MMP9), also called gelatinase B, is a zinc-dependent enzyme that breaks down collagen and other components of the extracellular matrix. MMP9 levels increase in severe COVID-19. Overactive MMP9 can damage lung tissue and worsen inflammation. Inhibition of MMP9 may prevent excessive tissue damage and help regulate the inflammatory response.
w.
The interleukin-6 (IL-6) pro-inflammatory cytokine (signaling molecule) has a complex role in the immune response and may trigger and perpetuate inflammation. Elevated IL-6 levels are associated with severe COVID-19 cases and cytokine storm. Anti-IL-6 therapies may be beneficial in reducing excessive inflammation in severe COVID-19 cases.
x.
The interleukin-10 (IL-10) anti-inflammatory cytokine helps regulate and dampen immune responses, preventing excessive inflammation. IL-10 levels can also be elevated in severe COVID-19. IL-10 could either help control harmful inflammation or potentially contribute to immune suppression.
y.
Vascular Endothelial Growth Factor A (VEGFA) promotes the growth of new blood vessels (angiogenesis) and has roles in inflammation and immune responses. VEGFA may contribute to blood vessel leakiness and excessive inflammation associated with severe COVID-19.
z.
RELA is a transcription factor subunit of NF-kB and is a key regulator of inflammation, driving pro-inflammatory gene expression. SARS-CoV-2 may hijack and modulate NF-kB pathways.
aa.
The interaction between the SARS-CoV-2 spike protein and the human ACE2 receptor is a primary method of viral entry, inhibiting this interaction can prevent the virus from attaching to and entering host cells, halting infection at an early stage.
ab.
Calu-3 is a human lung adenocarcinoma cell line with moderate ACE2 and TMPRSS2 expression and SARS-CoV-2 susceptibility. It provides a model of the human respiratory epithelium, but many not be ideal for modeling early stages of infection due to the moderate expression levels of ACE2 and TMPRSS2.
ac.
A549 is a human lung carcinoma cell line with low ACE2 expression and SARS-CoV-2 susceptibility. Viral entry/replication can be studied but the cells may not replicate all aspects of lung infection.
ad.
HEK293-ACE2+ is a human embryonic kidney cell line engineered for high ACE2 expression and SARS-CoV-2 susceptibility.
ae.
Huh-7 cells were derived from a liver tumor (hepatoma).
af.
Caco-2 cells come from a colorectal adenocarcinoma (cancer). They are valued for their ability to form a polarized cell layer with properties similar to the intestinal lining.
ag.
Vero E6 is an African green monkey kidney cell line with low/no ACE2 expression and high SARS-CoV-2 susceptibility. The cell line is easy to maintain and supports robust viral replication, however the monkey origin may not accurately represent human responses.
ah.
mTEC is a mouse tubular epithelial cell line.
ai.
RAW264.7 is a mouse macrophage cell line.
aj.
HLMEC (Human Lung Microvascular Endothelial Cells) are primary endothelial cells derived from the lung microvasculature. They are used to study endothelial function, inflammation, and viral interactions, particularly in the context of lung infections such as SARS-CoV-2. HLMEC express ACE2 and are susceptible to SARS-CoV-2 infection, making them a relevant model for studying viral entry and endothelial responses in the lung.
ak.
A mouse model expressing the human ACE2 receptor under the control of the K18 promoter.
al.
A mouse model of obesity and severe insulin resistance leading to type 2 diabetes due to a mutation in the leptin receptor gene that impairs satiety signaling.
am.
A mouse model commonly used in infectious disease and cancer research due to higher immune response and susceptibility to infection.
Singh et al., 29 Jan 2024, preprint, 7 authors.
Contact: khushboo.singh@amway.com.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
Unlocking the potential of phytochemicals in inhibiting SARS-CoV-2 M Pro protein - An in-silico and cell-based approach
doi:10.21203/rs.3.rs-3888947/v1
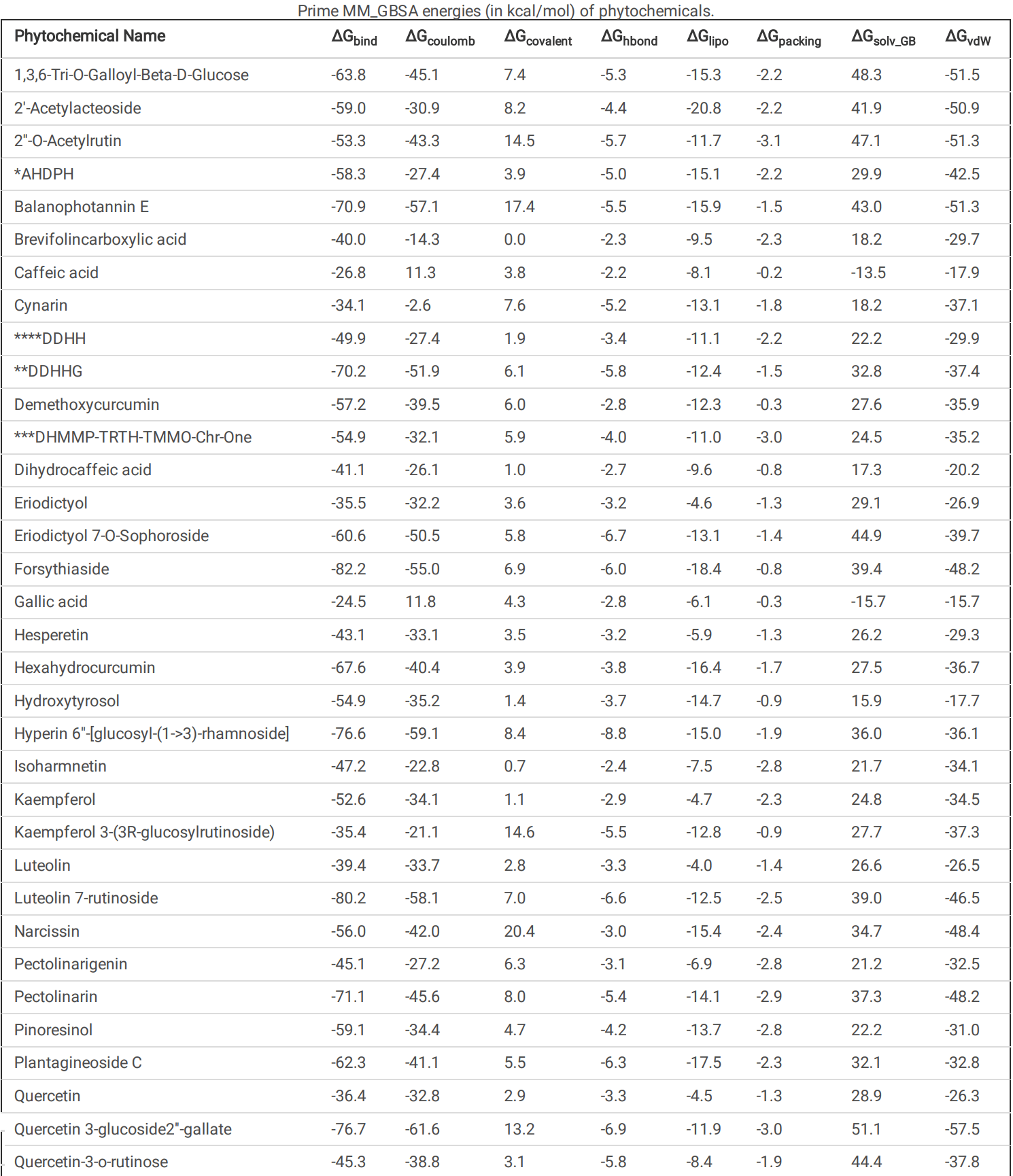
The main protease (M Pro ) of SARS-CoV-2 plays a crucial role in viral replication and is a prime target for therapeutic interventions. Phytochemicals, known for their antiviral properties, have been previously identi ed as potential M Pro inhibitors in several in silico studies. However, the e cacy of these remains in question owing to the inherent exibility of the M Pro binding site, posing challenges in selecting suitable protein structures for virtual screening. In this study, we conducted an extensive analysis of the M Pro binding pocket, utilizing molecular dynamics (MD) simulations to explore its conformational diversity. Based on pocket volume and shape-based clustering, ve representative protein conformations were selected for virtual screening. Virtual screening of a library of ~ 48,000 phytochemicals suggested 39 phytochemicals as potential M Pro inhibitors. Based on subsequent MM-GBSA binding energy calculations and ADMET property predictions, ve compounds were advanced to cell-based viral replication inhibition assays, with three compounds (demethoxycurcumin, shikonin, and withaferin A) exhibiting signi cant (EC50 < 10 uM) inhibition of SARS-CoV-2 replication. Our study provides an understanding of the binding interactions between these phytochemicals and M Pro , contributing signi cantly to the identi cation of promising M Pro inhibitors. Furthermore, beyond its impact on therapeutic development against SARS-CoV-2, this research highlights a crucial role of proper nutrition in the ght against viral infections. Phytochemical Name Docking scores Conformation 1 Conformation 2 Conformation 3 Conformation 4 Conformation 5 1,3,6-Tri-O-Galloyl-Beta-D-Glucose -7.6 -8.8 -11.1 -7.5 -10.1 2'-Acetylacteoside -8.6 -9.5 -12.1 -13.4 -7.6 2''-O-Acetylrutin -10.3 -9.6 -12.2 -10.8 -10.4 *AHDPH -8.1 -9.0 -11.6 -7.5 -9.0 Balanophotannin E -7.5 -11.0 -12.9 -9.9 -8.4 **DDHHG -9.7 -8.3 -10.9 -11.3 -8.6 ***DHMMP-TRTH-TMMO-Chr-One -9.7 -10.5 -10.7 -9.1 -9.9 Eriodictyol 7-O-Sophoroside -12.6 -9.3 -10.0 -11.1 -10.0 Forsythiaside -10.3 -12.6 -14.3 -14.6 -9.2 Hyperin 6''-[glucosyl-(1->3)-rhamnoside] -9.7 -10.9 -15.9 -12.1 -11.9 Kaempferol 3-(3R-glucosylrutinoside) -10.0 -10.6 -12.0 -11.1 -8.5 Luteolin 7-rutinoside -9.8 -9.4 -14.4 -12.0 -9.9 Narcissin -9.7 -10.5 -10.7 -9.1 -9.9 Pectolinarin -8.9 -7.7 -13.9 -8.5 -7.5 Plantagineoside C -9.4 -10.3 -13.3 -10.4 -9.3 Quercetin 3-glucoside2''-gallate -7.8 -9.2 -12.1 -10.6 -7.5 Quercetin-3-o-rutinose -12.2 -11.0 -11.1 -11.5 -11.4 Salvianolic Acid L (SAL) -9.1 -8.2 -13.3 -11.3 -7.6 Shikonin -8.1 -8.4 -8.6 -8.9 -9.5 Shimobashiric Acid C (SAC) -8.2 -8.7 -10.5 -9.6 -10. 2 *AHDPH = (3R,5R)-3-Acetoxy-5-Hydroxy-1,7-Bis(3,4-Dihydroxyphenyl)Heptane. **DDHHG = (3R,5R)-3,5-Dihydroxy-1-(3,4-Dihydroxyphenyl)-7-(4-Hydroxyphenyl)-Heptane 3-O-Beta-D-Glucopyranoside.
