In Silico Comparative Analysis of Ivermectin and Nirmatrelvir Inhibitors Interacting with the SARS-CoV-2 Main Protease
et al., Preprints, doi:10.20944/preprints202404.1825.v1, Apr 2024
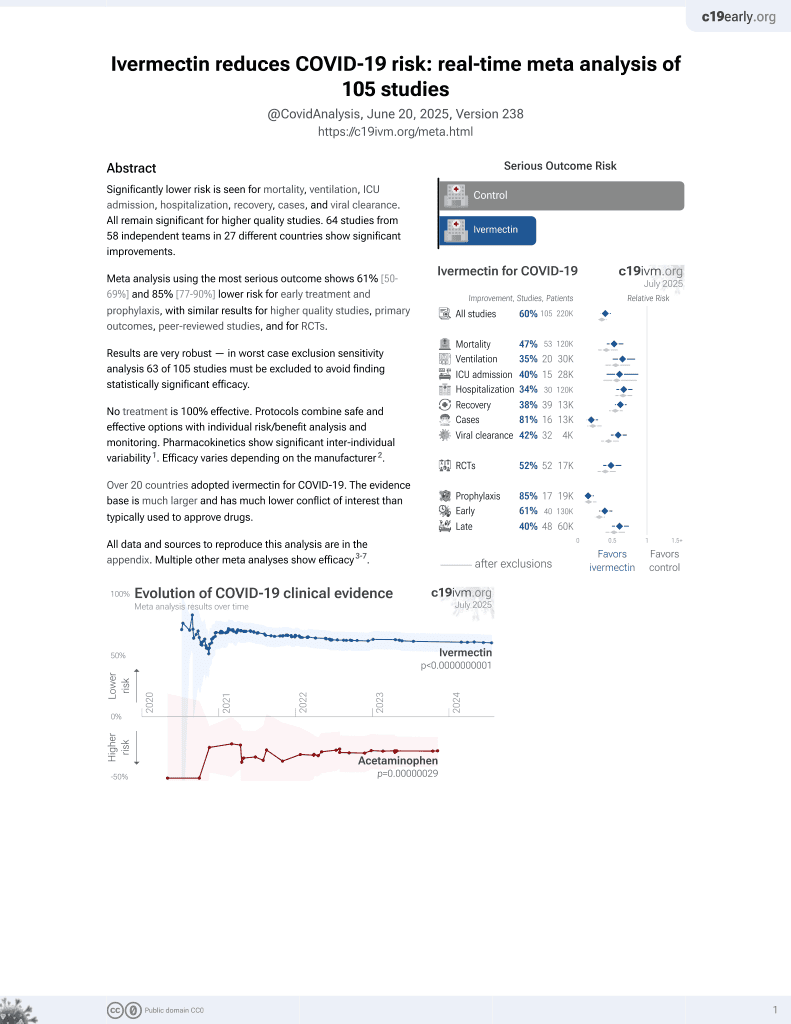
Ivermectin for COVID-19
4th treatment shown to reduce risk in
August 2020, now with p < 0.00000000001 from 106 studies, recognized in 24 countries.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
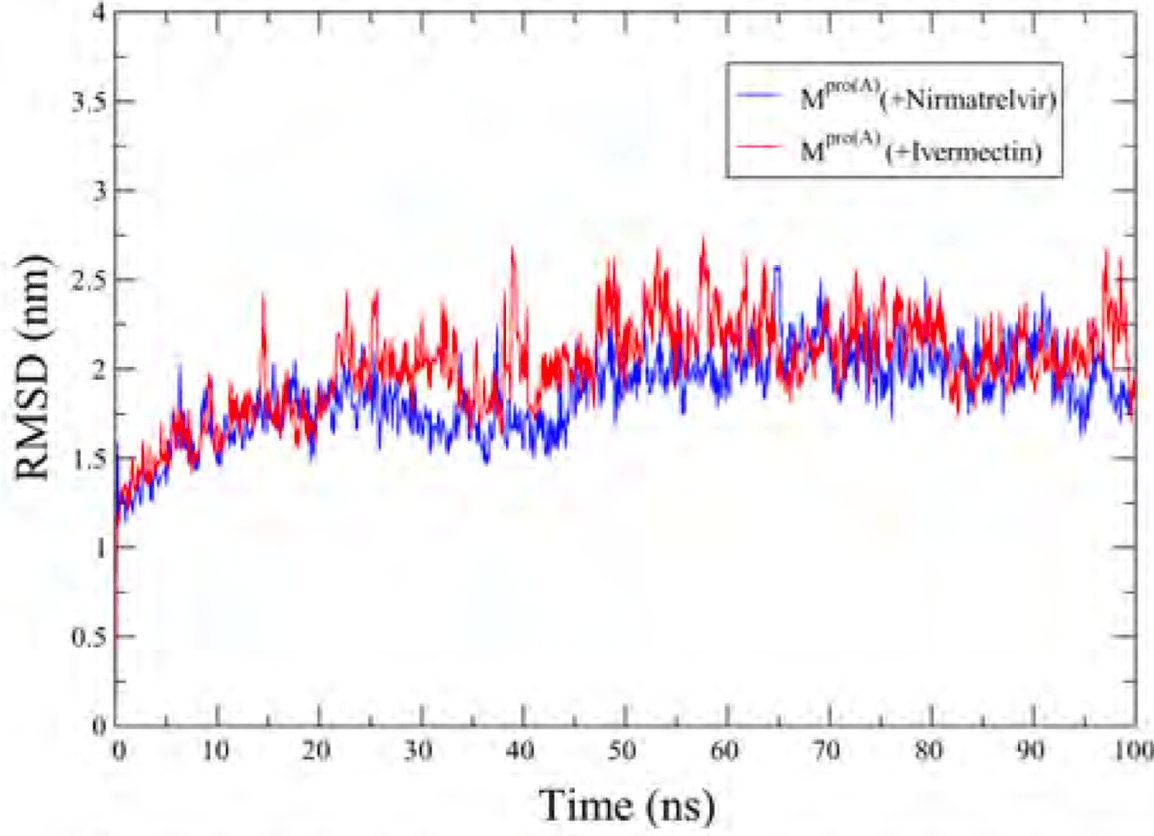
In silico study showing that ivermectin and nirmatrelvir interact with the SARS-CoV-2 main protease (Mpro). Authors used molecular docking and 100ns molecular dynamics simulations to investigate the binding interactions. Nirmatrelvir formed strong interactions with multiple Mpro amino acids, showing stable binding with an RMSD of around 2.0Å. Ivermectin also interacted with multiple amino acids, displaying an RMSD of 1.87Å. Ivermectin interacted with approximately the same set of amino acids at both 50ns and 100ns, demonstrating enduring interaction patterns over time. Both ligands stabilized Mpro, with ivermectin showing stability and persistent interactions despite forming fewer hydrogen bonds. Ivermectin primarily utilized alkyl and pi-alkyl interactions, indicating a different binding mode. Alkyl and pi-alkyl interactions are generally less specific which may improve the ability to retain efficacy for variants.
76 preclinical studies support the efficacy of ivermectin for COVID-19:
Ivermectin, better known for antiparasitic activity, is a broad spectrum antiviral with activity against many viruses including H7N773, Dengue38,74,75 , HIV-175, Simian virus 4076, Zika38,77,78 , West Nile78, Yellow Fever79,80, Japanese encephalitis79, Chikungunya80, Semliki Forest virus80, Human papillomavirus58, Epstein-Barr58, BK Polyomavirus81, and Sindbis virus80.
Ivermectin inhibits importin-α/β-dependent nuclear import of viral proteins73,75,76,82 , shows spike-ACE2 disruption at 1nM with microfluidic diffusional sizing39, binds to glycan sites on the SARS-CoV-2 spike protein preventing interaction with blood and epithelial cells and inhibiting hemagglutination42,83, shows dose-dependent inhibition of wildtype and omicron variants37, exhibits dose-dependent inhibition of lung