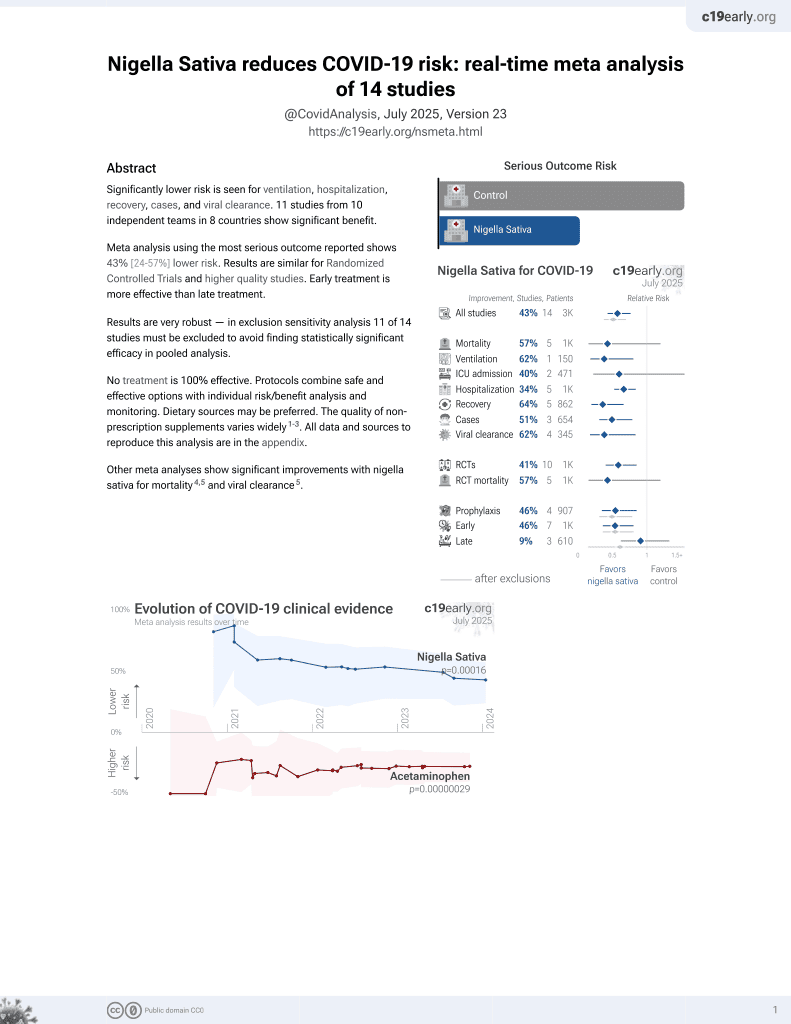
Inhibitory effect of thymoquinone from Nigella sativa against SARS-CoV-2 main protease. An in-silico study
et al., Brazilian Journal of Biology, doi:10.1590/1519-6984.25066, Jan 2022
13th treatment shown to reduce risk in
January 2021, now with p = 0.00016 from 14 studies.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
In silico analysis of thymoquinone from nigella sativa showing good binding affinity with SARS-CoV-2 Mpro.
22 preclinical studies support the efficacy of nigella sativa for COVID-19:
1.
Rahman et al., In Silico Screening of Potential Drug Candidate against Chain a of Coronavirus Binding Protein from Major Nigella Bioactive Compounds, Asian Journal of Advanced Research and Reports, doi:10.9734/ajarr/2024/v18i7697.
2.
Zafar Nayak Snehasis, S., Molecular Docking to Discover Potential Bio-Extract Substitutes for Hydroxychloroquine against COVID-19 and Malaria, International Journal of Science and Research (IJSR), doi:10.21275/SR24323192940.
3.
Alkafaas et al., A study on the effect of natural products against the transmission of B.1.1.529 Omicron, Virology Journal, doi:10.1186/s12985-023-02160-6.
4.
Ali et al., Computational Prediction of Nigella sativa Compounds as Potential Drug Agents for Targeting Spike Protein of SARS-CoV-2, Pakistan BioMedical Journal, doi:10.54393/pbmj.v6i3.853.
5.
Miraz et al., Nigelladine A among Selected Compounds from Nigella sativa Exhibits Propitious Interaction with Omicron Variant of SARS-CoV-2: An In Silico Study, International Journal of Clinical Practice, doi:10.1155/2023/9917306.
6.
Sherwani et al., Pharmacological Profile of Nigella sativa Seeds in Combating COVID-19 through In-Vitro and Molecular Docking Studies, Processes, doi:10.3390/pr10071346.
7.
Khan et al., Inhibitory effect of thymoquinone from Nigella sativa against SARS-CoV-2 main protease. An in-silico study, Brazilian Journal of Biology, doi:10.1590/1519-6984.25066.
8.
Esharkawy et al., In vitro Potential Antiviral SARS-CoV-19- Activity of Natural Product Thymohydroquinone and Dithymoquinone from Nigela sativia, Bioorganic Chemistry, doi:10.1016/j.bioorg.2021.105587.
9.
Banerjee et al., Nigellidine (Nigella sativa, black-cumin seed) docking to SARS CoV-2 nsp3 and host inflammatory proteins may inhibit viral replication/transcription, Natural Product Research, doi:10.1080/14786419.2021.2018430.
10.
