Identification of polyphenols as novel neuropilin-1 cendR pocket inhibitors to block SARS-CoV-2 entry and enhance variant resistance
et al., PLOS One, doi:10.1371/journal.pone.0345051, Mar 2026
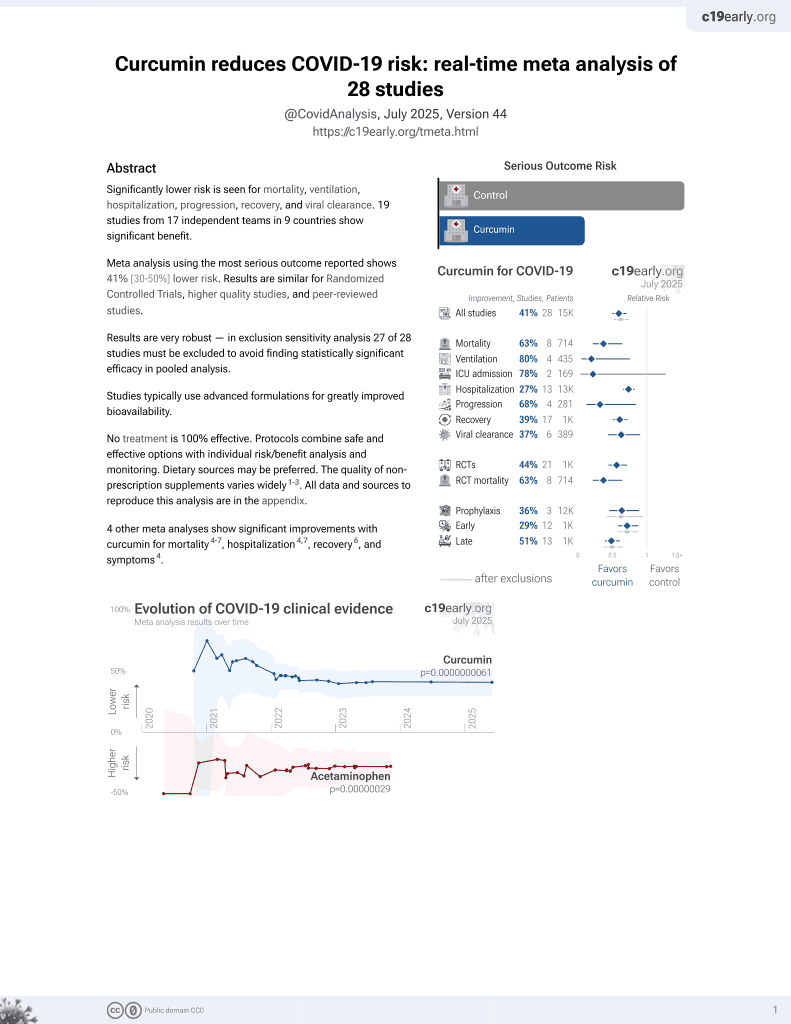
Curcumin for COVID-19
16th treatment shown to reduce risk in
February 2021, now with p = 0.0000000061 from 28 studies.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
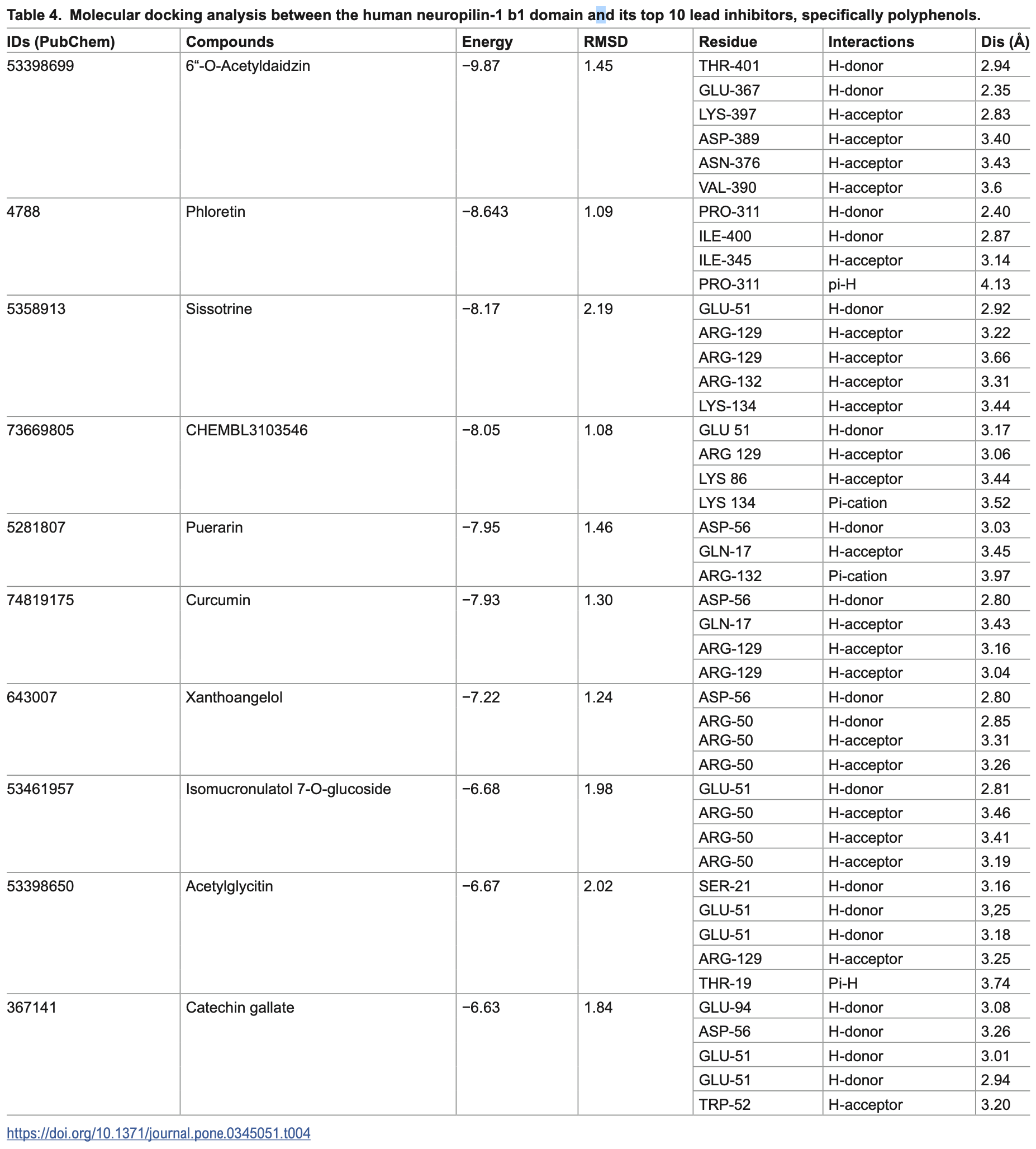
In silico study showing that polyphenols, particularly 6"-O-acetyldaidzin and phloretin, bind to the neuropilin-1 (NRP-1) b1 domain CendR pocket with higher affinity than the synthetic inhibitor EG01377, potentially blocking SARS-CoV-2 spike protein engagement. Curcumin showed a binding affinity of -7.93 kcal/mol, forming hydrogen bonds with ASP-56, GLN-17, and ARG-129, suggesting it may interfere with NRP-1-mediated SARS-CoV-2 entry.