For a comprehensive understanding of the model speci cations, validation, and performance, please refer to the AP11.0 user manual and relevant publications 74, 75 .
Cytoxicity Assay Vero cells were seeded using a multiDrop combi liquid dispenser (Thermo) into 384-well plates at a density of 500 cells/well suspended in 50 µL of media. Cells were allowed to recover and fully attach overnight (approximately 16 hours), at which point library compounds were dispensed into wells using an Echo 550 acoustic dispenser (Labcyte). A total of six nal concentrations where tested (50 µM, 25 µM, 12.5 µM, 6.25 µM, 3.125 µM, and 1.5625 µM) and wells were back lled with DMSO such that all wells contained a xed ratio of DMSO. Compounds were incubated with cells for 1 hour prior to addition of virus and then for an additional 24 hours, then xed with 10% formalin, permeabilized 0.1% Triton X-100, washed, and stained for SARS-CoV-2 N protein using a speci c antibody (Sino Biological # MM05) and uorescently labelled secondary antibody. Cells were counter stained with Hoechst 33342 to detect cell nuclei, washed, and imaged with a Cytation 1 (Biotek) automated. Each image was then analyzed with a custom work ow in Cell Pro ler (Broad Inst., Boston, MA) which involved counting of cell nuclei and infected cells. At least 4 replicates were used to construct dose response curves.
Statistics and data normalization The rate index is calculated from cell counts using the following formula: Where X c..
References
Abdusalam, Murugaiyah, Identi cation of Potential Inhibitors of 3CL Protease of SARS-CoV-2 From ZINC Database by Molecular Docking-Based Virtual Screening, Front. Mol. Biosci
Abraham, GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX
Agrawal, Blunden, Phytochemicals Against SARS-COV-2 Infection, Nat. Prod. Commun
Alici, Tahtaci, Demir, Design and various in silico studies of the novel curcumin derivatives as potential candidates against COVID-19 -associated main enzymes, Comput. Biol. Chem
Anand, Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra α-helical domain, EMBO J
Anand, Ziebuhr, Wadhwani, Mesters, Hilgenfeld, Coronavirus main proteinase (3CLpro) structure: basis for the design of anti-SARS drugs, Science
Banks, Integrated modeling program applied chemical theory (IMPACT), J. Comp. Chem
Berendsen, Postma, Van Gunsteren, Dinola, Haak, Molecular dynamics with coupling to an external bath, J. Chem. Phys
Bharadwaj, Macrolactin A as a Novel Inhibitory Agent for SARS-CoV-2 M pro : Bioinformatics Approach, Appl. Biochem. Biotechnol
Biancatelli, Berrill, Catravas, Marik, Quercetin and vitamin C: an experimental, synergistic therapy for the prevention and treatment of SARS-CoV-2 related disease (COVID-19), Front. Immunol
Bzówka, Mitusińska, Raczyńska, Samol, Tuszyński et al., Structural and Evolutionary Analysis Indicate That the SARS-CoV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design, Int. J. Mol. Sci
Cappelli, Manganelli, Lombardo, Gissi, Benfenati, Validation of quantitative structure-activity relationship models to predict water-solubility of organic compounds, Sci. Total Environ
Chakraborty, The Natural Products Withaferin A and Withanone from the Medicinal Herb Withania somnifera Are Covalent Inhibitors of the SARS-CoV-2 Main Protease, J. Nat. Prod
Cherrak, Merzouk, Mokhtari-Soulimane, Potential bioactive glycosylated avonoids as SARS-CoV-2 main protease inhibitors: A molecular docking and simulation studies, PLoS One
Da Fonseca, Screening of Potential Inhibitors Targeting the Main Protease Structure of SARS-CoV-2 via Molecular Docking, and Approach with Molecular Dynamics, RMSD, RMSF, H-Bond, SASA and MMGBSA, Mol. Biotechnol, doi:Preprintat10.1007/s12033-023-00831-x
Dai, Zhang, Jiang, Su, Li, Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease, Science
Darden, York, Pedersen, Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems, J. Chem. Phys
Dearden, In silico prediction of aqueous solubility, Expet Opin. Drug Discov
Dhawan, Anti-viral activity of Indian plants, Proc. Natl. Acad. Sci. India Sect. B Biol. Sci
Douangamath, Fearon, Gehrtz, Krojer, Lukacik, Crystallographic and electrophilic fragment screening of the SARS-CoV-2 main protease, Nat. Commun
Durdagi, Near-physiological-temperature serial crystallography reveals conformations of SARS-CoV-2 main protease active site for improved drug repurposing, Structure
Durrant, POVME 2.0: An Enhanced Tool for Determining Pocket Shape and Volume Characteristics, J. Chem. Theory Comput
Estrada, Topological analysis of SARS CoV-2 main protease, Chaos
Fei, Contribution of traditional Chinese medicine combined with conventional western medicine treatment for the novel coronavirus disease (COVID-19), current evidence with systematic review and meta-analysis, Phytother. Res
Flynn, Comprehensive tness landscape of SARS-CoV-2 Mpro reveals insights into viral resistance mechanisms, Elife
Galan, Phase 2 randomized study on chloroquine, hydroxychloroquine or ivermectin in hospitalized patients with severe manifestations of SARS-COV-2 infection, Pathog. Glob. Health
Ghosh, Structure-activity relationship (SAR) and molecular dynamics study of withaferin-A fragment derivatives as a potential therapeutic lead against the main protease (Mpro) of SARS-CoV-2, J. Mol. Model
Gorbalenya, The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-ncov and naming it SARS-COV-2, Nature Microbiol
Gossen, A Blueprint for High A nity SARS-CoV-2 Mpro Inhibitors from Activity-Based Compound Library Screening Guided by Analysis of Protein Dynamics, ACS Pharmacol. Transl. Sci
Gupta, Identi cation of potential natural inhibitors of SARS-CoV2 main protease by molecular docking and simulation studies, J. Biomol. Struct. Dyn
Gupta, Structure-Based Virtual Screening and Biochemical Validation to Discover Potential Inhibitor of the SARS-CoV-2 Main Protease, ACS Omega
Henrich, Beutler, Matching the power of high throughput screening to the chemical diversity of natural products, Nat. Prod. Rep
Hess, Bekker, Berendsen, Fraaije, Lincs, A linear constraint solver for molecular simulations, J. Comp. Chem
Huang, Mackerell, CHARMM36 all-atom additive protein force eld: validation based on comparison to NMR data, J. Comput. Chem
Humphrey, Dalke, Schulten, VMD -Visual Molecular Dynamics, J. Mol. Graphics
Issa, The Main Protease of SARS-CoV-2 as a Target for Phytochemicals against Coronavirus, Plants
Jamhour, Phytochemicals As a Potential Inhibitor of COVID-19: An In-Silico Perspective, Russ. J. Phys. Chem
Jin, Structure of M pro from SARS-CoV-2 and discovery of its inhibitors, Nature
Jorgensen, Chandrasekhar, Madura, Impey, Klein, Comparison of simple potential functions for simulating liquid water, J. Chem. Phys
Kaur, How do plants defend themselves against pathogens-Biochemical mechanisms and genetic interventions, Physiol. Mol. Biol. Plants