Rizvi et al., Identifying the Most Potent Dual-Targeting Compound(s) against 3CLprotease and NSP15exonuclease of SARS-CoV-2 from Nigella sativa: Virtual Screening via Physicochemical Properties, Docking and Dynamic Simulation Analysis, Processes, doi:10.3390/pr9101814.
11.
Mir et al., Identification of SARS-CoV-2 RNA-dependent RNA polymerase inhibitors from the major phytochemicals of Nigella sativa: An in silico approach, Saudi Journal of Biological Sciences, doi:10.1016/j.sjbs.2021.09.002.
12.
Hardianto et al., Exploring the Potency of Nigella sativa Seed in Inhibiting SARS-CoV-2 Main Protease Using Molecular Docking and Molecular Dynamics Simulations, Indonesian Journal of Chemistry, doi:10.22146/ijc.65951.
13.
Maiti et al., Active-site Molecular docking of Nigellidine to nucleocapsid/Nsp2/Nsp3/MPro of COVID-19 and to human IL1R and TNFR1/2 may stop viral-growth/cytokine-flood, and the drug source Nigella sativa (black cumin) seeds show potent antioxidant role in experimental rats, Research Square, doi:10.21203/rs.3.rs-26464/v1.
14.
Duru et al., In silico identification of compounds from Nigella sativa seed oil as potential inhibitors of SARS-CoV-2 targets, Bulletin of the National Research Centre, doi:10.1186/s42269-021-00517-x.
15.
Bouchentouf et al., Identification of Compounds from Nigella Sativa as New Potential Inhibitors of 2019 Novel Coronasvirus (Covid-19): Molecular Docking Study, ChemRxiv, doi:10.26434/chemrxiv.12055716.v1.
16.
Ali (B) et al., In vitro inhibitory effect of Nigella sativa L. extracts on SARS-COV-2 spike protein-ACE2 interaction, Current Therapeutic Research, doi:10.1016/j.curtheres.2024.100759.
17.
Bostancıklıoğlu et al., Nigella sativa, Anthemis hyaline and Citrus sinensis extracts reduce SARS-CoV-2 replication by fluctuating Rho GTPase, PI3K-AKT, and MAPK/ERK pathways in HeLa-CEACAM1a cells, Gene, doi:10.1016/j.gene.2024.148366.
Khan et al., 24 Jan 2022, peer-reviewed, 7 authors.
Contact: tahirmicrobiologist@gmail.com, muhammad.tahir8@imbb.uol.edu.pk, dqwei@sjtu.edu.cn.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
Inhibitory effect of thymoquinone from Nigella sativa against SARS-CoV-2 main protease. An in-silico study
Brazilian Journal of Biology, doi:10.1590/1519-6984.25066
Nigella sativa is known for the safety profile, containing a wealth of useful antiviral compounds. The main protease (Mpro, 3CLpro) of severe acute respiratory syndrome 2 (SARS-CoV-2) is being considered as one of the most attractive viral target, processing the polyproteins during viral pathogenesis and replication. In the current investigation we analyzed the potency of active component, thymoquinone (TQ) of Nigella sativa against SARS-CoV-2 Mpro. The structures of TQ and Mpro was retrieved from PubChem (CID10281) and Protein Data Bank (PDB ID 6MO3) respectively. The Mpro and TQ were docked and the complex was subjected to molecular dynamic (MD) simulations for a period 50ns. Protein folding effect was analyzed using radius of gyration (Rg) while stability and flexibility was measured, using root means square deviations (RMSD) and root means square fluctuation (RMSF) respectively. The simulation results shows that TQ is exhibiting good binding activity against SARS-CoV-2 Mpro, interacting many residues, present in the active site (His41, Cys145) and also the Glu166, facilitating the pocket shape. Further, experimental approaches are needed to validate the role of TQ against virus infection. The TQ is interfering with pocket maintaining residues as well as active site of virus Mpro which may be used as a potential inhibitor against SARS-CoV-2 for better management of COVID-19.