64 preclinical studies support the efficacy of curcumin for COVID-19:
In silico studies predict inhibition of SARS-CoV-2 with curcumin or metabolites via binding to the spikeA,3,7,8,13,18,20,26,29 (and specifically the receptor binding domainB,2,4,6,16,19,22 ), MproC,6-8,13,15,17-19,21,22,24,27,29,30,32,50 , RNA-dependent RNA polymeraseD,6-8,19,28 , PLproE,8, ACE2F,4,20,21,23 , nucleocapsidG,14,31 , nsp10H,31, and helicaseI,38 proteins, and inhibition of spike-ACE2 interactionJ,2,5 .
In vitro studies demonstrate inhibition of the spikeA,43 (and specifically the receptor binding domainB,53), MproC,25,43,50,52 , ACE2F,53, and TMPRSS2K,53 proteins, and inhibition of spike-ACE2 interactionJ,5,36 .
In vitro studies demonstrate efficacy in Calu-3L,51, A549M,43, A549-ATN,33, 293TO,9, HEK293-hACE2P,25,41 , 293T/hACE2/TMPRSS2Q,42, Vero E6R,3,15,19,29,41,43,45,47,49,51 , and SH-SY5YS,40 cells.
Curcumin decreases pro-inflammatory cytokines induced by SARS-CoV-2 in peripheral blood mononuclear cells49, alleviates SARS-CoV-2 spike protein-induced mitochondrial membrane damage and oxidative stress9, may limit COVID-19 induced cardiac damage by inhibiting the NF-κB signaling pathway which mediates the profibrotic effects of the SARS-CoV-2 spike protein on cardiac fibroblasts37, is predicted to inhibit the interaction between the SARS-CoV-2 spike protein receptor binding domain and the human ACE2 receptor for the delta and omicron variants16, lowers ACE2 and STAT3, curbing lung inflammation and ARDS in preclinical COVID-19 models34, inhibits SARS-CoV-2 ORF3a ion channel activity, which contributes to viral pathogenicity and cytotoxicity44, has direct virucidal action by disrupting viral envelope integrity46, may inhibit viral replication and modulate inflammatory pathways like NF-κB via SIRT1 activation54, and can function as a photosensitizer in photodynamic therapy to generate reactive oxygen species that damage the virus46.
1.
Ataya et al., Identification of polyphenols as novel neuropilin-1 cendR pocket inhibitors to block SARS-CoV-2 entry and enhance variant resistance, PLOS One, doi:10.1371/journal.pone.0345051.
2.
Jena et al., Halved but Potent: Exploring the Inhibitory Property of Curcumin Derivatives Against Evolving SARS-CoV-2 Strains, The Protein Journal, doi:10.1007/s10930-025-10309-1.
3.
Marzouk et al., Computational and Experimental Insights into the Antiviral Mechanism of Turmeric (Curcuma longa) against SARS-CoV-2 D614G, BIO Web of Conferences, doi:10.1051/bioconf/202519804002.
4.
Wu et al., Utilizing natural compounds as ligands to disrupt the binding of SARS-CoV-2 receptor-binding domain to angiotensin-converting enzyme 2, impeding viral infection, Phytochemistry Letters, doi:10.1016/j.phytol.2025.102999.
5.
Najimi et al., Phytochemical Inhibitors of SARS‐CoV‐2 Entry: Targeting the ACE2‐RBD Interaction with l‐Tartaric Acid, l‐Ascorbic Acid, and Curcuma longa Extract, ChemistrySelect, doi:10.1002/slct.202406035.
6.
Rajamanickam et al., Exploring the Potential of Siddha Formulation MilagaiKudineer-Derived Phytotherapeutics Against SARS-CoV-2: An In-Silico Investigation for Antiviral Intervention, Journal of Pharmacy and Pharmacology Research, doi:10.26502/fjppr.0105.
7.
Al balawi et al., Assessing multi-target antiviral and antioxidant activities of natural compounds against SARS-CoV-2: an integrated in vitro and in silico study, Bioresources and Bioprocessing, doi:10.1186/s40643-024-00822-z.
8.
Haque et al., Exploring potential therapeutic candidates against COVID-19: a molecular docking study, Discover Molecules, doi:10.1007/s44345-024-00005-5.
9.
Zhang et al., Computational Discovery of Mitochondrial Dysfunction Biomarkers in Severe SARS-CoV-2 Infection: Facilitating Pytomedicine Screening, Phytomedicine, doi:10.1016/j.phymed.2024.155784.
10.
Öztürkkan et al., In Silico investigation of the effects of curcuminoids on the spike protein of the omicron variant of SARS-CoV-2, Baku State University Journal of Chemistry and Material Sciences, 1:2, bsuj.bsu.edu.az/uploads/pdf/ec4204d62f7802de54e6092bf7860029.pdf.
11.
Yunze et al., Therapeutic effect and potential mechanism of curcumin, an active ingredient in Tongnao Decoction, on COVID-19 combined with stroke: a network pharmacology study and GEO database mining, Research Square, doi:10.21203/rs.3.rs-4329762/v1.
12.