Khaerunnisa, Kurniawan, Awaluddin, Suhartati, Soetjipto, Potential Inhibitor of COVID-19 Main Protease (M pro ) From Several Medicinal Plant Compounds by Molecular Docking Study, doi:10.20944/preprints202003.0226.v1
Khanna, Herbal Immune-boosters: Substantial warriors of pandemiccovid-19 battle, Phytomedicine
Kneller, Kovalevsky, Coates, Structural plasticity of the SARS-COV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography, Nature Commun
Lachance, Charting, navigating, and populating natural product chemical space for drug discovery, J. Med. Chem
Lawson, Maccoss, Heer, Importance of rigidity in designing small molecule drugs to tackle protein-protein interactions (ppis) through stabilization of desired conformers, J. Med. Chem
Li, Abel, Zhu, Cao, Zhao et al., The VSGB 2.0 model: a next-generation energy model for high-resolution protein structure modeling, Proteins
Ling, Traditional Chinese medicine is a resource for drug discovery against 2019 novel coronavirus (SARS-COV-2), J. Integr. Med
Ma, Disul ram, Carmofur, PX-12, Tideglusib, and Shikonin Are Nonspeci c Promiscuous SARS-CoV-2 Main Protease Inhibitors, ACS Pharmacol. Transl. Sci
Mani, Natural product-derived phytochemicals as potential agents against coronaviruses: a review, Virus Res
Mulu, The impact of curcumin-derived polyphenols on the structure and exibility COVID-19 main protease binding pocket: a molecular dynamics simulation study, PeerJ
Pettersen, UCSF Chimera -A visualization system for exploratory research and analysis, J. Comp. Chem
Remali, Aizat, A review on plant bioactive compounds and their modes of action against coronavirus infection, Front. Pharmacol
Ren, The newly emerged SARS-like coronavirus HCoV-EMC also has an "Achilles' heel": current effective inhibitor targeting a 3C-like protease, Protein Cell
Romano, Tatonetti, Informatics and computational methods in natural product drug discovery: A review and Perspectives, Front. Genet
Singh, Briggs, Impact of lymphoma-linked Asn11Tyr point mutation on the interaction between Bcl-2 and a BH3 mimetic: Insights from molecular dynamics simulation, Chem. Biol. Drug Design
Sztain, Amaro, Mccammon, Elucidation of cryptic and allosteric pockets within the SARS-CoV-2 protease, J. Chem. Inf. Model
Teli, Fragment-based design of SARS-CoV-2 Mpro inhibitors, Struct Chem
Vallejos, Ivermectin to prevent hospitalizations in patients with covid-19 (IVERCOR-covid19) a randomized, double-blind, placebocontrolled trial, BMC Infect. Dis
Wagner, POVME 3.0: Software for Mapping Binding Pocket Flexibility, J. Chem. Theory Comput
Wang, Structure of main protease from human coronavirus NL63: insights for wide spectrum anti-coronavirus drug design, Sci. Rep
Wu, A new coronavirus associated with human respiratory disease in China, Nature
Wu, Author Correction: A New Coronavirus Associated with Human Respiratory Disease in China, Nature
Xue, Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design, J. Virol
Yang, Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking, Front. Pharmacol
Yang, The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor, Proc. Natl. Acad. Sci
Zeng, CMAUP: A database of collective molecular activities of useful plants, Nucleic Acids Res
Zhang, Hilgenfeld, Crystal structure of SARS-CoV-2 Mpro in complex with the activity-based probe, biotin-PEG(4)-Abu-Tle-Leu-Glnvinylsulfone, doi:10.2210/pdb6Z2E/pdb
Zhang, Structure-Based Discovery and Structural Basis of a Novel Broad-Spectrum Natural Product against the Main Protease of Coronavirus, J. Virol
Zhou, A pneumonia outbreak associated with a new coronavirus of probable bat origin, Nature
DOI record:
{
"DOI": "10.21203/rs.3.rs-3888947/v1",
"URL": "http://dx.doi.org/10.21203/rs.3.rs-3888947/v1",
"abstract": "<jats:title>Abstract</jats:title>\n <jats:p>The main protease (M<jats:sup>Pro</jats:sup>) of SARS-CoV-2 plays a crucial role in viral replication and is a prime target for therapeutic interventions. Phytochemicals, known for their antiviral properties, have been previously identified as potential M<jats:sup>Pro</jats:sup> inhibitors in several in silico studies. However, the efficacy of these remains in question owing to the inherent flexibility of the M<jats:sup>Pro</jats:sup> binding site, posing challenges in selecting suitable protein structures for virtual screening. In this study, we conducted an extensive analysis of the M<jats:sup>Pro</jats:sup> binding pocket, utilizing molecular dynamics (MD) simulations to explore its conformational diversity. Based on pocket volume and shape-based clustering, five representative protein conformations were selected for virtual screening. Virtual screening of a library of ~ 48,000 phytochemicals suggested 39 phytochemicals as potential M<jats:sup>Pro</jats:sup> inhibitors. Based on subsequent MM-GBSA binding energy calculations and ADMET property predictions, five compounds were advanced to cell-based viral replication inhibition assays, with three compounds (demethoxycurcumin, shikonin, and withaferin A) exhibiting significant (EC50 < 10 uM) inhibition of SARS-CoV-2 replication. Our study provides an understanding of the binding interactions between these phytochemicals and M<jats:sup>Pro</jats:sup>, contributing significantly to the identification of promising M<jats:sup>Pro</jats:sup> inhibitors. Furthermore, beyond its impact on therapeutic development against SARS-CoV-2, this research highlights a crucial role of proper nutrition in the fight against viral infections.</jats:p>",
"accepted": {
"date-parts": [
[
2024,
1,
22
]
]
},
"author": [
{
"affiliation": [
{
"name": "Amway (United States)"
}
],
"family": "Singh",
"given": "Khushboo",
"sequence": "first"
},
{
"affiliation": [
{
"name": "Boston University"
}
],
"family": "Patten",
"given": "J. J.",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "The University of Texas Medical Branch at Galveston"
}
],
"family": "Dimet",
"given": "Andrea",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Boston University"
}
],
"family": "Davey",
"given": "Robert A.",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "The University of Texas Medical Branch at Galveston"
}
],
"family": "Watowich",
"given": "Stanley J.",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Amway (United States)"
}
],
"family": "Chandra",
"given": "Amit",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Amway (United States)"
}
],
"family": "Leverett",
"given": "Jesse",
"sequence": "additional"
}
],
"container-title": [],
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2024,
1,
29
]
],
"date-time": "2024-01-29T10:28:18Z",
"timestamp": 1706524098000
},
"deposited": {
"date-parts": [
[
2024,
1,
29
]
],
"date-time": "2024-01-29T10:28:52Z",
"timestamp": 1706524132000
},
"group-title": "In Review",
"indexed": {
"date-parts": [