References
Ahlatci, Kuzhan, Taysi, Demirtas, Alkis et al., Radiation-modifying abilities of Nigella sativa and thymoquinone on radiation-induced nitrosative stress in the brain tissue, Phytomedicine, doi:10.1016/j.phymed.2013.10.023
Amin, Hosseinzadeh, Black cumin (Nigella sativa) and its active constituent, thymoquinone: an overview on the analgesic and anti-inflammatory effects, Planta Medica, doi:10.1055/s-0035-1557838
Anand, Palm, Mesters, Siddell, Ziebuhr et al., Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain, The EMBO Journal, doi:10.1093/emboj/cdf327
Anand, Yang, Bartlam, Rao, Hilgenfeld, Coronavirus main proteinase: target for antiviral drug therapy, doi:10.1007/3-7643-7339-3_9
Bavi, Kumar, Choi, Lee, Exploration of novel inhibitors for Bruton's tyrosine kinase by 3D QSAR modeling and molecular dynamics simulation, PLoS One, doi:10.1371/journal.pone.0147190
Berendsen, Van Der Spoel, Van Drunen, GROMACS: a message-passing parallel molecular dynamics implementation, Computer Physics Communications, doi:10.1016/0010-4655(95)00042-E
Berhanu, Masunov, Molecular dynamic simulation of wild type and mutants of the polymorphic amyloid NNQNTF segments of elk prion: structural stability and thermodynamic of association, Biopolymers, doi:10.1002/bip.21611
Berman, Westbrook, Feng, Gilliland, Bhat et al., The Protein Data Bank, Nucleic Acids Research, doi:10.1093/nar/28.1.235
Bernstein, Koetzle, Williams, Meyer, Brice et al., The Protein Data Bank: a computerbased archival file for macromolecular structures, Journal of Molecular Biology, doi:10.1016/S0022-2836(77)80200-3
Brüstle, Beck, Schindler, King, Mitchell et al., Descriptors, physical properties, and druglikeness, Journal of Medicinal Chemistry, doi:10.1021/jm011027b
Chen, Shen, Computational analysis of amino acid mutation: a proteome wide perspective, Current Proteomics, doi:10.2174/157016409789973734
Chong, Lee, Kang, Park, Ham, Structural and thermodynamic investigations on the aggregation and folding of acylphosphatase by molecular dynamics simulations and solvation free energy analysis, Journal of the American Chemical Society, doi:10.1021/ja1116233
Daina, Michielin, Zoete, SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules, Scientific Reports, doi:10.1038/srep42717
Darden, York, Pedersen, Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems, The Journal of Chemical Physics, doi:10.1063/1.464397
Enomoto, Asano, Iwahori, Narui, Okada et al., Hematological studies on black cumin oil from the seeds of Nigella sativa L, Biological & Pharmaceutical Bulletin, doi:10.1248/bpb.24.307
Essmann, Perera, Berkowitz, Darden, Lee et al., A smooth particle mesh Ewald method, The Journal of Chemical Physics, doi:10.1063/1.470117
Gan, Huang, Huang, Rao, Zhao et al., Synthesis and activity of an octapeptide inhibitor designed for SARS coronavirus main proteinase, Peptides, doi:10.1016/j.peptides.2005.09.006
Götz, Williamson, Xu, Poole, Grand et al., Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born, Journal of Chemical Theory and Computation, doi:10.1021/ct200909j
He, Li, Zheng, Zhang, A molecular dynamics investigation into the mechanisms of alectinib resistance of three ALK mutants, Journal of Cellular Biochemistry, doi:10.1002/jcb.26666
Hollingsworth, Dror, Molecular dynamics simulation for all, Neuron, doi:10.1016/j.neuron.2018.08.011
Hosseinzadeh, Tafaghodi, Mosavi, Taghiabadi, Effect of aqueous and ethanolic extracts of Nigella sativa seeds on milk production in rats, Journal of Acupuncture and Meridian Studies, doi:10.1016/j.jams.2012.07.019
Islam, Biological activities and therapeutic promises of Nigella sativa L, International Journal of Pharma Sciences and Scientific Research, doi:10.25141/2471-6782-2016-6.0237
Ismail, Jusoh, Molecular docking and molecular dynamics simulation studies to predict flavonoid binding on the surface of DENV2 E protein, Interdisciplinary Sciences, Computational Life Sciences, doi:10.1007/s12539-016-0157-8
Jorgensen, Chandrasekhar, Madura, Impey, Klein, Comparison of simple potential functions for simulating liquid water, The Journal of Chemical Physics, doi:10.1063/1.445869
Khan, Ali, Wang, Irfan, Khan et al., Marine natural compounds as potents inhibitors against the main protease of SARS-CoV-2: a molecular dynamic study, Journal of Biomolecular Structure & Dynamics, doi:10.1080/07391102.2020.1769733
Khan, Ali, Zeb, Kaushik, Malik et al., Gibbs free energy calculation of mutation in PncA and RpsA associated with pyrazinamide resistance, Frontiers in Molecular Biosciences, doi:10.3389/fmolb.2020.00052
Khan, Chen, Tania, Zhang, Anticancer activities of Nigella sativa (black cumin), African Journal of Traditional, Complementary, and Alternative Medicines, doi:10.4314/ajtcam.v8i5S.10
Khan, Gastroprotective activity of Nigella sativa oil and its constituent, thymoquinone, against gastric mucosal injury induced by ischaemia/reperfusion in rats, Journal of Ethnopharmacology, doi:10.1016/S0378-8741(02)00324-0