Agamah et al., Network-based multi-omics-disease-drug associations reveal drug repurposing candidates for COVID-19 disease phases, ScienceOpen, doi:10.58647/DRUGARXIV.PR000010.v1.
13.
Boseila et al., Throat spray formulated with virucidal Pharmaceutical excipients as an effective early prophylactic or treatment strategy against pharyngitis post-exposure to SARS CoV-2, European Journal of Pharmaceutics and Biopharmaceutics, doi:10.1016/j.ejpb.2024.114279.
14.
Hidayah et al., Bioinformatics study of curcumin, demethoxycurcumin, bisdemethoxycurcumin and cyclocurcumin compounds in Curcuma longa as an antiviral agent via nucleocapsid on SARS-CoV-2 inhibition, International Conference on Organic and Applied Chemistry, doi:10.1063/5.0197724.
15.
Singh et al., Unlocking the potential of phytochemicals in inhibiting SARS-CoV-2 M Pro protein - An in-silico and cell-based approach, Research Square, doi:10.21203/rs.3.rs-3888947/v1.
16.
Kant et al., Structure-based drug discovery to identify SARS-CoV2 spike protein–ACE2 interaction inhibitors, Journal of Biomolecular Structure and Dynamics, doi:10.1080/07391102.2023.2300060.
17.
Naderi Beni et al., In silico studies of anti-oxidative and hot temperament-based phytochemicals as natural inhibitors of SARS-CoV-2 Mpro, PLOS ONE, doi:10.1371/journal.pone.0295014.
18.
Moschovou et al., Exploring the Binding Effects of Natural Products and Antihypertensive Drugs on SARS-CoV-2: An In Silico Investigation of Main Protease and Spike Protein, International Journal of Molecular Sciences, doi:10.3390/ijms242115894.
19.
Eleraky et al., Curcumin Transferosome-Loaded Thermosensitive Intranasal in situ Gel as Prospective Antiviral Therapy for SARS-Cov-2, International Journal of Nanomedicine, doi:10.2147/IJN.S423251.
20.
Singh (B) et al., Computational studies to analyze effect of curcumin inhibition on coronavirus D614G mutated spike protein, The Seybold Report, doi:10.17605/OSF.IO/TKEXJ.
21.
Thapa et al., In-silico Approach for Predicting the Inhibitory Effect of Home Remedies on Severe Acute Respiratory Syndrome Coronavirus-2, Makara Journal of Science, doi:10.7454/mss.v27i3.1609.
22.
Srivastava et al., Paradigm of Well-Orchestrated Pharmacokinetic Properties of Curcuminoids Relative to Conventional Drugs for the Inactivation of SARS-CoV-2 Receptors: An In Silico Approach, Stresses, doi:10.3390/stresses3030043.
23.
Alkafaas et al., A study on the effect of natural products against the transmission of B.1.1.529 Omicron, Virology Journal, doi:10.1186/s12985-023-02160-6.
24.
Winih Kinasih et al., Analisis in silico interaksi senyawa kurkuminoid terhadap enzim main protease 6LU7 dari SARS-CoV-2, Duta Pharma Journal, doi:10.47701/djp.v3i1.2904.
25.
Wu (B) et al., Potential Mechanism of Curcumin and Resveratrol against SARS-CoV-2, Research Square, doi:10.21203/rs.3.rs-2780614/v1.
26.
Nag et al., Curcumin inhibits spike protein of new SARS-CoV-2 variant of concern (VOC) Omicron, an in silico study, Computers in Biology and Medicine, doi:10.1016/j.compbiomed.2022.105552.
27.
Rampogu et al., Molecular Docking and Molecular Dynamics Simulations Discover Curcumin Analogue as a Plausible Dual Inhibitor for SARS-CoV-2, International Journal of Molecular Sciences, doi:10.3390/ijms23031771.
28.
Singh (C) et al., Potential of turmeric-derived compounds against RNA-dependent RNA polymerase of SARS-CoV-2: An in-silico approach, Computers in Biology and Medicine, doi:10.1016/j.compbiomed.2021.104965.
29.
Kandeil et al., Bioactive Polyphenolic Compounds Showing Strong Antiviral Activities against Severe Acute Respiratory Syndrome Coronavirus 2, Pathogens, doi:10.3390/pathogens10060758.
30.
Rajagopal et al., Activity of phytochemical constituents of Curcuma longa (turmeric) and Andrographis paniculata against coronavirus (COVID-19): an in silico approach, Future Journal of Pharmaceutical Sciences, doi:10.1186/s43094-020-00126-x.
31.
Suravajhala et al., Comparative Docking Studies on Curcumin with COVID-19 Proteins, Preprints, doi:10.20944/preprints202005.0439.v3.
32.
Sekiou et al., In-Silico Identification of Potent Inhibitors of COVID-19 Main Protease (Mpro) and Angiotensin Converting Enzyme 2 (ACE2) from Natural Products: Quercetin, Hispidulin, and Cirsimaritin Exhibited Better Potential Inhibition than Hydroxy-Chloroquine Against COVID-19 Main Protease Active Site and ACE2, ChemRxiv, doi:10.26434/chemrxiv.12181404.v1.
33.
Grüneberg et al., Dose-dependent antiviral effects of glycyrrhizin, curcumin, and harmaline against clinical SARS-CoV-2 isolates, including D614G, Omicron BA.5, and Omicron XBB.1, BMC Complementary Medicine and Therapies, doi:10.1186/s12906-026-05253-1.
34.
Aktay et al., Oral Administration of Water-Soluble Curcumin Complex Prevents ARDS With the Potential for COVID-19 Treatment, Phytotherapy Research, doi:10.1002/ptr.70046.
35.
Olubiyi et al., Novel dietary herbal preparations with inhibitory activities against multiple SARS-CoV-2 targets: A multidisciplinary investigation into antiviral activities, Food Chemistry Advances, doi:10.1016/j.focha.2025.100969.
36.
Emam et al., Establishment of in-house assay for screening of anti-SARS-CoV-2 protein inhibitors, AMB Express, doi:10.1186/s13568-024-01739-8.
37.
Van Tin et al., Spike Protein of SARS-CoV-2 Activates Cardiac Fibrogenesis through NLRP3 Inflammasomes and NF-κB Signaling, Cells, doi:10.3390/cells13161331.
38.