injury63,68, may inhibit SARS-CoV-2 via IMPase inhibition38, may inhibit SARS-CoV-2 induced formation of fibrin clots resistant to degradation10, inhibits SARS-CoV-2 3CLpro55, may inhibit SARS-CoV-2 RdRp activity1,29, may minimize viral myocarditis by inhibiting NF-κB/p65-mediated inflammation in macrophages62, may be beneficial for COVID-19 ARDS by blocking GSDMD and NET formation84, may interfere with SARS-CoV-2's immune evasion via ORF8 binding5, may inhibit SARS-CoV-2 by disrupting CD147 interaction85-88, may inhibit SARS-CoV-2 attachment to lipid rafts via spike NTD binding3, shows protection against inflammation, cytokine storm, and mortality in an LPS mouse model sharing key pathological features of severe COVID-1960,89, may be beneficial in severe COVID-19 by binding IGF1 to inhibit the promotion of inflammation, fibrosis, and cell proliferation that leads to lung damage9, significantly mitigates bleomycin-induced pulmonary fibrosis by reducing collagen accumulation and inflammatory cell infiltration61, improves oxidative stress markers while suppressing myofibroblast proliferation61, may minimize SARS-CoV-2 induced cardiac damage41,49, may counter immune evasion by inhibiting NSP15-TBK1/KPNA1 interaction and restoring IRF3 activation90, may disrupt SARS-CoV-2 N and ORF6 protein nuclear transport and their suppression of host interferon responses2, reduces TAZ/YAP nuclear import, relieving SARS-CoV-2-driven suppression of IRF3 and NF-κB antiviral pathways36, increases Bifidobacteria which play a key role in the immune system91, has immunomodulatory52 and anti-inflammatory72,92 properties, and has an extensive and very positive safety profile93.
Study covers ivermectin and paxlovid.
1.
Li et al., Drug–Target Interaction Prediction via Dual-Interaction Fusion, Molecules, doi:10.3390/molecules31030498.
2.
Gayozo et al., Binding affinities analysis of ivermectin, nucleocapsid and ORF6 proteins of SARS-CoV-2 to human importins α isoforms: A computational approach, Biotecnia, doi:10.18633/biotecnia.v27.2485.
3.
Lefebvre et al., Characterization and Fluctuations of an Ivermectin Binding Site at the Lipid Raft Interface of the N-Terminal Domain (NTD) of the Spike Protein of SARS-CoV-2 Variants, Viruses, doi:10.3390/v16121836.