Khan, Khan, Rehman, Wang, Akhtar et al., Structural and free energy landscape of novel mutations in ribosomal protein S1 (rpsA) associated with pyrazinamide resistance, Scientific Reports, doi:10.1038/s41598-019-44013-9
Khare, Sahu, Pandey, Samant, Current approaches for target-specific drug discovery using natural compounds against SARS-CoV-2 infection, Virus Research, doi:10.1016/j.virusres.2020.198169
Lee, Worrall, Vuckovic, Rosell, Gentile et al., Crystallographic structure of wildtype SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site, Nature Communications, doi:10.1038/s41467-020-19662-4
Li, Wang, Ren, Antiviral mechanisms of candidate chemical medicines and traditional Chinese medicines for SARS-CoV-2 infection, Virus Research, doi:10.1016/j.virusres.2020.198073
Liang, Characterization and inhibition of SARScoronavirus main protease, Current Topics in Medicinal Chemistry, doi:10.2174/156802606776287090
Lim, Shi, Mu, Song, Dynamically-driven enhancement of the catalytic machinery of the SARS 3C-like protease by the S284-T285-I286/A mutations on the extra domain, PLoS One, doi:10.1371/journal.pone.0101941
Liu, Shi, Zhou, Liu, Liu et al., Molecular dynamics simulations and novel drug discovery, Expert Opinion on Drug Discovery, doi:10.1080/17460441.2018.1403419
Liu, Yao, Molecular basis of the interaction for an essential subunit PA-PB1 in influenza virus RNA polymerase: insights from molecular dynamics simulation and free energy calculation, Molecular Pharmaceutics, doi:10.1021/mp900131p
Lobanov, Bogatyreva, Galzitskaya, Radius of gyration as an indicator of protein structure compactness, Molecular Biology, doi:10.1134/S0026893308040195
Nagasundaram, Zhu, Liu, Karthick, Chakraborty et al., Analysing the effect of mutation on protein function and discovering potential inhibitors of CDK4: molecular modelling and dynamics studies, PLoS One, doi:10.1371/journal.pone.0133969
Pillaiyar, Manickam, Namasivayam, Hayashi, Jung, An overview of severe acute respiratory syndrome-coronavirus (SARS-CoV) 3CL protease inhibitors: peptidomimetics and small molecule chemotherapy, Journal of Medicinal Chemistry, doi:10.1021/acs.jmedchem.5b01461
Rodrigues, Pires, Ascher, DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability, Nucleic Acids Research, doi:10.1093/nar/gky300
Salem, Hossain, Protective effect of black seed oil from Nigella sativa against murine cytomegalovirus infection, International Journal of Immunopharmacology, doi:10.1016/S0192-0561(00)00036-9
Salomon-Ferrer, Case, Walker, An overview of the Amber biomolecular simulation package, WIREs Computational Molecular Science, doi:10.1002/wcms.1121
Setlur, Naik, Skariyachan, Herbal lead as ideal bioactive compounds against probable drug targets of ebola virus in comparison with known chemical analogue: a computational drug discovery perspective, Interdisciplinary Sciences, Computational Life Sciences, doi:10.1007/s12539-016-0149-8
Shi, Song, The catalysis of the SARS 3C-like protease is under extensive regulation by its extra domain, The FEBS Journal, doi:10.1111/j.1742-4658.2006.05130.x
Smilgies, Molecular weightgyration radius relation of globular proteins: a comparison of light scattering, small-angle X-ray scattering and structure-based data, Journal of Applied Crystallography, doi:10.1107/S1600576715015551
Sun, Li, Shen, Tian, Xu et al., Assessing the performance of MM/PBSA and MM/ GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring, Physical Chemistry Chemical Physics, doi:10.1039/C4CP03179B
Sun, Li, Tian, Xu, Hou, Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set, Physical Chemistry Chemical Physics, doi:10.1039/C4CP01388C
Umar, Munir, Subhan, Azam, Nisa et al., Protective and antiviral activities of Nigella sativa against avian influenza (H9N2) in turkeys, Journal of the Saudi Society of Agricultural Sciences, doi:10.1016/j.jssas.2016.09.004
Ursu, Rayan, Goldblum, Oprea, Understanding drug-likeness, WIREs Computational Molecular Science, doi:10.1002/wcms.52
Varadwaj, Flavonoids as multi-target inhibitors for proteins associated with ebola virus: in silico Brazilian, Journal of Biology
Vilar, Cozza, Moro, Medicinal chemistry and the Molecular Operating Environment (MOE): application of QSAR and molecular docking to drug discovery, Current Topics in Medicinal Chemistry, doi:10.2174/156802608786786624
Vistoli, Pedretti, Testa, Assessing druglikeness -what are we missing?, Drug Discovery Today, doi:10.1016/j.drudis.2007.11.007
Walters, Murcko, Prediction of 'druglikeness, Advanced Drug Delivery Reviews, doi:10.1016/S0169-409X(02)00003-0
Xue, Yu, Yang, Xue, Wu et al., Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design, Journal of Virology, doi:10.1128/JVI.02114-07
Zhang, Lin, Sun, Curth, Drosten et al., Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors, Interdisciplinary Sciences, Computational Life Sciences, doi:10.1007/s12539-015-0109-8
DOI record:
{
"DOI": "10.1590/1519-6984.25066",
"ISSN": [
"1678-4375",
"1519-6984"
],