Li et al., Thermal shift assay (TSA)-based drug screening strategy for rapid discovery of inhibitors against the Nsp13 helicase of SARS-CoV-2, Animals and Zoonoses, doi:10.1016/j.azn.2024.06.001.
39.
Kamble et al., Nanoparticulate curcumin spray imparts prophylactic and therapeutic properties against SARS-CoV-2, Emergent Materials, doi:10.1007/s42247-024-00754-6.
40.
Nicoliche et al., Antiviral, anti-inflammatory and antioxidant effects of curcumin and curcuminoids in SH-SY5Y cells infected by SARS-CoV-2, Scientific Reports, doi:10.1038/s41598-024-61662-7.
41.
Nittayananta et al., A novel film spray containing curcumin inhibits SARS-CoV-2 and influenza virus infection and enhances mucosal immunity, Virology Journal, doi:10.1186/s12985-023-02282-x.
42.
Septisetyani et al., Curcumin and turmeric extract inhibited SARS-CoV-2 pseudovirus cell entry and Spike mediated cell fusion, bioRxiv, doi:10.1101/2023.09.28.560070.
43.
Mohd Abd Razak et al., In Vitro Anti-SARS-CoV-2 Activities of Curcumin and Selected Phenolic Compounds, Natural Product Communications, doi:10.1177/1934578X231188861.
44.
Fam et al., Channel activity of SARS-CoV-2 viroporin ORF3a inhibited by adamantanes and phenolic plant metabolites, Scientific Reports, doi:10.1038/s41598-023-31764-9.
45.
Teshima et al., Antiviral activity of curcumin and its analogs selected by an artificial intelligence-supported activity prediction system in SARS-CoV-2-infected VeroE6 cells, Natural Product Research, doi:10.1080/14786419.2023.2194647.
46.
Zupin et al., Optimization of Anti-SARS-CoV-2 Treatments Based on Curcumin, Used Alone or Employed as a Photosensitizer, Viruses, doi:10.3390/v14102132.
47.
Leka et al., In vitro antiviral activity against SARS-CoV-2 of common herbal medicinal extracts and their bioactive compounds, Phytotherapy Research, doi:10.1002/ptr.7463.
48.
Goc et al., Inhibitory effects of specific combination of natural compounds against SARS-CoV-2 and its Alpha, Beta, Gamma, Delta, Kappa, and Mu variants, European Journal of Microbiology and Immunology, doi:10.1556/1886.2021.00022.
49.
Marín-Palma et al., Curcumin Inhibits In Vitro SARS-CoV-2 Infection In Vero E6 Cells through Multiple Antiviral Mechanisms, Molecules, doi:10.3390/molecules26226900.
50.
Bahun et al., Inhibition of the SARS-CoV-2 3CLpro main protease by plant polyphenols, Food Chemistry, doi:10.1016/j.foodchem.2021.131594.
51.
Bormann et al., Turmeric Root and Its Bioactive Ingredient Curcumin Effectively Neutralize SARS-CoV-2 In Vitro, Viruses, doi:10.3390/v13101914.
52.
Guijarro-Real et al., Potential In Vitro Inhibition of Selected Plant Extracts against SARS-CoV-2 Chymotripsin-Like Protease (3CLPro) Activity, Foods, doi:10.3390/foods10071503.
a.
The trimeric spike (S) protein is a glycoprotein that mediates viral entry by binding to the host ACE2 receptor, is critical for SARS-CoV-2's ability to infect host cells, and is a target of neutralizing antibodies. Inhibition of the spike protein prevents viral attachment, halting infection at the earliest stage.
b.
The receptor binding domain is a specific region of the spike protein that binds ACE2 and is a major target of neutralizing antibodies. Focusing on the precise binding site allows highly specific disruption of viral attachment with reduced potential for off-target effects.
c.
The main protease or Mpro, also known as 3CLpro or nsp5, is a cysteine protease that cleaves viral polyproteins into functional units needed for replication. Inhibiting Mpro disrupts the SARS-CoV-2 lifecycle within the host cell, preventing the creation of new copies.
d.
RNA-dependent RNA polymerase (RdRp), also called nsp12, is the core enzyme of the viral replicase-transcriptase complex that copies the positive-sense viral RNA genome into negative-sense templates for progeny RNA synthesis. Inhibiting RdRp blocks viral genome replication and transcription.
e.
The papain-like protease (PLpro) has multiple functions including cleaving viral polyproteins and suppressing the host immune response by deubiquitination and deISGylation of host proteins. Inhibiting PLpro may block viral replication and help restore normal immune responses.
f.
The angiotensin converting enzyme 2 (ACE2) protein is a host cell transmembrane protein that serves as the cellular receptor for the SARS-CoV-2 spike protein. ACE2 is expressed on many cell types, including epithelial cells in the lungs, and allows the virus to enter and infect host cells. Inhibition may affect ACE2's physiological function in blood pressure control.
g.
The nucleocapsid (N) protein binds and encapsulates the viral genome by coating the viral RNA. N enables formation and release of infectious virions and plays additional roles in viral replication and pathogenesis. N is also an immunodominant antigen used in diagnostic assays.
h.
Non-structural protein 10 (nsp10) serves as an RNA chaperone and stabilizes conformations of nsp12 and nsp14 in the replicase-transcriptase complex, which synthesizes new viral RNAs. Nsp10 disruption may destabilize replicase-transcriptase complex activity.
i.
The helicase, or nsp13, protein unwinds the double-stranded viral RNA, a crucial step in replication and transcription. Inhibition may prevent viral genome replication and the creation of new virus components.
j.
The interaction between the SARS-CoV-2 spike protein and the human ACE2 receptor is a primary method of viral entry, inhibiting this interaction can prevent the virus from attaching to and entering host cells, halting infection at an early stage.
k.
Transmembrane protease serine 2 (TMPRSS2) is a host cell protease that primes the spike protein, facilitating cellular entry. TMPRSS2 activity helps enable cleavage of the spike protein required for membrane fusion and virus entry. Inhibition may especially protect respiratory epithelial cells, buy may have physiological effects.
l.
Calu-3 is a human lung adenocarcinoma cell line with moderate ACE2 and TMPRSS2 expression and SARS-CoV-2 susceptibility. It provides a model of the human respiratory epithelium, but many not be ideal for modeling early stages of infection due to the moderate expression levels of ACE2 and TMPRSS2.
m.