4.
Haque et al., Exploring potential therapeutic candidates against COVID-19: a molecular docking study, Discover Molecules, doi:10.1007/s44345-024-00005-5.
5.
Bagheri-Far et al., Non-spike protein inhibition of SARS-CoV-2 by natural products through the key mediator protein ORF8, Molecular Biology Research Communications, doi:10.22099/mbrc.2024.50245.2001.
6.
de Oliveira Só et al., In Silico Comparative Analysis of Ivermectin and Nirmatrelvir Inhibitors Interacting with the SARS-CoV-2 Main Protease, Preprints, doi:10.20944/preprints202404.1825.v1.
7.
Agamah et al., Network-based multi-omics-disease-drug associations reveal drug repurposing candidates for COVID-19 disease phases, ScienceOpen, doi:10.58647/DRUGARXIV.PR000010.v1.
8.
Oranu et al., Validation of the binding affinities and stabilities of ivermectin and moxidectin against SARS-CoV-2 receptors using molecular docking and molecular dynamics simulation, GSC Biological and Pharmaceutical Sciences, doi:10.30574/gscbps.2024.26.1.0030.
9.
Zhao et al., Identification of the shared gene signatures between pulmonary fibrosis and pulmonary hypertension using bioinformatics analysis, Frontiers in Immunology, doi:10.3389/fimmu.2023.1197752.
10.
Vottero et al., Computational Prediction of the Interaction of Ivermectin with Fibrinogen, Molecular Sciences, doi:10.3390/ijms241411449.
11.
Chellasamy et al., Docking and molecular dynamics studies of human ezrin protein with a modelled SARS-CoV-2 endodomain and their interaction with potential invasion inhibitors, Journal of King Saud University - Science, doi:10.1016/j.jksus.2022.102277.
12.
Umar et al., Inhibitory potentials of ivermectin, nafamostat, and camostat on spike protein and some nonstructural proteins of SARS-CoV-2: Virtual screening approach, Jurnal Teknologi Laboratorium, doi:10.29238/teknolabjournal.v11i1.344.
13.
Alvarado et al., Interaction of the New Inhibitor Paxlovid (PF-07321332) and Ivermectin With the Monomer of the Main Protease SARS-CoV-2: A Volumetric Study Based on Molecular Dynamics, Elastic Networks, Classical Thermodynamics and SPT, Computational Biology and Chemistry, doi:10.1016/j.compbiolchem.2022.107692.
14.
Aminpour et al., In Silico Analysis of the Multi-Targeted Mode of Action of Ivermectin and Related Compounds, Computation, doi:10.3390/computation10040051.
15.
Parvez et al., Insights from a computational analysis of the SARS-CoV-2 Omicron variant: Host–pathogen interaction, pathogenicity, and possible drug therapeutics, Immunity, Inflammation and Disease, doi:10.1002/iid3.639.
16.
Francés-Monerris et al., Microscopic interactions between ivermectin and key human and viral proteins involved in SARS-CoV-2 infection, Physical Chemistry Chemical Physics, doi:10.1039/D1CP02967C.
17.
González-Paz et al., Comparative study of the interaction of ivermectin with proteins of interest associated with SARS-CoV-2: A computational and biophysical approach, Biophysical Chemistry, doi:10.1016/j.bpc.2021.106677.
18.
González-Paz (B) et al., Structural Deformability Induced in Proteins of Potential Interest Associated with COVID-19 by binding of Homologues present in Ivermectin: Comparative Study Based in Elastic Networks Models, Journal of Molecular Liquids, doi:10.1016/j.molliq.2021.117284.
19.
Rana et al., Ivermectin and Doxycycline combination as a promising anti-viral drug candidate: an in-silico and DFT study, In Silico Pharmacology, doi:10.1007/s40203-026-00576-w.
20.
Muthusamy et al., Virtual Screening Reveals Potential Anti-Parasitic Drugs Inhibiting the Receptor Binding Domain of SARS-CoV-2 Spike protein, Journal of Virology & Antiviral Research, www.scitechnol.com/abstract/virtual-screening-reveals-potential-antiparasitic-drugs-inhibiting-the-receptor-binding-domain-of-sarscov2-spike-protein-16398.html.
21.
Qureshi et al., Mechanistic insights into the inhibitory activity of FDA approved ivermectin against SARS-CoV-2: old drug with new implications, Journal of Biomolecular Structure and Dynamics, doi:10.1080/07391102.2021.1906750.
22.
Schöning et al., Highly-transmissible Variants of SARS-CoV-2 May Be More Susceptible to Drug Therapy Than Wild Type Strains, Research Square, doi:10.21203/rs.3.rs-379291/v1.
23.
Bello et al., Elucidation of the inhibitory activity of ivermectin with host nuclear importin α and several SARS-CoV-2 targets, Journal of Biomolecular Structure and Dynamics, doi:10.1080/07391102.2021.1911857.
24.
Udofia et al., In silico studies of selected multi-drug targeting against 3CLpro and nsp12 RNA-dependent RNA-polymerase proteins of SARS-CoV-2 and SARS-CoV, Network Modeling Analysis in Health Informatics and Bioinformatics, doi:10.1007/s13721-021-00299-2.
25.