"URL": "http://dx.doi.org/10.1590/1519-6984.25066",
"abstract": "<jats:p>Abstract Nigella sativa is known for the safety profile, containing a wealth of useful antiviral compounds. The main protease (Mpro, 3CLpro) of severe acute respiratory syndrome 2 (SARS-CoV-2) is being considered as one of the most attractive viral target, processing the polyproteins during viral pathogenesis and replication. In the current investigation we analyzed the potency of active component, thymoquinone (TQ) of Nigella sativa against SARS-CoV-2 Mpro. The structures of TQ and Mpro was retrieved from PubChem (CID10281) and Protein Data Bank (PDB ID 6MO3) respectively. The Mpro and TQ were docked and the complex was subjected to molecular dynamic (MD) simulations for a period 50ns. Protein folding effect was analyzed using radius of gyration (Rg) while stability and flexibility was measured, using root means square deviations (RMSD) and root means square fluctuation (RMSF) respectively. The simulation results shows that TQ is exhibiting good binding activity against SARS-CoV-2 Mpro, interacting many residues, present in the active site (His41, Cys145) and also the Glu166, facilitating the pocket shape. Further, experimental approaches are needed to validate the role of TQ against virus infection. The TQ is interfering with pocket maintaining residues as well as active site of virus Mpro which may be used as a potential inhibitor against SARS-CoV-2 for better management of COVID-19.</jats:p>",
"alternative-id": [
"S1519-69842024000100270"
],
"author": [
{
"ORCID": "http://orcid.org/0000-0003-1158-2133",
"affiliation": [
{
"name": "The University of Lahore, Pakistan"
}
],
"authenticated-orcid": false,
"family": "Khan",
"given": "M. T.",
"sequence": "first"
},
{
"ORCID": "http://orcid.org/0000-0002-1420-3525",
"affiliation": [
{
"name": "Shanghai Jiao Tong University, China"
}
],
"authenticated-orcid": false,
"family": "Ali",
"given": "A.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-2591-7021",
"affiliation": [
{
"name": "Shanghai Jiao Tong University, China"
}
],
"authenticated-orcid": false,
"family": "Wei",
"given": "X.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-0104-5484",
"affiliation": [
{
"name": "University of The Punjab, Pakistan"
}
],
"authenticated-orcid": false,
"family": "Nadeem",
"given": "T.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0003-4908-3313",
"affiliation": [
{
"name": "King Khalid University, Saudi Arabia"
}
],
"authenticated-orcid": false,
"family": "Muhammad",
"given": "S.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-6793-3038",
"affiliation": [
{
"name": "King Khalid University, Saudi Arabia"
}
],
"authenticated-orcid": false,
"family": "Al-Sehemi",
"given": "A. G.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-6770-7102",
"affiliation": [
{
"name": "Shanghai Jiao Tong University, China; Peng Cheng Laboratory, China"
}
],
"authenticated-orcid": false,
"family": "Wei",
"given": "Dongqing",
"sequence": "additional"
}
],
"container-title": "Brazilian Journal of Biology",
"container-title-short": "Braz. J. Biol.",
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2022,
4,
15
]
],
"date-time": "2022-04-15T19:24:49Z",
"timestamp": 1650050689000
},
"deposited": {
"date-parts": [
[
2022,
4,
15
]
],
"date-time": "2022-04-15T19:25:39Z",
"timestamp": 1650050739000
},
"indexed": {
"date-parts": [
[
2022,
4,
16
]
],
"date-time": "2022-04-16T10:11:09Z",
"timestamp": 1650103869212
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2024
]
]
},
"license": [
{
"URL": "http://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
1,
1
]
],
"date-time": "2024-01-01T00:00:00Z",
"timestamp": 1704067200000
}
},
{
"URL": "http://creativecommons.org/licenses/by/4.0/",
"content-version": "am",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
1,
1
]
],
"date-time": "2024-01-01T00:00:00Z",
"timestamp": 1704067200000
}
},
{
"URL": "http://creativecommons.org/licenses/by/4.0/",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2024,
1,
1
]
],
"date-time": "2024-01-01T00:00:00Z",
"timestamp": 1704067200000
}
}
],
"link": [
{
"URL": "http://www.scielo.br/scielo.php?script=sci_pdf&pid=S1519-69842024000100270&tlng=en",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "530",
"original-title": [
"Efeito inibitório da timoquinona de Nigella sativa contra a principal protease do SARS-CoV-2. Um estudo in silico"
],
"prefix": "10.1590",
"published": {
"date-parts": [
[
2024
]
]
},
"published-online": {
"date-parts": [
[
2024
]
]
},
"publisher": "FapUNIFESP (SciELO)",
"reference": [
{
"DOI": "10.1016/j.phymed.2013.10.023",
"article-title": "Radiation-modifying abilities of Nigella sativa and thymoquinone on radiation-induced nitrosative stress in the brain tissue",
"author": "AHLATCI A.",
"doi-asserted-by": "crossref",
"first-page": "740",
"issue": "5",
"journal-title": "Phytomedicine",
"key": "ref1",
"volume": "21",
"year": "2014"
},
{