A549 is a human lung carcinoma cell line with low ACE2 expression and SARS-CoV-2 susceptibility. Viral entry/replication can be studied but the cells may not replicate all aspects of lung infection.
n.
A549-AT is a human lung carcinoma cell line stably transfected with ACE2 and TMPRSS2 receptors. Unlike the parental line, this overexpression ensures stable infection and enhanced viral entry, allowing for the evaluation of antiviral efficacy against various SARS-CoV-2 variants.
o.
293T is a human embryonic kidney cell line that can be engineered for high ACE2 expression and SARS-CoV-2 susceptibility. 293T cells are easily transfected and support high protein expression.
p.
HEK293-hACE2 is a human embryonic kidney cell line with high ACE2 expression and SARS-CoV-2 susceptibility. Cells have been transfected with a plasmid to express the human ACE2 (hACE2) protein.
q.
293T/hACE2/TMPRSS2 is a human embryonic kidney cell line engineered for high ACE2 and TMPRSS2 expression, which mimics key aspects of human infection. 293T/hACE2/TMPRSS2 cells are very susceptible to SARS-CoV-2 infection.
r.
Vero E6 is an African green monkey kidney cell line with low/no ACE2 expression and high SARS-CoV-2 susceptibility. The cell line is easy to maintain and supports robust viral replication, however the monkey origin may not accurately represent human responses.
s.
SH-SY5Y is a human neuroblastoma cell line that exhibits neuronal phenotypes. It is commonly used as an in vitro model for studying neurotoxicity, neurodegenerative diseases, and neuronal differentiation.
Ataya et al., 23 Mar 2026, peer-reviewed, 4 authors.
Contact: fataya@ksu.edu.sa.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
Abstract:
Citation: Ataya F , Alamro A, Alghamdi A, Fouad D (2026) Identification of polyphenols as novel neuropilin-1 cendR pocket inhibitors to block SARS-CoV-2 entry and enhance variant resistance. PLoS One 21(3): e0345051. https:// doi.org/10.1371/journal.pone.0345051
Editor: Yusuf Oloruntoyin Ayipo, Kwara State University, NIGERIA
Received:
November 10, 2025
Accepted:
February 12, 2026
Published:
March 23, 2026
Copyright: © 2026 Ataya et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Data availability statement : All relevant data are included within the paper and its supplementary file. No additional datasets were generated or deposited in external repositories.
Funding: FA received funding from King Abdulaziz City for Science and Technology (KACST) - Projects, grant number
RESEARCH ARTICLE
Identification of polyphenols as novel neuropilin-1 cendR pocket inhibitors to block SARS-CoV-2 entry and enhance variant resistance
Farid Ataya 1 * , Abir Alamro 1 , Amani Alghamdi 1 , Dalia Fouad 2
- 1 Department of Biochemistry, College of Science, King Saud University, Riyadh, Saudi Arabia,
- 2 Department of Zoology, College of Science, King Saud University, Riyadh, Saudi Arabia
[* fataya@ksu.edu.sa](mailto:fataya@ksu.edu.sa)
Abstract
Neuropilin-1 (NRP-1) functions as an essential co-receptor for SARS-CoV-2, facilitating viral entry by binding the spike protein's C-end rule (CendR) motif in its b1 domain, yet it has received less attention than ACE2 in therapeutic development. This in silico study evaluates plant-derived polyphenols as potential selective inhibitors of the NRP-1 CendR pocket to disrupt SARS-CoV-2 engagement, addressing limitations of synthetic inhibitors like EG01377, which exhibit modest affinity (-5.83 kcal/mol) and potential off-target risks. High-throughput molecular docking of 10,000 phytochemicals using AutoDock Vina identified 10 polyphenols with binding affinities ranging from -9.87 to -6.63 kcal/mol, led by 6'-O-acetyldaidzin (-9.87 kcal/mol) and phloretin (-8.64 kcal/mol), forming stable hydrogen bonds and π-cation interactions with critical residues (e.g., THR-401, GLU-367 for 6'-O-acetyldaidzin; PRO-311, ILE400 for phloretin), as visualized in Discovery Studio. Notably, four of the top inhibitors are isoflavonoid derivatives, highlighting a chemical class enrichment. Molecular dynamics simulations over 100 ns using Desmond indicated moderate complex stability (RMSD: 0.6-3.8 Å; RMSF <0.5 Å at binding site ). ADMET-Tox profiling via SwissADME and ProTox-II revealed drug-like properties, including high gastrointestinal absorption (>70% for leads) and low toxicity (classes 4-5), though 6'-Oacetyldaidzin shows limited bioavailability due to its high H-bond acceptor count (10) and large size, suggesting need for formulation optimization. The NRP-1 b1 homology model, built with SWISS-MODEL, exhibited high fidelity (GMQE: 0.79; Ramachandran favored regions: 90.3% ). This focused computational screening of polyphenols against NRP-1 complements prior studies and identifies candidates for experimental validation as potential SARS-CoV-2 inhibitors. Limitations include the in silico nature, and lack of membrane/sialic acid models, necessitating in vitro and in vivo testing against SARS-CoV-2..