[
2024,
1,
30
]
],
"date-time": "2024-01-30T00:22:24Z",
"timestamp": 1706574144551
},
"institution": [
{
"name": "Research Square"
}
],
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2024,
1,
29
]
]
},
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "unspecified",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
1,
29
]
],
"date-time": "2024-01-29T00:00:00Z",
"timestamp": 1706486400000
}
}
],
"link": [
{
"URL": "https://www.researchsquare.com/article/rs-3888947/v1",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.researchsquare.com/article/rs-3888947/v1.html",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "8761",
"original-title": [],
"posted": {
"date-parts": [
[
2024,
1,
29
]
]
},
"prefix": "10.21203",
"published": {
"date-parts": [
[
2024,
1,
29
]
]
},
"publisher": "Research Square Platform LLC",
"reference": [
{
"DOI": "10.1038/s41586-020-2202-3",
"article-title": "Author Correction: A New Coronavirus Associated with Human Respiratory Disease in China",
"author": "Wu F",
"doi-asserted-by": "crossref",
"first-page": "E7",
"journal-title": "Nature.",
"key": "ref1",
"unstructured": "Wu, F., et al. Author Correction: A New Coronavirus Associated with Human Respiratory Disease in China. Nature. 580, E7 (2020).",
"volume": "580",
"year": "2020"
},
{
"DOI": "10.1038/s41564-020-0695-z",
"article-title": "The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-ncov and naming it SARS-COV-2",
"author": "Gorbalenya AE",
"doi-asserted-by": "crossref",
"first-page": "536",
"journal-title": "Nature Microbiol",
"key": "ref2",
"unstructured": "Gorbalenya, A.E. et al. The species severe acute respiratory syndrome-related coronavirus: Classifying 2019-ncov and naming it SARS-COV-2. Nature Microbiol. 5, 536–544 (2020).",
"volume": "5",
"year": "2020"
},
{
"DOI": "10.1080/20477724.2021.1890887",
"article-title": "Phase 2 randomized study on chloroquine, hydroxychloroquine or ivermectin in hospitalized patients with severe manifestations of SARS-COV-2 infection",
"author": "Galan LE",
"doi-asserted-by": "crossref",
"first-page": "235",
"journal-title": "Pathog. Glob. Health.",
"key": "ref3",
"unstructured": "Galan, L.E. et al. Phase 2 randomized study on chloroquine, hydroxychloroquine or ivermectin in hospitalized patients with severe manifestations of SARS-COV-2 infection. Pathog. Glob. Health. 115, 235–242 (2021).",
"volume": "115",
"year": "2021"
},
{
"DOI": "10.1186/s12879-021-06348-5",
"article-title": "Ivermectin to prevent hospitalizations in patients with covid-19 (IVERCOR-covid19) a randomized, double-blind, placebo-controlled trial",
"author": "Vallejos J",
"doi-asserted-by": "crossref",
"first-page": "635",
"journal-title": "BMC Infect. Dis",
"key": "ref4",
"unstructured": "Vallejos, J. et al. Ivermectin to prevent hospitalizations in patients with covid-19 (IVERCOR-covid19) a randomized, double-blind, placebo-controlled trial. BMC Infect. Dis. 21, 635 (2021)",
"volume": "21",
"year": "2021"
},
{
"author": "Dhawan BN",
"key": "ref5",
"unstructured": "Dhawan, B.N. Anti-viral activity of Indian plants. Proc. Natl. Acad. Sci. India Sect. B Biol. Sci. 82, 209–224 (2012).",
"year": "2012"
},
{
"article-title": "Phytochemical, antioxidant and microbial inhibitory effects of spondias mombin leaf and stem bark extracts",
"first-page": "14",
"journal-title": "IOSR J. Pharm. Biol. Sci.",
"key": "ref6",
"unstructured": "H.C.C, M. et al. Phytochemical, antioxidant and microbial inhibitory effects of spondias mombin leaf and stem bark extracts. IOSR J. Pharm. Biol. Sci. 9, pp. 14–17 (2014).",
"volume": "9",
"year": "2014"
},
{
"DOI": "10.1016/j.phymed.2020.153361",
"article-title": "Herbal Immune-boosters: Substantial warriors of pandemiccovid-19 battle",
"author": "Khanna K",
"doi-asserted-by": "crossref",
"first-page": "153361",
"journal-title": "Phytomedicine",
"key": "ref7",
"unstructured": "Khanna, K. et al. Herbal Immune-boosters: Substantial warriors of pandemiccovid-19 battle. Phytomedicine. 85, 153361 (2021).",
"volume": "85",
"year": "2021"
},
{
"DOI": "10.1016/j.joim.2020.02.004",
"article-title": "Traditional Chinese medicine is a resource for drug discovery against 2019 novel coronavirus (SARS-COV-2)",
"author": "Ling C",
"doi-asserted-by": "crossref",
"first-page": "8788",
"journal-title": "J. Integr. Med.",
"key": "ref8",
"unstructured": "Ling, C. Traditional Chinese medicine is a resource for drug discovery against 2019 novel coronavirus (SARS-COV-2). J. Integr. Med. 18, 8788 (2020).",
"volume": "18",
"year": "2020"
},
{
"DOI": "10.3389/fphar.2020.589044",
"article-title": "A review on plant bioactive compounds and their modes of action against coronavirus infection",
"author": "Remali J",
"doi-asserted-by": "crossref",
"first-page": "589044",
"journal-title": "Front. Pharmacol.",
"key": "ref9",
"unstructured": "Remali, J. & Aizat, W.M. A review on plant bioactive compounds and their modes of action against coronavirus infection. Front. Pharmacol. 11, 589044 (2021).",
"volume": "11",
"year": "2021"
},
{
"article-title": "Contribution of traditional Chinese medicine combined with conventional western medicine treatment for the novel coronavirus disease (COVID-19), current evidence with systematic review and meta‐analysis",
"author": "Fei J",
"journal-title": "Phytother. Res.",
"key": "ref10",
"unstructured": "Fei, J. et al. Contribution of traditional Chinese medicine combined with conventional western medicine treatment for the novel coronavirus disease (COVID-19), current evidence with systematic review and meta‐analysis. Phytother. Res. 35, (2021).",
"volume": "35",
"year": "2021"
},
{
"DOI": "10.1021/acs.jmedchem.7b01120",
"article-title": "Importance of rigidity in designing small molecule drugs to tackle protein–protein interactions (ppis) through stabilization of desired conformers",
"author": "Lawson AD",
"doi-asserted-by": "crossref",
"first-page": "4283",
"journal-title": "J. Med. Chem.",
"key": "ref11",
"unstructured": "Lawson, A.D., MacCoss, M. & Heer, J.P. Importance of rigidity in designing small molecule drugs to tackle protein–protein interactions (ppis) through stabilization of desired conformers. J. Med. Chem. 61, 4283–4289 (2017).",
"volume": "61",
"year": "2017"
},
{
"DOI": "10.1039/c3np70052f",
"article-title": "Matching the power of high throughput screening to the chemical diversity of natural products",
"author": "Henrich CJ",
"doi-asserted-by": "crossref",
"first-page": "1284",
"journal-title": "Nat. Prod. Rep.",
"key": "ref12",
"unstructured": "Henrich, C.J. & Beutler, J.A. Matching the power of high throughput screening to the chemical diversity of natural products. Nat. Prod. Rep. 30, 1284 (2013).",
"volume": "30",
"year": "2013"
},
{
"DOI": "10.3389/fgene.2019.00368",
"article-title": "Informatics and computational methods in natural product drug discovery: A review and Perspectives",