Choudhury et al., Exploring the binding efficacy of ivermectin against the key proteins of SARS-CoV-2 pathogenesis: an in silico approach, Future Medicine, doi:10.2217/fvl-2020-0342.
26.
Kern et al., Modeling of SARS-CoV-2 Treatment Effects for Informed Drug Repurposing, Frontiers in Pharmacology, doi:10.3389/fphar.2021.625678.
27.
Saha et al., The Binding mechanism of ivermectin and levosalbutamol with spike protein of SARS-CoV-2, Structural Chemistry, doi:10.1007/s11224-021-01776-0.
28.
Eweas et al., Molecular Docking Reveals Ivermectin and Remdesivir as Potential Repurposed Drugs Against SARS-CoV-2, Frontiers in Microbiology, doi:10.3389/fmicb.2020.592908.
29.
Parvez (B) et al., Prediction of potential inhibitors for RNA-dependent RNA polymerase of SARS-CoV-2 using comprehensive drug repurposing and molecular docking approach, International Journal of Biological Macromolecules, doi:10.1016/j.ijbiomac.2020.09.098.
30.
Francés-Monerris (B) et al., Has Ivermectin Virus-Directed Effects against SARS-CoV-2? Rationalizing the Action of a Potential Multitarget Antiviral Agent, ChemRxiv, doi:10.26434/chemrxiv.12782258.v1.
31.
Kalhor et al., Repurposing of the approved small molecule drugs in order to inhibit SARS-CoV-2 S protein and human ACE2 interaction through virtual screening approaches, Journal of Biomolecular Structure and Dynamics, doi:10.1080/07391102.2020.1824816.
32.
Swargiary, A., Ivermectin as a promising RNA-dependent RNA polymerase inhibitor and a therapeutic drug against SARS-CoV2: Evidence from in silico studies, Research Square, doi:10.21203/rs.3.rs-73308/v1.
33.
Maurya, D., A Combination of Ivermectin and Doxycycline Possibly Blocks the Viral Entry and Modulate the Innate Immune Response in COVID-19 Patients, American Chemical Society (ACS), doi:10.26434/chemrxiv.12630539.v1.
34.
Lehrer et al., Ivermectin Docks to the SARS-CoV-2 Spike Receptor-binding Domain Attached to ACE2, In Vivo, 34:5, 3023-3026, doi:10.21873/invivo.12134.
35.
Suravajhala et al., Comparative Docking Studies on Curcumin with COVID-19 Proteins, Preprints, doi:10.20944/preprints202005.0439.v3.
36.
Kofler et al., M-Motif, a potential non-conventional NLS in YAP/TAZ and other cellular and viral proteins that inhibits classic protein import, iScience, doi:10.1016/j.isci.2025.112105.
37.
Shahin et al., The selective effect of Ivermectin on different human coronaviruses; in-vitro study, Research Square, doi:10.21203/rs.3.rs-4180797/v1.
38.
Jitobaom et al., Identification of inositol monophosphatase as a broad‐spectrum antiviral target of ivermectin, Journal of Medical Virology, doi:10.1002/jmv.29552.
39.
Fauquet et al., Microfluidic Diffusion Sizing Applied to the Study of Natural Products and Extracts That Modulate the SARS-CoV-2 Spike RBD/ACE2 Interaction, Molecules, doi:10.3390/molecules28248072.
40.
García-Aguilar et al., In Vitro Analysis of SARS-CoV-2 Spike Protein and Ivermectin Interaction, International Journal of Molecular Sciences, doi:10.3390/ijms242216392.
41.
Liu et al., SARS-CoV-2 viral genes Nsp6, Nsp8, and M compromise cellular ATP levels to impair survival and function of human pluripotent stem cell-derived cardiomyocytes, Stem Cell Research & Therapy, doi:10.1186/s13287-023-03485-3.
42.
Boschi et al., SARS-CoV-2 Spike Protein Induces Hemagglutination: Implications for COVID-19 Morbidities and Therapeutics and for Vaccine Adverse Effects, bioRxiv, doi:10.1101/2022.11.24.517882.
43.
De Forni et al., Synergistic drug combinations designed to fully suppress SARS-CoV-2 in the lung of COVID-19 patients, PLoS ONE, doi:10.1371/journal.pone.0276751.
44.
Saha (B) et al., Manipulation of Spray-Drying Conditions to Develop an Inhalable Ivermectin Dry Powder, Pharmaceutics, doi:10.3390/pharmaceutics14071432.
45.
Jitobaom (B) et al., Synergistic anti-SARS-CoV-2 activity of repurposed anti-parasitic drug combinations, BMC Pharmacology and Toxicology, doi:10.1186/s40360-022-00580-8.
46.
Croci et al., Liposomal Systems as Nanocarriers for the Antiviral Agent Ivermectin, International Journal of Biomaterials, doi:10.1155/2016/8043983.
47.
Zheng et al., Red blood cell-hitchhiking mediated pulmonary delivery of ivermectin: Effects of nanoparticle properties, International Journal of Pharmaceutics, doi:10.1016/j.ijpharm.2022.121719.
48.
Delandre et al., Antiviral Activity of Repurposing Ivermectin against a Panel of 30 Clinical SARS-CoV-2 Strains Belonging to 14 Variants, Pharmaceuticals, doi:10.3390/ph15040445.