DOI record:
{
"DOI": "10.1371/journal.pone.0345051",
"ISSN": [
"1932-6203"
],
"URL": "http://dx.doi.org/10.1371/journal.pone.0345051",
"abstract": "<jats:p>\n Neuropilin-1 (NRP-1) functions as an essential co-receptor for SARS-CoV-2, facilitating viral entry by binding the spike protein’s C-end rule (CendR) motif in its b1 domain, yet it has received less attention than ACE2 in therapeutic development. This\n <jats:italic>in silico</jats:italic>\n study evaluates plant-derived polyphenols as potential selective inhibitors of the NRP-1 CendR pocket to disrupt SARS-CoV-2 engagement, addressing limitations of synthetic inhibitors like EG01377, which exhibit modest affinity (−5.83 kcal/mol) and potential off-target risks. High-throughput molecular docking of 10,000 phytochemicals using AutoDock Vina identified 10 polyphenols with binding affinities ranging from −9.87 to −6.63 kcal/mol, led by 6“-O-acetyldaidzin (-9.87 kcal/mol) and phloretin (-8.64 kcal/mol), forming stable hydrogen bonds and π-cation interactions with critical residues (e.g., THR-401, GLU-367 for 6”-O-acetyldaidzin; PRO-311, ILE-400 for phloretin), as visualized in Discovery Studio. Notably, four of the top inhibitors are isoflavonoid derivatives, highlighting a chemical class enrichment. Molecular dynamics simulations over 100 ns using Desmond indicated moderate complex stability (RMSD: 0.6–3.8 Å; RMSF <0.5 Å at binding site\n <jats:bold>).</jats:bold>\n ADMET-Tox profiling via SwissADME and ProTox-II revealed drug-like properties, including high gastrointestinal absorption (>70% for leads) and low toxicity (classes 4–5), though 6”-O-acetyldaidzin shows limited bioavailability due to its high H-bond acceptor count (10) and large size, suggesting need for formulation optimization. The NRP-1 b1 homology model, built with SWISS-MODEL, exhibited high fidelity (GMQE: 0.79; Ramachandran favored regions: 90.3%\n <jats:bold>).</jats:bold>\n This focused computational screening of polyphenols against NRP-1 complements prior studies and identifies candidates for experimental validation as potential SARS-CoV-2 inhibitors. Limitations include the\n <jats:italic>in silico</jats:italic>\n nature, and lack of membrane/sialic acid models, necessitating\n <jats:italic>in vitro</jats:italic>\n and\n <jats:italic>in vivo</jats:italic>\n testing against SARS-CoV-2 variants.\n </jats:p>",
"author": [
{
"ORCID": "https://orcid.org/0009-0006-7036-3888",
"affiliation": [],
"authenticated-orcid": true,
"family": "Ataya",
"given": "Farid",
"sequence": "first"
},
{
"affiliation": [],
"family": "Alamro",
"given": "Abir",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Alghamdi",
"given": "Amani",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Fouad",
"given": "Dalia",
"sequence": "additional"
}
],
"container-title": "PLOS One",
"container-title-short": "PLoS One",
"content-domain": {
"crossmark-restriction": false,
"domain": [
"www.plosone.org"
]
},
"created": {
"date-parts": [
[
2026,
3,
23
]
],
"date-time": "2026-03-23T17:23:26Z",
"timestamp": 1774286606000
},
"deposited": {
"date-parts": [
[
2026,
3,
23
]
],
"date-time": "2026-03-23T17:23:34Z",
"timestamp": 1774286614000
},
"editor": [
{
"affiliation": [],
"family": "Ayipo",
"given": "Yusuf Oloruntoyin",
"sequence": "first"
}
],
"funder": [
{
"DOI": "10.13039/501100004919",
"award": [
"5-21-01-001-0003"
],
"award-info": [
{
"award-number": [
"5-21-01-001-0003"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100004919",
"id-type": "DOI"
}
],
"name": "King Abdulaziz City for Science and Technology"
}
],
"indexed": {
"date-parts": [
[
2026,
3,
23
]
],
"date-time": "2026-03-23T18:21:14Z",
"timestamp": 1774290074519,
"version": "3.50.1"
},
"is-referenced-by-count": 0,
"issue": "3",
"issued": {
"date-parts": [
[
2026,
3,
23
]
]
},
"journal-issue": {
"issue": "3",
"published-online": {
"date-parts": [
[
2026,
3,
23
]
]
}
},
"language": "en",
"license": [
{
"URL": "http://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
3,
23
]
],
"date-time": "2026-03-23T00:00:00Z",
"timestamp": 1774224000000
}
}
],
"link": [
{
"URL": "https://dx.plos.org/10.1371/journal.pone.0345051",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "340",
"original-title": [],
"page": "e0345051",
"prefix": "10.1371",
"published": {
"date-parts": [
[
2026,
3,
23
]
]
},
"published-online": {
"date-parts": [
[
2026,
3,
23
]
]
},
"publisher": "Public Library of Science (PLoS)",
"reference": [
{
"article-title": "COVID-19 epidemiological update – September 2025",
"author": "World Health Organization",
"journal-title": "WHO",
"key": "pone.0345051.ref001",
"year": "2025"
},
{
"DOI": "10.1126/science.abb2507",
"article-title": "Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation",
"author": "D Wrapp",
"doi-asserted-by": "crossref",
"first-page": "1260",
"issue": "6483",
"journal-title": "Science",
"key": "pone.0345051.ref002",
"volume": "367",
"year": "2020"
},
{
"DOI": "10.1007/s00335-021-09880-6",
"article-title": "SNPs at 3’UTR of APOL1 and miR-6741-3p target sites associated with kidney diseases more susceptible to SARS-COV-2 infection: in silco and in vitro studies",
"author": "M Safdar",
"doi-asserted-by": "crossref",
"first-page": "389",
"issue": "5",
"journal-title": "Mamm Genome",
"key": "pone.0345051.ref003",
"volume": "32",
"year": "2021"
},
{
"DOI": "10.1126/science.abd3072",
"article-title": "Neuropilin-1 is a host factor for SARS-CoV-2 infection",
"author": "JL Daly",
"doi-asserted-by": "crossref",
"first-page": "861",
"issue": "6518",
"journal-title": "Science",
"key": "pone.0345051.ref004",
"volume": "370",
"year": "2020"
},
{
"DOI": "10.1126/science.abd2985",
"article-title": "Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity",
"author": "L Cantuti-Castelvetri",