"author": "Romano JD",
"doi-asserted-by": "crossref",
"first-page": "368",
"journal-title": "Front. Genet.",
"key": "ref13",
"unstructured": "Romano, J.D. & Tatonetti, N.P. Informatics and computational methods in natural product drug discovery: A review and Perspectives. Front. Genet. 10, 368 (2019).",
"volume": "10",
"year": "2019"
},
{
"article-title": "A database of collective molecular activities of useful plants",
"author": "Zeng X",
"first-page": "D1118-D1127",
"journal-title": "Nucleic Acids Res",
"key": "ref14",
"unstructured": "Zeng, X. et al. CMAUP: A database of collective molecular activities of useful plants. Nucleic Acids Res. 47, D1118-D1127 (2018).",
"volume": "CMAUP",
"year": "2018"
},
{
"DOI": "10.1038/s41586-020-2012-7",
"article-title": "A pneumonia outbreak associated with a new coronavirus of probable bat origin",
"author": "Zhou P",
"doi-asserted-by": "crossref",
"first-page": "270",
"journal-title": "Nature",
"key": "ref15",
"unstructured": "Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 579, 270–273 (2020).",
"volume": "579",
"year": "2020"
},
{
"DOI": "10.1038/s41586-020-2008-3",
"article-title": "A new coronavirus associated with human respiratory disease in China",
"author": "Wu F",
"doi-asserted-by": "crossref",
"first-page": "265",
"journal-title": "Nature.",
"key": "ref16",
"unstructured": "Wu, F. et al. A new coronavirus associated with human respiratory disease in China. Nature. 579, 265–269 (2020).",
"volume": "579",
"year": "2020"
},
{
"DOI": "10.1093/emboj/cdf327",
"article-title": "Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra α-helical domain",
"author": "Anand K",
"doi-asserted-by": "crossref",
"first-page": "3213",
"journal-title": "EMBO J.",
"key": "ref17",
"unstructured": "Anand, K. et al. Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra α-helical domain. EMBO J. 21, 3213–3224 (2002).",
"volume": "21",
"year": "2002"
},
{
"author": "Yang H",
"key": "ref18",
"unstructured": "Yang, H. et al. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc. Natl. Acad. Sci. 100, 13190–13195 (2003).",
"year": "2003"
},
{
"DOI": "10.1128/JVI.02114-07",
"article-title": "Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design",
"author": "Xue X",
"doi-asserted-by": "crossref",
"first-page": "2515",
"journal-title": "J. Virol.",
"key": "ref19",
"unstructured": "Xue, X. et al. Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design. J. Virol. 82, 2515–2527 (2008).",
"volume": "82",
"year": "2008"
},
{
"DOI": "10.1007/s13238-013-2841-3",
"article-title": "The newly emerged SARS-like coronavirus HCoV-EMC also has an \"Achilles' heel\": current effective inhibitor targeting a 3C-like protease",
"author": "Ren Z",
"doi-asserted-by": "crossref",
"first-page": "248",
"journal-title": "Protein Cell.",
"key": "ref20",
"unstructured": "Ren, Z. et al. The newly emerged SARS-like coronavirus HCoV-EMC also has an \"Achilles' heel\": current effective inhibitor targeting a 3C-like protease. Protein Cell. 4, 248–250 (2013).",
"volume": "4",
"year": "2013"
},
{
"DOI": "10.1038/srep22677",
"article-title": "Structure of main protease from human coronavirus NL63: insights for wide spectrum anti-coronavirus drug design",
"author": "Wang F",
"doi-asserted-by": "crossref",
"first-page": "22677",
"journal-title": "Sci. Rep.",
"key": "ref21",
"unstructured": "Wang, F. et al. Structure of main protease from human coronavirus NL63: insights for wide spectrum anti-coronavirus drug design. Sci. Rep. 6, 22677 (2016).",
"volume": "6",
"year": "2016"
},
{
"DOI": "10.1038/s41586-020-2223-y",
"article-title": "Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors.",
"author": "Jin Z",
"doi-asserted-by": "crossref",
"first-page": "289",
"journal-title": "Nature",
"key": "ref22",
"unstructured": "Jin, Z. et al. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature. 582, 289–293 (2020).",
"volume": "582",
"year": "2020"
},
{
"DOI": "10.1038/s41467-020-16954-7",
"article-title": "Structural plasticity of the SARS-COV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography",
"author": "Kneller DW",
"doi-asserted-by": "crossref",
"first-page": "3202",
"journal-title": "Nature Commun",
"key": "ref23",
"unstructured": "Kneller, D.W., Kovalevsky, A. & Coates, L. Structural plasticity of the SARS-COV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography. Nature Commun. 11, 3202 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.3390/ijms21093099",
"article-title": "Structural and Evolutionary Analysis Indicate That the SARS-CoV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design",
"author": "Bzówka M",
"doi-asserted-by": "crossref",
"first-page": "3099",
"journal-title": "Int. J. Mol. Sci.",
"key": "ref24",
"unstructured": "Bzówka, M., Mitusińska, K., Raczyńska, A., Samol, A., Tuszyński, J.A., & Góra, A. Structural and Evolutionary Analysis Indicate That the SARS-CoV-2 Mpro Is a Challenging Target for Small-Molecule Inhibitor Design. Int. J. Mol. Sci. 21, 3099 (2020).",
"volume": "21",
"year": "2020"
},
{
"DOI": "10.1021/acsptsci.0c00215",
"article-title": "A Blueprint for High Affinity SARS-CoV-2 Mpro Inhibitors from Activity-Based Compound Library Screening Guided by Analysis of Protein Dynamics",
"author": "Gossen J",
"doi-asserted-by": "crossref",
"first-page": "1079",
"journal-title": "ACS Pharmacol. Transl. Sci.",
"key": "ref25",
"unstructured": "Gossen, J. et al. A Blueprint for High Affinity SARS-CoV-2 Mpro Inhibitors from Activity-Based Compound Library Screening Guided by Analysis of Protein Dynamics. ACS Pharmacol. Transl. Sci. 4, 1079–1095 (2021).",
"volume": "4",
"year": "2021"
},
{
"DOI": "10.7554/eLife.77433",
"article-title": "Comprehensive fitness landscape of SARS-CoV-2 Mpro reveals insights into viral resistance mechanisms",
"author": "Flynn JM",
"doi-asserted-by": "crossref",
"first-page": "e77433",
"journal-title": "Elife.",
"key": "ref26",
"unstructured": "Flynn, J.M. et al. Comprehensive fitness landscape of SARS-CoV-2 Mpro reveals insights into viral resistance mechanisms. Elife. 11, e77433 (2022).",
"volume": "11",
"year": "2022"
},
{
"DOI": "10.1063/5.0013029",
"article-title": "Topological analysis of SARS CoV-2 main protease",
"author": "Estrada E",
"doi-asserted-by": "crossref",
"first-page": "061102",
"journal-title": "Chaos",
"key": "ref27",
"unstructured": "Estrada, E. Topological analysis of SARS CoV-2 main protease. Chaos. 30, 061102 (2020).",
"volume": "30",
"year": "2020"
},
{
"DOI": "10.1007/s12010-021-03608-7",
"article-title": "Macrolactin A as a Novel Inhibitory Agent for SARS-CoV-2 Mpro: Bioinformatics Approach",
"author": "Bharadwaj KK",
"doi-asserted-by": "crossref",
"first-page": "3371",
"journal-title": "Appl. Biochem. Biotechnol.",
"key": "ref28",
"unstructured": "Bharadwaj, K.K. et al. Macrolactin A as a Novel Inhibitory Agent for SARS-CoV-2 Mpro: Bioinformatics Approach. Appl. Biochem. Biotechnol. 193, 3371–3394 (2021).",