"article-title": "Black cumin (Nigella sativa) and its active constituent, thymoquinone: an overview on the analgesic and anti-inflammatory effects",
"author": "AMIN B.",
"first-page": "8",
"issue": "1-2",
"journal-title": "Planta Medica",
"key": "ref2",
"volume": "82",
"year": "2016"
},
{
"DOI": "10.1093/emboj/cdf327",
"article-title": "Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain",
"author": "ANAND K.",
"doi-asserted-by": "crossref",
"first-page": "3213",
"issue": "13",
"journal-title": "The EMBO Journal",
"key": "ref3",
"volume": "21",
"year": "2002"
},
{
"author": "ANAND K.",
"first-page": "173",
"key": "ref4",
"series-title": "Coronaviruses with special emphasis on first insights concerning SARS",
"volume-title": "Coronavirus main proteinase: target for antiviral drug therapy",
"year": "2005"
},
{
"DOI": "10.1371/journal.pone.0147190",
"article-title": "Exploration of novel inhibitors for Bruton’s tyrosine kinase by 3D QSAR modeling and molecular dynamics simulation",
"author": "BAVI R.",
"doi-asserted-by": "crossref",
"issue": "1",
"journal-title": "PLoS One",
"key": "ref5",
"volume": "11",
"year": "2016"
},
{
"DOI": "10.1016/0010-4655(95)00042-E",
"article-title": "GROMACS: a message-passing parallel molecular dynamics implementation",
"author": "BERENDSEN H.J.C.",
"doi-asserted-by": "crossref",
"first-page": "43",
"issue": "1-3",
"journal-title": "Computer Physics Communications",
"key": "ref6",
"volume": "91",
"year": "1995"
},
{
"DOI": "10.1002/bip.21611",
"article-title": "Molecular dynamic simulation of wild type and mutants of the polymorphic amyloid NNQNTF segments of elk prion: structural stability and thermodynamic of association",
"author": "BERHANU W.M.",
"doi-asserted-by": "crossref",
"first-page": "573",
"issue": "9",
"journal-title": "Biopolymers",
"key": "ref7",
"volume": "95",
"year": "2011"
},
{
"DOI": "10.1093/nar/28.1.235",
"article-title": "The Protein Data Bank",
"author": "BERMAN H.M.",
"doi-asserted-by": "crossref",
"first-page": "235",
"issue": "1",
"journal-title": "Nucleic Acids Research",
"key": "ref8",
"volume": "28",
"year": "2000"
},
{
"DOI": "10.1016/S0022-2836(77)80200-3",
"article-title": "The Protein Data Bank: a computer-based archival file for macromolecular structures",
"author": "BERNSTEIN F.C.",
"doi-asserted-by": "crossref",
"first-page": "535",
"issue": "3",
"journal-title": "Journal of Molecular Biology",
"key": "ref9",
"volume": "112",
"year": "1977"
},
{
"DOI": "10.1021/jm011027b",
"article-title": "Descriptors, physical properties, and drug-likeness",
"author": "BRÜSTLE M.",
"doi-asserted-by": "crossref",
"first-page": "3345",
"issue": "16",
"journal-title": "Journal of Medicinal Chemistry",
"key": "ref10",
"volume": "45",
"year": "2002"
},
{
"DOI": "10.2174/157016409789973734",
"article-title": "Computational analysis of amino acid mutation: a proteome wide perspective",
"author": "CHEN J.",
"doi-asserted-by": "crossref",
"first-page": "228",
"issue": "4",
"journal-title": "Current Proteomics",
"key": "ref11",
"volume": "6",
"year": "2009"
},
{
"DOI": "10.1021/ja1116233",
"article-title": "Structural and thermodynamic investigations on the aggregation and folding of acylphosphatase by molecular dynamics simulations and solvation free energy analysis",
"author": "CHONG S.-H.",
"doi-asserted-by": "crossref",
"first-page": "7075",
"issue": "18",
"journal-title": "Journal of the American Chemical Society",
"key": "ref12",
"volume": "133",
"year": "2011"
},
{
"DOI": "10.1038/srep42717",
"article-title": "SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules",
"author": "DAINA A.",
"doi-asserted-by": "crossref",
"first-page": "42717",
"issue": "1",
"journal-title": "Scientific Reports",
"key": "ref13",
"volume": "7",
"year": "2017"
},
{
"DOI": "10.1063/1.464397",
"article-title": "Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems",
"author": "DARDEN T.",
"doi-asserted-by": "crossref",
"first-page": "10089",
"issue": "12",
"journal-title": "The Journal of Chemical Physics",
"key": "ref14",
"volume": "98",
"year": "1993"
},
{
"DOI": "10.1016/S0378-8741(02)00324-0",
"article-title": "Gastroprotective activity of Nigella sativa oil and its constituent, thymoquinone, against gastric mucosal injury induced by ischaemia/reperfusion in rats",
"author": "EL-ABHAR H.S.",
"doi-asserted-by": "crossref",
"first-page": "251",
"issue": "2-3",
"journal-title": "Journal of Ethnopharmacology",
"key": "ref15",
"volume": "84",
"year": "2003"
},
{
"DOI": "10.1248/bpb.24.307",
"article-title": "Hematological studies on black cumin oil from the seeds of Nigella sativa L",
"author": "ENOMOTO S.",
"doi-asserted-by": "crossref",
"first-page": "307",
"issue": "3",
"journal-title": "Biological & Pharmaceutical Bulletin",
"key": "ref16",
"volume": "24",
"year": "2001"
},
{
"DOI": "10.1063/1.470117",
"article-title": "A smooth particle mesh Ewald method",
"author": "ESSMANN U.",
"doi-asserted-by": "crossref",
"first-page": "8577",
"issue": "19",
"journal-title": "The Journal of Chemical Physics",
"key": "ref17",
"volume": "103",
"year": "1995"