"doi-asserted-by": "crossref",
"first-page": "856",
"issue": "6518",
"journal-title": "Science",
"key": "pone.0345051.ref005",
"volume": "370",
"year": "2020"
},
{
"article-title": "ACE2 Expression Patterns Across Mammals and Key Findings for SARS-CoV-2 Model Development for Human and Animal Research",
"author": "A Ali",
"first-page": "312",
"issue": "1",
"journal-title": "Pak Vet J",
"key": "pone.0345051.ref006",
"volume": "45",
"year": "2025"
},
{
"article-title": "In-silico natural product database mining for novel neuropilin-1 inhibitors: molecular docking, molecular dynamics and binding energy computations",
"author": "MAA Ibrahim",
"issue": "1",
"journal-title": "Journal of Taibah University for Science",
"key": "pone.0345051.ref007",
"volume": "17",
"year": "2023"
},
{
"article-title": "Neuropilin 1: A Novel Entry Factor for SARS-CoV-2 Infection and a Potential Therapeutic Target",
"author": "E Chekol Abebe",
"first-page": "143",
"journal-title": "Biologics",
"key": "pone.0345051.ref008",
"volume": "15",
"year": "2021"
},
{
"DOI": "10.1016/j.drudis.2018.10.004",
"article-title": "Targeting VEGF-neuropilin interactions: a promising antitumor strategy",
"author": "K Peng",
"doi-asserted-by": "crossref",
"first-page": "656",
"issue": "2",
"journal-title": "Drug Discov Today",
"key": "pone.0345051.ref009",
"volume": "24",
"year": "2019"
},
{
"DOI": "10.3390/nu17142325",
"article-title": "Polyphenols as antiviral agents: their potential against a range of virus types",
"author": "N Coşkun",
"doi-asserted-by": "crossref",
"first-page": "2325",
"issue": "14",
"journal-title": "Nutrients",
"key": "pone.0345051.ref010",
"volume": "17",
"year": "2025"
},
{
"DOI": "10.1002/ptr.8123",
"article-title": "Flavonoids derived from medicinal plants as a COVID-19 treatment",
"author": "M Sopjani",
"doi-asserted-by": "crossref",
"first-page": "1589",
"issue": "3",
"journal-title": "Phytother Res",
"key": "pone.0345051.ref011",
"volume": "38",
"year": "2024"
},
{
"DOI": "10.1016/j.phrs.2020.105224",
"article-title": "Natural product derived phytochemicals in managing acute lung injury by multiple mechanisms",
"author": "Y-Q He",
"doi-asserted-by": "crossref",
"first-page": "105224",
"journal-title": "Pharmacol Res",
"key": "pone.0345051.ref012",
"volume": "163",
"year": "2021"
},
{
"DOI": "10.3390/futurepharmacol2040034",
"article-title": "Computational Screening of Plant-Derived Natural Products against SARS-CoV-2 Variants",
"author": "WA Ansari",
"doi-asserted-by": "crossref",
"first-page": "558",
"issue": "4",
"journal-title": "Future Pharmacology",
"key": "pone.0345051.ref013",
"volume": "2",
"year": "2022"
},
{
"DOI": "10.4103/2225-4110.124335",
"article-title": "Antiviral natural products and herbal medicines",
"author": "L-T Lin",
"doi-asserted-by": "crossref",
"first-page": "24",
"issue": "1",
"journal-title": "J Tradit Complement Med",
"key": "pone.0345051.ref014",
"volume": "4",
"year": "2014"
},
{
"DOI": "10.1186/s13099-023-00535-2",
"article-title": "In-silico computational approaches to study microbiota impacts on diseases and pharmacotherapy",
"author": "H Shokri Garjan",
"doi-asserted-by": "crossref",
"first-page": "10",
"issue": "1",
"journal-title": "Gut Pathog",
"key": "pone.0345051.ref015",
"volume": "15",
"year": "2023"
},
{
"article-title": "Dose-dependent effects and safety assessment of inactivated covid-19 vaccine: a comprehensive preclinical study",
"author": "K Sonmez",
"first-page": "605",
"issue": "2",
"journal-title": "Pak Vet J",
"key": "pone.0345051.ref016",
"volume": "45",
"year": "2025"
},
{
"DOI": "10.1093/jas/skae248",
"article-title": "In silico analysis of polyphenols modulate bovine PPARγ to increase milk fat synthesis in dairy cattle via the MAPK signaling pathways",
"author": "M Safdar",
"doi-asserted-by": "crossref",
"journal-title": "J Anim Sci",
"key": "pone.0345051.ref017",
"volume": "102",
"year": "2024"
},
{
"DOI": "10.1038/srep42717",
"article-title": "SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules",
"author": "A Daina",
"doi-asserted-by": "crossref",
"first-page": "42717",
"journal-title": "Sci Rep",
"key": "pone.0345051.ref018",
"volume": "7",
"year": "2017"
},
{
"DOI": "10.1093/nar/gkl842",
"article-title": "NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins",
"author": "KD Pruitt",
"doi-asserted-by": "crossref",
"journal-title": "Nucleic Acids Res",
"key": "pone.0345051.ref019",
"volume": "35",
"year": "2007"
},
{
"DOI": "10.1093/nar/gky427",
"article-title": "SWISS-MODEL: homology modelling of protein structures and complexes",
"author": "A Waterhouse",
"doi-asserted-by": "crossref",
"journal-title": "Nucleic Acids Res",
"key": "pone.0345051.ref020",
"volume": "46",
"year": "2018"
},
{
"DOI": "10.1093/bioinformatics/btz828",
"article-title": "QMEANDisCo-distance constraints applied on model quality estimation",
"author": "G Studer",
"doi-asserted-by": "crossref",
"first-page": "1765",
"issue": "6",
"journal-title": "Bioinformatics",
"key": "pone.0345051.ref021",
"volume": "36",
"year": "2020"
},
{
"DOI": "10.1385/1-59259-890-0:571",
"article-title": "Protein Identification and Analysis Tools on the ExPASy Server",
"author": "E Gasteiger",
"doi-asserted-by": "crossref",
"first-page": "571",
"journal-title": "The Proteomics Protocols Handbook. Humana Press",
"key": "pone.0345051.ref022",
"year": "2005"
},
{
"article-title": "PubChem in 2021: new data content and improved web interfaces",
"author": "S Kim",
"journal-title": "Nucleic Acids Res",
"key": "pone.0345051.ref023",
"volume": "49",
"year": "2021"
},
{
"DOI": "10.1093/database/bat070",
"article-title": "Phenol-Explorer 3.0: a major update of the Phenol-Explorer database to incorporate data on the effects of food processing on polyphenol content",
"author": "JA Rothwell",