49.
Liu (B) et al., Genome-wide analyses reveal the detrimental impacts of SARS-CoV-2 viral gene Orf9c on human pluripotent stem cell-derived cardiomyocytes, Stem Cell Reports, doi:10.1016/j.stemcr.2022.01.014.
50.
Segatori et al., Effect of Ivermectin and Atorvastatin on Nuclear Localization of Importin Alpha and Drug Target Expression Profiling in Host Cells from Nasopharyngeal Swabs of SARS-CoV-2- Positive Patients, Viruses, doi:10.3390/v13102084.
51.
Jitobaom (C) et al., Favipiravir and Ivermectin Showed in Vitro Synergistic Antiviral Activity against SARS-CoV-2, Research Square, doi:10.21203/rs.3.rs-941811/v1.
52.
Munson et al., Niclosamide and ivermectin modulate caspase-1 activity and proinflammatory cytokine secretion in a monocytic cell line, British Society For Nanomedicine Early Career Researcher Summer Meeting, 2021, web.archive.org/web/20230401070026/https://michealmunson.github.io/COVID.pdf.
53.
Mountain Valley MD, Mountain Valley MD Receives Successful Results From BSL-4 COVID-19 Clearance Trial on Three Variants Tested With Ivectosol™, 5/18, www.globenewswire.com/en/news-release/2021/05/18/2231755/0/en/Mountain-Valley-MD-Receives-Successful-Results-From-BSL-4-COVID-19-Clearance-Trial-on-Three-Variants-Tested-With-Ivectosol.html.
54.
Yesilbag et al., Ivermectin also inhibits the replication of bovine respiratory viruses (BRSV, BPIV-3, BoHV-1, BCoV and BVDV) in vitro, Virus Research, doi:10.1016/j.virusres.2021.198384.
55.
Mody et al., Identification of 3-chymotrypsin like protease (3CLPro) inhibitors as potential anti-SARS-CoV-2 agents, Communications Biology, doi:10.1038/s42003-020-01577-x.
56.
Jeffreys et al., Remdesivir-ivermectin combination displays synergistic interaction with improved in vitro activity against SARS-CoV-2, International Journal of Antimicrobial Agents, doi:10.1016/j.ijantimicag.2022.106542.
57.
Surnar et al., Clinically Approved Antiviral Drug in an Orally Administrable Nanoparticle for COVID-19, ACS Pharmacol. Transl. Sci., doi:10.1021/acsptsci.0c00179.
58.
Li (B) et al., Quantitative proteomics reveals a broad-spectrum antiviral property of ivermectin, benefiting for COVID-19 treatment, J. Cellular Physiology, doi:10.1002/jcp.30055.
59.
Caly et al., The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro, Antiviral Research, doi:10.1016/j.antiviral.2020.104787.
60.
Zhang et al., Ivermectin inhibits LPS-induced production of inflammatory cytokines and improves LPS-induced survival in mice, Inflammation Research, doi:10.1007/s00011-008-8007-8.
61.
Zaher et al., Testing the Possible Protective Effect of Ivermectin on Bleomycin-Induced Pulmonary Fibrosis in Albino Rats: Histological and Immunohistochemical Study, Medicina, doi:10.3390/medicina62030560.
62.
Gao et al., Ivermectin ameliorates acute myocarditis via the inhibition of importin-mediated nuclear translocation of NF-κB/p65, International Immunopharmacology, doi:10.1016/j.intimp.2024.112073.
63.
Abd-Elmawla et al., Suppression of NLRP3 inflammasome by ivermectin ameliorates bleomycin-induced pulmonary fibrosis, Journal of Zhejiang University-SCIENCE B, doi:10.1631/jzus.B2200385.
64.
Uematsu et al., Prophylactic administration of ivermectin attenuates SARS-CoV-2 induced disease in a Syrian Hamster Model, The Journal of Antibiotics, doi:10.1038/s41429-023-00623-0.
65.
Albariqi et al., Pharmacokinetics and Safety of Inhaled Ivermectin in Mice as a Potential COVID-19 Treatment, International Journal of Pharmaceutics, doi:10.1016/j.ijpharm.2022.121688.
66.
Errecalde et al., Safety and Pharmacokinetic Assessments of a Novel Ivermectin Nasal Spray Formulation in a Pig Model, Journal of Pharmaceutical Sciences, doi:10.1016/j.xphs.2021.01.017.
67.
Madrid et al., Safety of oral administration of high doses of ivermectin by means of biocompatible polyelectrolytes formulation, Heliyon, doi:10.1016/j.heliyon.2020.e05820.
68.
Ma et al., Ivermectin contributes to attenuating the severity of acute lung injury in mice, Biomedicine & Pharmacotherapy, doi:10.1016/j.biopha.2022.113706.
69.
de Melo et al., Attenuation of clinical and immunological outcomes during SARS-CoV-2 infection by ivermectin, EMBO Mol. Med., doi:10.15252/emmm.202114122.
70.
Arévalo et al., Ivermectin reduces in vivo coronavirus infection in a mouse experimental model, Scientific Reports, doi:10.1038/s41598-021-86679-0.
71.