"volume": "193",
"year": "2021"
},
{
"DOI": "10.3389/fmolb.2020.603037",
"article-title": "Identification of Potential Inhibitors of 3CL Protease of SARS-CoV-2 From ZINC Database by Molecular Docking-Based Virtual Screening",
"author": "Abdusalam AAA",
"doi-asserted-by": "crossref",
"first-page": "603037",
"journal-title": "Front. Mol. Biosci.",
"key": "ref29",
"unstructured": "Abdusalam, A.A.A., & Murugaiyah, V. Identification of Potential Inhibitors of 3CL Protease of SARS-CoV-2 From ZINC Database by Molecular Docking-Based Virtual Screening. Front. Mol. Biosci. 7, 603037 (2020).",
"volume": "7",
"year": "2020"
},
{
"DOI": "10.1021/acsomega.0c04808",
"article-title": "Structure-Based Virtual Screening and Biochemical Validation to Discover a Potential Inhibitor of the SARS-CoV-2 Main Protease",
"author": "Gupta A",
"doi-asserted-by": "crossref",
"first-page": "33151",
"journal-title": "ACS Omega",
"key": "ref30",
"unstructured": "Gupta, A. et al. Structure-Based Virtual Screening and Biochemical Validation to Discover a Potential Inhibitor of the SARS-CoV-2 Main Protease. ACS Omega. 5, 33151–33161 (2020).",
"volume": "5",
"year": "2020"
},
{
"DOI": "10.1021/ct500381c",
"article-title": "POVME 2.0: An Enhanced Tool for Determining Pocket Shape and Volume Characteristics",
"author": "Durrant JD",
"doi-asserted-by": "crossref",
"first-page": "5047",
"journal-title": "J. Chem. Theory Comput",
"key": "ref31",
"unstructured": "Durrant, J.D. et al. POVME 2.0: An Enhanced Tool for Determining Pocket Shape and Volume Characteristics. J. Chem. Theory Comput. 10, 5047–5056 (2014).",
"volume": "10",
"year": "2014"
},
{
"DOI": "10.1021/acs.jctc.7b00500",
"article-title": "POVME 3.0: Software for Mapping Binding Pocket Flexibility",
"author": "Wagner JR",
"doi-asserted-by": "crossref",
"first-page": "4584",
"journal-title": "J. Chem. Theory Comput.",
"key": "ref32",
"unstructured": "Wagner, J.R. et al. POVME 3.0: Software for Mapping Binding Pocket Flexibility. J. Chem. Theory Comput. 13, 4584–4592 (2017).",
"volume": "13",
"year": "2017"
},
{
"DOI": "10.1016/j.str.2021.07.007",
"article-title": "Near-physiological-temperature serial crystallography reveals conformations of SARS-CoV-2 main protease active site for improved drug repurposing",
"author": "Durdagi S",
"doi-asserted-by": "crossref",
"first-page": "1382",
"journal-title": "Structure.",
"key": "ref33",
"unstructured": "Durdagi, S. et al. Near-physiological-temperature serial crystallography reveals conformations of SARS-CoV-2 main protease active site for improved drug repurposing. Structure. 29, 1382–1396 (2021).",
"volume": "29",
"year": "2021"
},
{
"DOI": "10.1021/acs.jcim.1c00140",
"article-title": "Elucidation of cryptic and allosteric pockets within the SARS-CoV-2 protease",
"author": "Sztain T",
"doi-asserted-by": "crossref",
"first-page": "3495",
"journal-title": "J. Chem. Inf. Model",
"key": "ref34",
"unstructured": "Sztain, T., Amaro, R., & McCammon, J.A. Elucidation of cryptic and allosteric pockets within the SARS-CoV-2 protease. J. Chem. Inf. Model. 61, 3495–3501 (2021).",
"volume": "61",
"year": "2021"
},
{
"DOI": "10.3389/fphar.2022.962863",
"article-title": "Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking",
"author": "Yang X",
"doi-asserted-by": "crossref",
"first-page": "962863",
"journal-title": "Front. Pharmacol.",
"key": "ref35",
"unstructured": "Yang, X. et al. Screening of potential inhibitors targeting the main protease structure of SARS-CoV-2 via molecular docking. Front. Pharmacol. 13, 962863 (2022).",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.1007/s12033-023-00831-x",
"author": "Fonseca AM",
"doi-asserted-by": "publisher",
"key": "ref36",
"unstructured": "da Fonseca, A.M. et al. Screening of Potential Inhibitors Targeting the Main Protease Structure of SARS-CoV-2 via Molecular Docking, and Approach with Molecular Dynamics, RMSD, RMSF, H-Bond, SASA and MMGBSA. Mol. Biotechnol. Preprint at 10.1007/s12033-023-00831-x (2023).",
"year": "2023"
},
{
"DOI": "10.1007/s11224-022-02031-w",
"article-title": "Fragment-based design of SARS-CoV-2 Mpro inhibitors",
"author": "Teli DM",
"doi-asserted-by": "crossref",
"first-page": "2155",
"journal-title": "Struct Chem",
"key": "ref37",
"unstructured": "Teli, D.M. et al. Fragment-based design of SARS-CoV-2 Mpro inhibitors. Struct Chem. 33, 2155–2168 (2022).",
"volume": "33",
"year": "2022"
},
{
"DOI": "10.1134/S0036024422070251",
"article-title": "Phytochemicals As a Potential Inhibitor of COVID-19: An In-Silico Perspective",
"author": "Jamhour RMAQ",
"doi-asserted-by": "crossref",
"first-page": "1589",
"journal-title": "Russ. J. Phys. Chem.",
"key": "ref38",
"unstructured": "Jamhour, R.M.A.Q. et al. Phytochemicals As a Potential Inhibitor of COVID-19: An In-Silico Perspective. Russ. J. Phys. Chem. 96, 1589–97 (2022).",
"volume": "96",
"year": "2022"
},
{
"article-title": "Phytochemicals Against SARS-COV-2 Infection",
"author": "Agrawal PK",
"journal-title": "Nat. Prod. Commun.",
"key": "ref39",
"unstructured": "Agrawal, P.K., & Blunden, G. Phytochemicals Against SARS-COV-2 Infection. Nat. Prod. Commun. 18, (2023).",
"volume": "18",
"year": "2023"
},
{
"DOI": "10.3389/fimmu.2020.01451",
"article-title": "Quercetin and vitamin C: an experimental, synergistic therapy for the prevention and treatment of SARS-CoV-2 related disease (COVID-19)",
"author": "Biancatelli RMLC",
"doi-asserted-by": "crossref",
"first-page": "1451",
"journal-title": "Front. Immunol.",
"key": "ref40",
"unstructured": "Biancatelli, R.M.L.C., Berrill, M., Catravas, J., & Marik, P.E. Quercetin and vitamin C: an experimental, synergistic therapy for the prevention and treatment of SARS-CoV-2 related disease (COVID-19). Front. Immunol. 11, 1451 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1016/j.virusres.2020.197989",
"article-title": "Natural product-derived phytochemicals as potential agents against coronaviruses: a review",
"author": "Mani JS",
"doi-asserted-by": "crossref",
"first-page": "197989",
"journal-title": "Virus Res.",
"key": "ref41",
"unstructured": "Mani J.S et al. Natural product-derived phytochemicals as potential agents against coronaviruses: a review. Virus Res. 284, 197989. (2020).",
"volume": "284",
"year": "2020"
},
{
"DOI": "10.3390/plants11141862",
"article-title": "The Main Protease of SARS-CoV-2 as a Target for Phytochemicals against Coronavirus",
"author": "Issa SS",
"doi-asserted-by": "crossref",
"first-page": "1862",
"journal-title": "Plants",
"key": "ref42",
"unstructured": "Issa S.S. et al. The Main Protease of SARS-CoV-2 as a Target for Phytochemicals against Coronavirus. Plants. 11, 1862 (2022).",
"volume": "11",
"year": "2022"
},
{
"key": "ref43",
"unstructured": "AP11.0, SimulationsPlus LLC, Lancaster, CA."