"doi-asserted-by": "crossref",
"journal-title": "Database (Oxford)",
"key": "pone.0345051.ref024",
"volume": "2013",
"year": "2013"
},
{
"DOI": "10.1021/acs.jcim.5b00559",
"article-title": "ZINC 15--Ligand Discovery for Everyone",
"author": "T Sterling",
"doi-asserted-by": "crossref",
"first-page": "2324",
"issue": "11",
"journal-title": "J Chem Inf Model",
"key": "pone.0345051.ref025",
"volume": "55",
"year": "2015"
},
{
"author": "BIOVIA",
"key": "pone.0345051.ref026",
"volume-title": "Discovery Studio modeling environment, release 2021",
"year": "2021"
},
{
"DOI": "10.1186/1758-2946-3-33",
"article-title": "Open Babel: An open chemical toolbox",
"author": "NM O’Boyle",
"doi-asserted-by": "crossref",
"first-page": "33",
"journal-title": "J Cheminform",
"key": "pone.0345051.ref027",
"volume": "3",
"year": "2011"
},
{
"DOI": "10.1007/978-1-4939-2269-7_19",
"article-title": "Small-molecule library screening by docking with PyRx",
"author": "S Dallakyan",
"doi-asserted-by": "crossref",
"first-page": "243",
"journal-title": "Methods Mol Biol",
"key": "pone.0345051.ref028",
"volume": "1263",
"year": "2015"
},
{
"DOI": "10.1002/jcc.21256",
"article-title": "AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility",
"author": "GM Morris",
"doi-asserted-by": "crossref",
"first-page": "2785",
"issue": "16",
"journal-title": "J Comput Chem",
"key": "pone.0345051.ref029",
"volume": "30",
"year": "2009"
},
{
"DOI": "10.1002/jcc.21334",
"article-title": "AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading",
"author": "O Trott",
"doi-asserted-by": "crossref",
"first-page": "455",
"issue": "2",
"journal-title": "J Comput Chem",
"key": "pone.0345051.ref030",
"volume": "31",
"year": "2010"
},
{
"DOI": "10.1093/nar/gky318",
"article-title": "ProTox-II: a webserver for the prediction of toxicity of chemicals",
"author": "P Banerjee",
"doi-asserted-by": "crossref",
"journal-title": "Nucleic Acids Res",
"key": "pone.0345051.ref031",
"volume": "46",
"year": "2018"
},
{
"author": "L Schrödinger",
"key": "pone.0345051.ref032",
"volume-title": "Desmond molecular dynamics system, release 2021-4",
"year": "2021"
},
{
"DOI": "10.1063/1.445869",
"article-title": "Comparison of simple potential functions for simulating liquid water",
"author": "WL Jorgensen",
"doi-asserted-by": "crossref",
"first-page": "926",
"issue": "2",
"journal-title": "J Chem Phys",
"key": "pone.0345051.ref033",
"volume": "79",
"year": "1983"
},
{
"DOI": "10.1021/ct900587b",
"article-title": "Prediction of Absolute Solvation Free Energies using Molecular Dynamics Free Energy Perturbation and the OPLS Force Field",
"author": "D Shivakumar",
"doi-asserted-by": "crossref",
"first-page": "1509",
"issue": "5",
"journal-title": "J Chem Theory Comput",
"key": "pone.0345051.ref034",
"volume": "6",
"year": "2010"
},
{
"article-title": "Evaluating the cardioprotective efficacy of diosmin against LPS-induced cardiac dysfunction: in silico docking and experimental investigations",
"author": "B Yan",
"first-page": "1201",
"issue": "4",
"journal-title": "Pak Vet J",
"key": "pone.0345051.ref035",
"volume": "44",
"year": "2024"
},
{
"DOI": "10.1111/rda.70033",
"article-title": "Phytoestrogens Modulate Bovine G Protein‐Coupled Receptors and Play a Critical Role in Regulating Reproductive Functions in Animals",
"author": "M Safdar",
"doi-asserted-by": "crossref",
"issue": "3",
"journal-title": "Reproduction in Domestic Animals",
"key": "pone.0345051.ref036",
"volume": "60",
"year": "2025"
},
{
"DOI": "10.3906/biy-2012-52",
"article-title": "In silico drug repositioning against human NRP1 to block SARS-CoV-2 host entry",
"author": "Ş Gül",
"doi-asserted-by": "crossref",
"first-page": "442",
"issue": "4",
"journal-title": "Turk J Biol",
"key": "pone.0345051.ref037",
"volume": "45",
"year": "2021"
},
{
"DOI": "10.1021/acs.jmedchem.8b00210",
"article-title": "Small Molecule Neuropilin-1 Antagonists Combine Antiangiogenic and Antitumor Activity with Immune Modulation through Reduction of Transforming Growth Factor Beta (TGFβ) Production in Regulatory T-Cells",
"author": "J Powell",
"doi-asserted-by": "crossref",
"first-page": "4135",
"issue": "9",
"journal-title": "J Med Chem",
"key": "pone.0345051.ref038",
"volume": "61",
"year": "2018"
},
{
"DOI": "10.1080/07391102.2023.2268185",
"article-title": "Identification of phytochemicals as putative ligands for the targeted modulation of peroxisome proliferator-activated receptor α (PPARα) in animals",
"author": "F-U Hassan",
"doi-asserted-by": "crossref",
"first-page": "12164",
"issue": "22",
"journal-title": "J Biomol Struct Dyn",
"key": "pone.0345051.ref039",
"volume": "42",
"year": "2024"
},
{
"article-title": "A review on the use of phytochemicals for the control of zoonotic Giardiasis",
"author": "BS Alawfi",
"first-page": "592",
"issue": "3",
"journal-title": "Pak Vet J",
"key": "pone.0345051.ref040",
"volume": "44",
"year": "2024"
},
{
"article-title": "Efficacy and comparative toxicity of phytochemical compounds extracted from aromatic perennial trees and herbs against vector borne Culex pipiens (Diptera: Culicidae) and Hyalomma dromedarii (Acari: Ixodidae) as green insecticides",
"author": "MM Baz",
"first-page": "55",
"issue": "1",
"journal-title": "Pak Vet J",
"key": "pone.0345051.ref041",
"volume": "44",
"year": "2024"
}
],
"reference-count": 41,
"references-count": 41,
"relation": {},
"resource": {
"primary": {
"URL": "https://dx.plos.org/10.1371/journal.pone.0345051"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Identification of polyphenols as novel neuropilin-1 cendR pocket inhibitors to block SARS-CoV-2 entry and enhance variant resistance",
"type": "journal-article",
"update-policy": "https://doi.org/10.1371/journal.pone.corrections_policy",
"volume": "21"
}