Chaccour et al., Nebulized ivermectin for COVID-19 and other respiratory diseases, a proof of concept, dose-ranging study in rats, Scientific Reports, doi:10.1038/s41598-020-74084-y.
72.
Yan et al., Anti-inflammatory effects of ivermectin in mouse model of allergic asthma, Inflammation Research, doi:10.1007/s00011-011-0307-8.
73.
Götz et al., Influenza A viruses escape from MxA restriction at the expense of efficient nuclear vRNP import, Scientific Reports, doi:10.1038/srep23138.
74.
Tay et al., Nuclear localization of dengue virus (DENV) 1–4 non-structural protein 5; protection against all 4 DENV serotypes by the inhibitor Ivermectin, Antiviral Research, doi:10.1016/j.antiviral.2013.06.002.
75.
Wagstaff et al., Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus, Biochemical Journal, doi:10.1042/BJ20120150.
76.
Wagstaff (B) et al., An AlphaScreen®-Based Assay for High-Throughput Screening for Specific Inhibitors of Nuclear Import, SLAS Discovery, doi:10.1177/1087057110390360.
77.
Barrows et al., A Screen of FDA-Approved Drugs for Inhibitors of Zika Virus Infection, Cell Host & Microbe, doi:10.1016/j.chom.2016.07.004.
78.
Yang et al., The broad spectrum antiviral ivermectin targets the host nuclear transport importin α/β1 heterodimer, Antiviral Research, doi:10.1016/j.antiviral.2020.104760.
79.
Mastrangelo et al., Ivermectin is a potent inhibitor of flavivirus replication specifically targeting NS3 helicase activity: new prospects for an old drug, Journal of Antimicrobial Chemotherapy, doi:10.1093/jac/dks147.
80.
Varghese et al., Discovery of berberine, abamectin and ivermectin as antivirals against chikungunya and other alphaviruses, Antiviral Research, doi:10.1016/j.antiviral.2015.12.012.
81.
Bennett et al., Role of a nuclear localization signal on the minor capsid Proteins VP2 and VP3 in BKPyV nuclear entry, Virology, doi:10.1016/j.virol.2014.10.013.
82.
Kosyna et al., The importin α/β-specific inhibitor Ivermectin affects HIF-dependent hypoxia response pathways, Biological Chemistry, doi:10.1515/hsz-2015-0171.
83.
Scheim et al., Sialylated Glycan Bindings from SARS-CoV-2 Spike Protein to Blood and Endothelial Cells Govern the Severe Morbidities of COVID-19, International Journal of Molecular Sciences, doi:10.3390/ijms242317039.
84.
Liu (C) et al., Crosstalk between neutrophil extracellular traps and immune regulation: insights into pathobiology and therapeutic implications of transfusion-related acute lung injury, Frontiers in Immunology, doi:10.3389/fimmu.2023.1324021.
85.
Shouman et al., SARS-CoV-2-associated lymphopenia: possible mechanisms and the role of CD147, Cell Communication and Signaling, doi:10.1186/s12964-024-01718-3.
86.
Scheim (B), D., Ivermectin for COVID-19 Treatment: Clinical Response at Quasi-Threshold Doses Via Hypothesized Alleviation of CD147-Mediated Vascular Occlusion, SSRN, doi:10.2139/ssrn.3636557.
87.
Scheim (C), D., From Cold to Killer: How SARS-CoV-2 Evolved without Hemagglutinin Esterase to Agglutinate and Then Clot Blood Cells, Center for Open Science, doi:10.31219/osf.io/sgdj2.
88.
Behl et al., CD147-spike protein interaction in COVID-19: Get the ball rolling with a novel receptor and therapeutic target, Science of The Total Environment, doi:10.1016/j.scitotenv.2021.152072.
89.
DiNicolantonio et al., Ivermectin may be a clinically useful anti-inflammatory agent for late-stage COVID-19, Open Heart, doi:10.1136/openhrt-2020-001350.
90.
Mothae et al., SARS-CoV-2 host-pathogen interactome: insights into more players during pathogenesis, Virology, doi:10.1016/j.virol.2025.110607.
91.
Hazan et al., Treatment with Ivermectin Increases the Population of Bifidobacterium in the Gut, ACG 2023, acg2023posters.eventscribe.net/posterspeakers.asp.
de Oliveira Só et al., 28 Apr 2024, preprint, 8 authors.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
In Silico Comparative Analysis of Ivermectin and Nirmatrelvir Inhibitors Interacting with the SARS-CoV-2 Main Protease
doi:10.20944/preprints202404.1825.v1
Exploring therapeutic options is crucial in the ongoing COVID-19 pandemic caused by SARS-CoV-2. Nirmatrelvir, a potent inhibitor targeting the SARS-CoV-2 M pro , shows promise as an antiviral treatment. Additionally, Ivermectin, a broad-spectrum antiparasitic drug, has demonstrated effectiveness against the virus in laboratory settings. However, its clinical implications are still debated. Using computational methods such as molecular docking and 100 ns molecular dynamics simulations, we investigated how Nirmatrelvir and Ivermectin interact with SARS-CoV-2 M pro(A) . Calculations using density functional theory have been instrumental in elucidating the behavior of isolated molecules, primarily by analyzing the frontier molecular orbitals. Our analysis revealed distinct binding patterns: Nirmatrelvir formed strong interactions with amino acids like MET49, MET165, HIS41, HIS163, HIS164, PHE140, CYS145, GLU166, and ASN142, showing stable binding with a root mean square deviation (RMSD) of around 2.0 Å. On the other hand, Ivermectin interacted with THR237, THR239, LEU271, LEU272, and LEU287, displaying an RMSD of 1.87 Å, indicating enduring interactions. Both ligands stabilized M pro (A) , with Ivermectin showing stability and persistent interactions despite forming fewer hydrogen bonds. These findings offer detailed insights into how Nirmatrelvir and Ivermectin bind to the SARS-CoV-2 main protease, providing valuable information for potential therapeutic strategies against COVID-19.