},
{
"DOI": "10.1016/j.compbiolchem.2022.107657",
"article-title": "Design and various in silico studies of the novel curcumin derivatives as potential candidates against COVID-19 -associated main enzymes",
"author": "Alici H",
"doi-asserted-by": "crossref",
"journal-title": "Comput. Biol. Chem.",
"key": "ref44",
"unstructured": "Alici, H., Tahtaci, H., & Demir, K. Design and various in silico studies of the novel curcumin derivatives as potential candidates against COVID-19 -associated main enzymes. Comput. Biol. Chem. (2022).",
"year": "2022"
},
{
"DOI": "10.7717/peerj.11590",
"article-title": "The impact of curcumin-derived polyphenols on the structure and flexibility COVID-19 main protease binding pocket: a molecular dynamics simulation study",
"author": "Mulu A",
"doi-asserted-by": "crossref",
"first-page": "e11590",
"journal-title": "PeerJ",
"key": "ref45",
"unstructured": "Mulu, A. et al. The impact of curcumin-derived polyphenols on the structure and flexibility COVID-19 main protease binding pocket: a molecular dynamics simulation study. PeerJ. 9, e11590 (2021).",
"volume": "9",
"year": "2021"
},
{
"DOI": "10.20944/preprints202003.0226.v1",
"author": "Khaerunnisa S",
"doi-asserted-by": "publisher",
"key": "ref46",
"unstructured": "Khaerunnisa, S., Kurniawan, H., Awaluddin, R., Suhartati, S., & Soetjipto, S. Potential Inhibitor of COVID-19 Main Protease (Mpro) From Several Medicinal Plant Compounds by Molecular Docking Study. Preprint at https://doi.org/10.20944/preprints202003.0226.v1 (2020).",
"year": "2020"
},
{
"DOI": "10.1007/s00894-021-04703-6",
"article-title": "Structure-activity relationship (SAR) and molecular dynamics study of withaferin-A fragment derivatives as a potential therapeutic lead against the main protease (Mpro) of SARS-CoV-2",
"author": "Ghosh A",
"doi-asserted-by": "crossref",
"first-page": "97",
"journal-title": "J. Mol. Model.",
"key": "ref47",
"unstructured": "Ghosh, A. et al. Structure-activity relationship (SAR) and molecular dynamics study of withaferin-A fragment derivatives as a potential therapeutic lead against the main protease (Mpro) of SARS-CoV-2. J. Mol. Model. 27, 97 (2021).",
"volume": "27",
"year": "2021"
},
{
"DOI": "10.1021/acs.jnatprod.2c00521",
"article-title": "The Natural Products Withaferin A and Withanone from the Medicinal Herb Withania somnifera Are Covalent Inhibitors of the SARS-CoV-2 Main Protease",
"author": "Chakraborty S",
"doi-asserted-by": "crossref",
"first-page": "2340",
"journal-title": "J. Nat. Prod.",
"key": "ref48",
"unstructured": "Chakraborty, S. et al. The Natural Products Withaferin A and Withanone from the Medicinal Herb Withania somnifera Are Covalent Inhibitors of the SARS-CoV-2 Main Protease. J. Nat. Prod. 85, 2340–2350 (2022).",
"volume": "85",
"year": "2022"
},
{
"DOI": "10.1128/JVI.01253-21",
"article-title": "Structure-Based Discovery and Structural Basis of a Novel Broad-Spectrum Natural Product against the Main Protease of Coronavirus",
"author": "Zhang Y",
"doi-asserted-by": "crossref",
"first-page": "e0125321",
"journal-title": "J. Virol.",
"key": "ref49",
"unstructured": "Zhang, Y. et al. Structure-Based Discovery and Structural Basis of a Novel Broad-Spectrum Natural Product against the Main Protease of Coronavirus. J. Virol. 96, e0125321 (2022).",
"volume": "96",
"year": "2022"
},
{
"DOI": "10.1021/acsptsci.0c00130",
"article-title": "PX-12, Tideglusib, and Shikonin Are Nonspecific Promiscuous SARS-CoV-2 Main Protease Inhibitors",
"author": "Ma C",
"doi-asserted-by": "crossref",
"first-page": "1265",
"journal-title": "ACS Pharmacol. Transl. Sci.",
"key": "ref50",
"unstructured": "Ma, C. et al. Ebselen, Disulfiram, Carmofur, PX-12, Tideglusib, and Shikonin Are Nonspecific Promiscuous SARS-CoV-2 Main Protease Inhibitors. ACS Pharmacol. Transl. Sci. 3, 1265–1277 (2020).",
"volume": "3",
"year": "2020"
},
{
"DOI": "10.1080/07391102.2020.1776157",
"article-title": "Identification of potential natural inhibitors of SARS-CoV2 main protease by molecular docking and simulation studies",
"author": "Gupta S",
"doi-asserted-by": "crossref",
"first-page": "4334",
"journal-title": "J. Biomol. Struct. Dyn.",
"key": "ref51",
"unstructured": "Gupta, S. et al. Identification of potential natural inhibitors of SARS-CoV2 main protease by molecular docking and simulation studies. J. Biomol. Struct. Dyn. 39, 4334–4345 (2021).",
"volume": "39",
"year": "2021"
},
{
"DOI": "10.1021/jm300288g",
"article-title": "Charting, navigating, and populating natural product chemical space for drug discovery",
"author": "Lachance H",
"doi-asserted-by": "crossref",
"first-page": "5989",
"journal-title": "J. Med. Chem.",
"key": "ref52",
"unstructured": "Lachance, H. et al. Charting, navigating, and populating natural product chemical space for drug discovery. J. Med. Chem. 55, 5989–6001 (2012).",
"volume": "55",
"year": "2012"
},
{
"DOI": "10.1007/s12298-022-01146-y",
"article-title": "How do plants defend themselves against pathogens-Biochemical mechanisms and genetic interventions",
"author": "Kaur S",
"doi-asserted-by": "crossref",
"first-page": "485",
"journal-title": "Physiol. Mol. Biol. Plants",
"key": "ref53",
"unstructured": "Kaur, S. et al. How do plants defend themselves against pathogens-Biochemical mechanisms and genetic interventions. Physiol. Mol. Biol. Plants. 28, 485–504 (2022).",
"volume": "28",
"year": "2022"
},
{
"key": "ref54",
"unstructured": "Protein Data Bank. Retrieved August 29, 2023, from https://www.rcsb.org"
},
{
"DOI": "10.2210/pdb6Z2E/pdb",
"article-title": "Crystal structure of SARS-CoV-2 Mpro in complex with the activity-based probe, biotin-PEG(4)",
"author": "Zhang L",
"doi-asserted-by": "publisher",
"key": "ref55",
"unstructured": "Zhang, L., & Hilgenfeld, R. Crystal structure of SARS-CoV-2 Mpro in complex with the activity-based probe, biotin-PEG(4)-Abu-Tle-Leu-Gln-vinylsulfone. https://doi.org/10.2210/pdb6Z2E/pdb (2020).",
"year": "2020"
},
{
"DOI": "10.1126/science.abb4489",
"article-title": "Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease",
"author": "Dai W",
"doi-asserted-by": "crossref",
"first-page": "1331",
"journal-title": "Science.",
"key": "ref56",
"unstructured": "Dai, W., Zhang, B., Jiang, X.M., Su, H., & Li, J. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science. 368, 1331–1335 (2020).",
"volume": "368",
"year": "2020"
},
{
"DOI": "10.1126/science.1085658",
"article-title": "Coronavirus main proteinase (3CLpro) structure: basis for the design of anti-SARS drugs",
"author": "Anand K",
"doi-asserted-by": "crossref",
"first-page": "1763",
"journal-title": "Science",
"key": "ref57",
"unstructured": "Anand, K., Ziebuhr, J., Wadhwani, P., Mesters, J.R., & Hilgenfeld, R. Coronavirus main proteinase (3CLpro) structure: basis for the design of anti-SARS drugs. Science. 300, 1763–1767 (2020).",
"volume": "300",
"year": "2020"
},
{
"DOI": "10.1371/journal.pone.0240653",
"article-title": "Potential bioactive glycosylated flavonoids as SARS-CoV-2 main protease inhibitors: A molecular docking and simulation studies",
"author": "Cherrak SA",
"doi-asserted-by": "crossref",
"first-page": "e0240653",
"journal-title": "PLoS One",
"key": "ref58",
"unstructured": "Cherrak, S.A., Merzouk, H., & Mokhtari-Soulimane, N. Potential bioactive glycosylated flavonoids as SARS-CoV-2 main protease inhibitors: A molecular docking and simulation studies. PLoS One. 15, e0240653 (2020).",