},
{
"DOI": "10.1016/j.peptides.2005.09.006",
"article-title": "Synthesis and activity of an octapeptide inhibitor designed for SARS coronavirus main proteinase",
"author": "GAN Y.-R.",
"doi-asserted-by": "crossref",
"first-page": "622",
"issue": "4",
"journal-title": "Peptides",
"key": "ref18",
"volume": "27",
"year": "2006"
},
{
"DOI": "10.1021/ct200909j",
"article-title": "Routine microsecond molecular dynamics simulations with AMBER on GPUs. 1. Generalized born",
"author": "GÖTZ A.W.",
"doi-asserted-by": "crossref",
"first-page": "1542",
"issue": "5",
"journal-title": "Journal of Chemical Theory and Computation",
"key": "ref19",
"volume": "8",
"year": "2012"
},
{
"DOI": "10.1002/jcb.26666",
"article-title": "A molecular dynamics investigation into the mechanisms of alectinib resistance of three ALK mutants",
"author": "HE M.",
"doi-asserted-by": "crossref",
"first-page": "5332",
"issue": "7",
"journal-title": "Journal of Cellular Biochemistry",
"key": "ref20",
"volume": "119",
"year": "2018"
},
{
"DOI": "10.1016/j.neuron.2018.08.011",
"article-title": "Molecular dynamics simulation for all",
"author": "HOLLINGSWORTH S.A.",
"doi-asserted-by": "crossref",
"first-page": "1129",
"issue": "6",
"journal-title": "Neuron",
"key": "ref21",
"volume": "99",
"year": "2018"
},
{
"DOI": "10.1016/j.jams.2012.07.019",
"article-title": "Effect of aqueous and ethanolic extracts of Nigella sativa seeds on milk production in rats",
"author": "HOSSEINZADEH H.",
"doi-asserted-by": "crossref",
"first-page": "18",
"issue": "1",
"journal-title": "Journal of Acupuncture and Meridian Studies",
"key": "ref22",
"volume": "6",
"year": "2013"
},
{
"DOI": "10.25141/2471-6782-2016-6.0237",
"article-title": "Biological activities and therapeutic promises of Nigella sativa L",
"author": "ISLAM M.T.",
"doi-asserted-by": "crossref",
"first-page": "237",
"issue": "6",
"journal-title": "International Journal of Pharma Sciences and Scientific Research",
"key": "ref23",
"volume": "2",
"year": "2016"
},
{
"DOI": "10.1007/s12539-016-0157-8",
"article-title": "Molecular docking and molecular dynamics simulation studies to predict flavonoid binding on the surface of DENV2 E protein",
"author": "ISMAIL N.A.",
"doi-asserted-by": "crossref",
"first-page": "499",
"issue": "4",
"journal-title": "Interdisciplinary Sciences, Computational Life Sciences",
"key": "ref24",
"volume": "9",
"year": "2017"
},
{
"DOI": "10.1063/1.445869",
"article-title": "Comparison of simple potential functions for simulating liquid water",
"author": "JORGENSEN W.L.",
"doi-asserted-by": "crossref",
"first-page": "926",
"issue": "2",
"journal-title": "The Journal of Chemical Physics",
"key": "ref25",
"volume": "79",
"year": "1983"
},
{
"DOI": "10.4314/ajtcam.v8i5S.10",
"article-title": "Anticancer activities of Nigella sativa (black cumin)",
"author": "KHAN A.",
"doi-asserted-by": "crossref",
"first-page": "226",
"issue": "5S",
"journal-title": "African Journal of Traditional, Complementary, and Alternative Medicines",
"key": "ref26",
"volume": "8",
"year": "2011"
},
{
"DOI": "10.1080/07391102.2020.1769733",
"article-title": "Marine natural compounds as potents inhibitors against the main protease of SARS-CoV-2: a molecular dynamic study",
"author": "KHAN M.T.",
"doi-asserted-by": "crossref",
"first-page": "3627",
"issue": "10",
"journal-title": "Journal of Biomolecular Structure & Dynamics",
"key": "ref27",
"volume": "39",
"year": "2021"
},
{
"DOI": "10.3389/fmolb.2020.00052",
"article-title": "Gibbs free energy calculation of mutation in PncA and RpsA associated with pyrazinamide resistance",
"author": "KHAN M.T.",
"doi-asserted-by": "crossref",
"first-page": "52",
"journal-title": "Frontiers in Molecular Biosciences",
"key": "ref28",
"volume": "7",
"year": "2020"
},
{
"DOI": "10.1038/s41598-019-44013-9",
"article-title": "Structural and free energy landscape of novel mutations in ribosomal protein S1 (rpsA) associated with pyrazinamide resistance",
"author": "KHAN M.T.",
"doi-asserted-by": "crossref",
"first-page": "7482",
"issue": "1",
"journal-title": "Scientific Reports",
"key": "ref29",
"volume": "9",
"year": "2019"
},
{
"DOI": "10.1016/j.virusres.2020.198169",
"article-title": "Current approaches for target-specific drug discovery using natural compounds against SARS-CoV-2 infection",
"author": "KHARE P.",
"doi-asserted-by": "crossref",
"journal-title": "Virus Research",
"key": "ref30",
"volume": "290",
"year": "2020"
},
{
"DOI": "10.1038/s41467-020-19662-4",
"article-title": "Crystallographic structure of wild-type SARS-CoV-2 main protease acyl-enzyme intermediate with physiological C-terminal autoprocessing site",
"author": "LEE J.",
"doi-asserted-by": "crossref",
"first-page": "5877",
"issue": "1",
"journal-title": "Nature Communications",
"key": "ref31",
"volume": "11",
"year": "2020"
},
{
"article-title": "Antiviral mechanisms of candidate chemical medicines and traditional Chinese medicines for SARS-CoV-2 infection",
"author": "LI C.",
"journal-title": "Virus Research",