Institutional Review Board Statement: Not applicable.
Informed Consent Statement: Not applicable.
Conflicts of Interest: The authors declare no conflicts of interest.
References
Agost-Beltrán, De La Hoz-Rodríguez, Bou-Iserte, Rodríguez, Fernández-De-La Pradilla et al., Advances in the development of SARS-CoV-2 Mpro inhibitors, Molecules
Ahmad, Batool, Ain, Kim, Choi, Exploring the binding mechanism of PF-07321332 SARS-CoV-2 protease inhibitor through molecular dynamics and binding free energy simulations, International journal of molecular sciences
Alugubelli, Geng, Yang, Shaabani, Khatua et al., A systematic exploration of boceprevir-based main protease inhibitors as SARS-CoV-2 antivirals, European Journal of Medicinal Chemistry
Andi, Kumaran, Kreitler, Soares, Keereetaweep et al., Hepatitis C virus NS3/4A inhibitors and other drug-like compounds as covalent binders of SARS-CoV-2 main protease, Scientific reports
Asadi, Airborne Infectious Disease Transmission Via Expiratory Aerosol Particles and Aerosolized Fomites
Atzrodt, Maknojia, Mccarthy, Oldfield, Po et al., A Guide to COVID-19: a global pandemic caused by the novel coronavirus SARS-CoV-2, The FEBS journal
Banerjee, Perera, Tillekeratne, Potential SARS-CoV-2 main protease inhibitors, Drug Discovery Today
Becke, A new mixing of Hartree-Fock and local density-functional theories, The Journal of chemical physics
Buonfrate, Chesini, Martini, Roncaglioni, Fernandez et al., High-dose ivermectin for early treatment of COVID-19 (COVER study): a randomised, double-blind, multicentre, phase II, dose-finding, proof-of-concept clinical trial, International journal of antimicrobial agents
Cannalire, Cerchia, Beccari, Di Leva, Summa, Targeting SARS-CoV-2 proteases and polymerase for COVID-19 treatment: state of the art and future opportunities, Journal of medicinal chemistry
Cobos-Campos, Apiñaniz, Parraza, Cordero, García et al., Potential use of ivermectin for the treatment and prophylaxis of SARS-CoV-2 infection, Current Research in Translational Medicine
Di Chio, Previti, Amendola, Ravichandran, Wagner et al., Development of novel dipeptide nitriles as inhibitors of rhodesain of Trypanosoma brucei rhodesiense, European Journal of Medicinal Chemistry
Frisch, Trucks, Schlegel, Scuseria, Robb et al., None, Gaussian
Fukunishi, Yamashita, Mashimo, Nakamura, Prediction of protein-compound binding energies from known activity data: docking-score-based method and its applications, Molecular Informatics
Hill, Garratt, Levi, Falconer, Ellis et al., Retracted: meta-analysis of randomized trials of ivermectin to treat SARS-CoV-2 infection. Open forum infectious diseases
Huey, Morris, Forli, Using AutoDock 4 and AutoDock vina with AutoDockTools: a tutorial
Humphrey, Dalke, Schulten, Vmd, Visual molecular dynamics, Journal of Molecular Graphics, doi:10.1016/0263-7855(96)00018-5
Jayaweera, Perera, Gunawardana, Manatunge, Transmission of COVID-19 virus by droplets and aerosols: A critical review on the unresolved dichotomy, Environmental research
Jin, Du, Xu, Deng, Liu et al., Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors, Nature
Klauda, Venable, Freites, O'connor, Tobias et al., Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types, The journal of physical chemistry B
Lin, Zeng, Duan, Yang, Ma et al., Molecular mechanism of ensitrelvir inhibiting SARS-CoV-2 main protease and its variants, Communications Biology
Louten, Virus transmission and epidemiology, Essential human virology
Miura, Malla, Owen, Tumber, Brewitz et al., In vitro selection of macrocyclic peptide inhibitors containing cyclic γ2, 4-amino acids targeting the SARS-CoV-2 main protease, Nature Chemistry
Parr, Density functional theory, Annual Review of Physical Chemistry
Pellis, Scarabel, Stage, Overton, Chappell et al., Challenges in control of COVID-19: short doubling time and long delay to effect of interventions, Philosophical Transactions of the Royal Society B
Phillips, Zheng, Kumar, Kalé, Namd, Biomolecular simulation on thousands of processors, doi:10.1109/SC.2002.10019
Rajeev, Prathiviraj, Kiran, Selvin, Zoonotic evolution and implications of microbiome in viral transmission and infection, Virus research
Sham, Schlüter, Density-functional theory of the energy gap, Physical review letters
Sharun, Dhama, Patel, Pathak, Tiwari et al., a new candidate therapeutic against SARS-CoV-2/COVID-19
Stewart, Application of the PM6 method to modeling proteins, Journal of molecular modeling
Tyndall, S-217622, a 3CL protease inhibitor and clinical candidate for SARS-CoV-2, Journal of Medicinal Chemistry