"volume": "15",
"year": "2020"
},
{
"DOI": "10.1038/s41467-020-18709-w",
"article-title": "Crystallographic and electrophilic fragment screening of the SARS-CoV-2 main protease",
"author": "Douangamath A",
"doi-asserted-by": "crossref",
"first-page": "5047",
"journal-title": "Nat. Commun.",
"key": "ref59",
"unstructured": "Douangamath, A., Fearon, D., Gehrtz, P., Krojer, T., & Lukacik, P. Crystallographic and electrophilic fragment screening of the SARS-CoV-2 main protease. Nat. Commun. 11, 5047 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1016/j.softx.2015.06.001",
"article-title": "High performance molecular simulations through multi-level parallelism from laptops to supercomputers",
"author": "Abraham MJ",
"doi-asserted-by": "crossref",
"first-page": "19",
"journal-title": "SoftwareX",
"key": "ref60",
"unstructured": "Abraham, M.J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX. 1, 19–25 (2015).",
"volume": "GROMACS",
"year": "2015"
},
{
"DOI": "10.1002/jcc.23354",
"article-title": "CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data",
"author": "Huang J",
"doi-asserted-by": "crossref",
"first-page": "2135",
"journal-title": "J. Comput. Chem.",
"key": "ref61",
"unstructured": "Huang, J., & MacKerell, A.D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).",
"volume": "34",
"year": "2013"
},
{
"DOI": "10.1063/1.445869",
"article-title": "Comparison of simple potential functions for simulating liquid water",
"author": "Jorgensen WL",
"doi-asserted-by": "crossref",
"first-page": "926",
"journal-title": "J. Chem. Phys.",
"key": "ref62",
"unstructured": "Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W., & Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926 (1983).",
"volume": "79",
"year": "1983"
},
{
"DOI": "10.1063/1.464397",
"article-title": "Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems",
"author": "Darden T",
"doi-asserted-by": "crossref",
"first-page": "10089",
"journal-title": "J. Chem. Phys.",
"key": "ref63",
"unstructured": "Darden, T., York, D., & Pedersen, L. Particle mesh Ewald: An N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).",
"volume": "98",
"year": "1993"
},
{
"DOI": "10.1111/cbdd.13653",
"article-title": "Impact of lymphoma-linked Asn11Tyr point mutation on the interaction between Bcl-2 and a BH3 mimetic: Insights from molecular dynamics simulation",
"author": "Singh K",
"doi-asserted-by": "crossref",
"first-page": "435",
"journal-title": "Chem. Biol. Drug Design.",
"key": "ref64",
"unstructured": "Singh,K. & Briggs, J.M. Impact of lymphoma-linked Asn11Tyr point mutation on the interaction between Bcl-2 and a BH3 mimetic: Insights from molecular dynamics simulation. Chem. Biol. Drug Design. 95, 435–450 (2020).",
"volume": "95",
"year": "2020"
},
{
"DOI": "10.1063/1.448118",
"article-title": "Molecular dynamics with coupling to an external bath",
"author": "Berendsen H",
"doi-asserted-by": "crossref",
"first-page": "3684",
"journal-title": "J. Chem. Phys.",
"key": "ref65",
"unstructured": "Berendsen, H., Postma, J., van Gunsteren, W., DiNola, A., & Haak, J. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).",
"volume": "81",
"year": "1984"
},
{
"DOI": "10.1002/(SICI)1096-987X(199709)18:12<1463::AID-JCC4>3.0.CO;2-H",
"article-title": "E. M. LINCS: A linear constraint solver for molecular simulations",
"author": "Hess B",
"doi-asserted-by": "crossref",
"journal-title": "J. Comp. Chem.",
"key": "ref66",
"unstructured": "Hess, B., Bekker, H., Berendsen, H. J. C., & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Comp. Chem. 18, (1997).",
"volume": "18",
"year": "1997"
},
{
"DOI": "10.1016/0263-7855(96)00018-5",
"article-title": "VMD - Visual Molecular Dynamics",
"author": "Humphrey W",
"doi-asserted-by": "crossref",
"first-page": "33",
"journal-title": "J. Mol. Graphics.",
"key": "ref67",
"unstructured": "Humphrey, W., Dalke, A., & Schulten, K. VMD - Visual Molecular Dynamics. J. Mol. Graphics. 14, 33–38 (1996).",
"volume": "14",
"year": "1996"
},
{
"DOI": "10.1002/jcc.20084",
"article-title": "UCSF Chimera - A visualization system for exploratory research and analysis",
"author": "Pettersen E",
"doi-asserted-by": "crossref",
"first-page": "1605",
"journal-title": "J. Comp. Chem.",
"key": "ref68",
"unstructured": "Pettersen, E. et al. UCSF Chimera - A visualization system for exploratory research and analysis. J. Comp. Chem. 25, 1605–1612 (2004).",
"volume": "25",
"year": "2004"
},
{
"author": "Schrödinger Release 2022-4",
"key": "ref69",
"unstructured": "Schrödinger Release 2022-4: Protein Preparation Wizard; Epik, Schrödinger, LLC, New York, NY, 2022.",
"volume-title": "Protein Preparation Wizard; Epik, Schrödinger",
"year": "2022"
},
{
"DOI": "10.1002/jcc.20292",
"article-title": "Integrated modeling program applied chemical theory (IMPACT)",
"author": "Banks JL",
"doi-asserted-by": "crossref",
"first-page": "1752",
"journal-title": "J. Comp. Chem.",
"key": "ref70",
"unstructured": "Banks, J.L. et al. Integrated modeling program applied chemical theory (IMPACT). J. Comp. Chem. 26, 1752–1780 (2005).",
"volume": "26",
"year": "2005"
},
{
"key": "ref71",
"unstructured": "Schrödinger Release 2022-4: Glide, Schrödinger, LLC, New York, NY, 2022"
},
{
"author": "Schrödinger",
"key": "ref72",
"unstructured": "Schrödinger Release 2022-4: LigPrep, Schrödinger, LLC, New York, NY, 2022",
"year": "2022"
},
{
"DOI": "10.1002/prot.23106",
"article-title": "The VSGB 2.0 model: a next-generation energy model for high-resolution protein structure modeling",
"author": "Li J",
"doi-asserted-by": "crossref",
"first-page": "2794",
"journal-title": "Proteins.",
"key": "ref73",
"unstructured": "Li, J., Abel, R., Zhu, K., Cao, Y., Zhao, S., & Friesner RA. The VSGB 2.0 model: a next-generation energy model for high-resolution protein structure modeling. Proteins. 79, 2794–2812 (2011).",
"volume": "79",
"year": "2011"
},
{
"DOI": "10.1016/j.scitotenv.2013.06.081",
"article-title": "Validation of quantitative structure-activity relationship models to predict water-solubility of organic compounds",
"author": "Cappelli CI",
"doi-asserted-by": "crossref",
"first-page": "781",
"journal-title": "Sci. Total Environ",
"key": "ref74",
"unstructured": "Cappelli, C.I., Manganelli, S., Lombardo, A., Gissi, A., & Benfenati, E. Validation of quantitative structure-activity relationship models to predict water-solubility of organic compounds. Sci. Total Environ. 463–464, 781–789 (2013).",
"volume": "463–464",
"year": "2013"
},
{
"DOI": "10.1517/17460441.1.1.31",
"article-title": "In silico prediction of aqueous solubility",
"author": "Dearden JC",
"doi-asserted-by": "crossref",
"first-page": "31",
"journal-title": "Expet Opin. Drug Discov",
"key": "ref75",
"unstructured": "Dearden, J.C. In silico prediction of aqueous solubility. Expet Opin. Drug Discov. 1, 31–52 (2006).",
"volume": "1",
"year": "2006"
}
],
"reference-count": 75,
"references-count": 75,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.researchsquare.com/article/rs-3888947/v1"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subtitle": [],
"subtype": "preprint",
"title": "Unlocking the potential of phytochemicals in inhibiting SARS-CoV-2 M Pro protein - An in-silico and cell-based approach",
"type": "posted-content"
}
singh11