Weigend, Ahlrichs, Balanced basis sets of split valence, triple zeta valence and quadruple zeta valence quality for H to Rn: Design and assessment of accuracy, Physical Chemistry Chemical Physics
Weiss, Sankaran, Emergence of epidemic diseases: Zoonoses and other origins, Faculty Reviews
Worldometer, COVID-19 CORONAVIRUS PANDEMIC
Yamamoto, Yasuo, Sekijima, Screening for inhibitors of main protease in sars-cov-2: In silico and in vitro approach avoiding peptidyl secondary amides, Journal of Chemical Information and Modeling
Zaidi, Dehgani-Mobaraki, The mechanisms of action of ivermectin against SARS-CoV-2-an extensive review, The Journal of antibiotics
Zhao, Fang, Zhang, Zhang, Zhao et al., Crystal structure of SARS-CoV-2 main protease in complex with protease inhibitor PF-07321332, Protein & cell
DOI record:
{
"DOI": "10.20944/preprints202404.1825.v1",
"URL": "http://dx.doi.org/10.20944/preprints202404.1825.v1",
"abstract": "<jats:p>Exploring therapeutic options is crucial in the ongoing COVID-19 pandemic caused by SARS-CoV-2. Nirmatrelvir, a potent inhibitor targeting the SARS-CoV-2 Mpro, shows promise as an antiviral treatment. Additionally, Ivermectin, a broad-spectrum antiparasitic drug, has demonstrated effectiveness against the virus in laboratory settings. However, its clinical implications are still debated. Using computational methods such as molecular docking and 100 ns molecular dynamics simulations, we investigated how Nirmatrelvir and Ivermectin interact with SARS-CoV-2 Mpro(A). Calculations using density functional theory have been instrumental in elucidating the behavior of isolated molecules, primarily by analyzing the frontier molecular orbitals. Our analysis revealed distinct binding patterns: Nirmatrelvir formed strong interactions with amino acids like MET49, MET165, HIS41, HIS163, HIS164, PHE140, CYS145, GLU166, and ASN142, showing stable binding with a root mean square deviation (RMSD) of around 2.0 Å. On the other hand, Ivermectin interacted with THR237, THR239, LEU271, LEU272, and LEU287, displaying an RMSD of 1.87 Å, indicating enduring interactions. Both ligands stabilized Mpro(A), with Ivermectin showing stability and persistent interactions despite forming fewer hydrogen bonds. These findings offer detailed insights into how Nirmatrelvir and Ivermectin bind to the SARS-CoV-2 main protease, providing valuable information for potential therapeutic strategies against COVID-19.</jats:p>",
"accepted": {
"date-parts": [
[
2024,
4,
26
]
]
},
"author": [
{
"affiliation": [],
"family": "de Oliveira Só",
"given": "Yuri Alves",
"sequence": "first"
},
{
"affiliation": [],
"family": "Bezerra",
"given": "Kelton Silva",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0003-3823-7436",
"affiliation": [],
"authenticated-orcid": false,
"family": "Gargano",
"given": "Ricardo",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Mendonça",
"given": "Fabio L.L",
"sequence": "additional"
},
{
"affiliation": [],
"family": "de Souto",
"given": "Janeusa Trindade",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Fulco",
"given": "Umberto L.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Pereira Junior",
"given": "Marcelo Lopes",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Júnior",
"given": "Luiz Antônio Ribeiro",
"sequence": "additional"
}
],
"container-title": [],
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2024,
4,
29
]
],
"date-time": "2024-04-29T08:42:45Z",
"timestamp": 1714380165000
},
"deposited": {
"date-parts": [
[
2024,
4,
29
]
],
"date-time": "2024-04-29T08:48:59Z",
"timestamp": 1714380539000
},
"group-title": "Biology and Life Sciences",
"indexed": {
"date-parts": [
[
2024,
4,
30
]
],
"date-time": "2024-04-30T00:29:13Z",
"timestamp": 1714436953418
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2024,
4,
28
]
]
},
"license": [
{
"URL": "http://creativecommons.org/licenses/by/4.0",
"content-version": "unspecified",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
4,
28
]
],
"date-time": "2024-04-28T00:00:00Z",
"timestamp": 1714262400000
}
}
],
"member": "1968",
"original-title": [],
"posted": {
"date-parts": [
[
2024,
4,
28
]
]
},
"prefix": "10.20944",
"published": {
"date-parts": [
[
2024,
4,
28
]
]
},
"publisher": "MDPI AG",
"reference-count": 0,
"references-count": 0,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.preprints.org/manuscript/202404.1825/v1"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"subtype": "preprint",
"title": "In Silico Comparative Analysis of Ivermectin and Nirmatrelvir Inhibitors Interacting with the SARS-CoV-2 Main Protease",
"type": "posted-content"
}