"key": "ref32",
"volume": "286",
"year": "2020"
},
{
"DOI": "10.2174/156802606776287090",
"article-title": "Characterization and inhibition of SARS-coronavirus main protease",
"author": "LIANG P.-H.",
"doi-asserted-by": "crossref",
"first-page": "361",
"issue": "4",
"journal-title": "Current Topics in Medicinal Chemistry",
"key": "ref33",
"volume": "6",
"year": "2006"
},
{
"article-title": "Dynamically-driven enhancement of the catalytic machinery of the SARS 3C-like protease by the S284-T285-I286/A mutations on the extra domain",
"author": "LIM L.",
"issue": "7",
"journal-title": "PLoS One",
"key": "ref34",
"volume": "9",
"year": "2014"
},
{
"DOI": "10.1021/mp900131p",
"article-title": "Molecular basis of the interaction for an essential subunit PA-PB1 in influenza virus RNA polymerase: insights from molecular dynamics simulation and free energy calculation",
"author": "LIU H.",
"doi-asserted-by": "crossref",
"first-page": "75",
"issue": "1",
"journal-title": "Molecular Pharmaceutics",
"key": "ref35",
"volume": "7",
"year": "2010"
},
{
"DOI": "10.1080/17460441.2018.1403419",
"article-title": "Molecular dynamics simulations and novel drug discovery",
"author": "LIU X.",
"doi-asserted-by": "crossref",
"first-page": "23",
"issue": "1",
"journal-title": "Expert Opinion on Drug Discovery",
"key": "ref36",
"volume": "13",
"year": "2018"
},
{
"DOI": "10.1134/S0026893308040195",
"article-title": "Radius of gyration as an indicator of protein structure compactness",
"author": "LOBANOV M.Y.",
"doi-asserted-by": "crossref",
"first-page": "623",
"journal-title": "Molecular Biology",
"key": "ref37",
"volume": "42",
"year": "2008"
},
{
"article-title": "Analysing the effect of mutation on protein function and discovering potential inhibitors of CDK4: molecular modelling and dynamics studies",
"author": "NAGASUNDARAM N.",
"issue": "8",
"journal-title": "PLoS One",
"key": "ref38",
"volume": "10",
"year": "2015"
},
{
"DOI": "10.1021/acs.jmedchem.5b01461",
"article-title": "An overview of severe acute respiratory syndrome-coronavirus (SARS-CoV) 3CL protease inhibitors: peptidomimetics and small molecule chemotherapy",
"author": "PILLAIYAR T.",
"doi-asserted-by": "crossref",
"first-page": "6595",
"issue": "14",
"journal-title": "Journal of Medicinal Chemistry",
"key": "ref39",
"volume": "59",
"year": "2016"
},
{
"DOI": "10.1007/s12539-015-0109-8",
"article-title": "Flavonoids as multi-target inhibitors for proteins associated with ebola virus: in silico discovery using virtual screening and molecular docking studies",
"author": "RAJ U.",
"doi-asserted-by": "crossref",
"first-page": "132",
"issue": "2",
"journal-title": "Interdisciplinary Sciences, Computational Life Sciences",
"key": "ref40",
"volume": "8",
"year": "2016"
},
{
"DOI": "10.1093/nar/gky300",
"article-title": "DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability",
"author": "RODRIGUES C.H.",
"doi-asserted-by": "crossref",
"first-page": "W350",
"issue": "W1",
"journal-title": "Nucleic Acids Research",
"key": "ref41",
"volume": "46",
"year": "2018"
},
{
"DOI": "10.1016/S0192-0561(00)00036-9",
"article-title": "Protective effect of black seed oil from Nigella sativa against murine cytomegalovirus infection",
"author": "SALEM M.L.",
"doi-asserted-by": "crossref",
"first-page": "729",
"issue": "9",
"journal-title": "International Journal of Immunopharmacology",
"key": "ref42",
"volume": "22",
"year": "2000"
},
{
"DOI": "10.1002/wcms.1121",
"article-title": "An overview of the Amber biomolecular simulation package",
"author": "SALOMON‐FERRER R.",
"doi-asserted-by": "crossref",
"first-page": "198",
"issue": "2",
"journal-title": "WIREs Computational Molecular Science",
"key": "ref43",
"volume": "3",
"year": "2013"
},
{
"DOI": "10.1007/s12539-016-0149-8",
"article-title": "Herbal lead as ideal bioactive compounds against probable drug targets of ebola virus in comparison with known chemical analogue: a computational drug discovery perspective",
"author": "SETLUR A.S.",
"doi-asserted-by": "crossref",
"first-page": "254",
"issue": "2",
"journal-title": "Interdisciplinary Sciences, Computational Life Sciences",
"key": "ref44",
"volume": "9",
"year": "2017"
},
{
"DOI": "10.1111/j.1742-4658.2006.05130.x",
"article-title": "The catalysis of the SARS 3C-like protease is under extensive regulation by its extra domain",
"author": "SHI J.",
"doi-asserted-by": "crossref",
"first-page": "1035",
"issue": "5",
"journal-title": "The FEBS Journal",
"key": "ref45",
"volume": "273",
"year": "2006"
},
{
"DOI": "10.1107/S1600576715015551",
"article-title": "Molecular weight–gyration radius relation of globular proteins: a comparison of light scattering, small-angle X-ray scattering and structure-based data",
"author": "SMILGIES D.-M.",
"doi-asserted-by": "crossref",
"first-page": "1604",
"journal-title": "Journal of Applied Crystallography",
"key": "ref46",
"volume": "48",
"year": "2015"
},
{
"DOI": "10.1039/C4CP03179B",
"article-title": "Assessing the performance of MM/PBSA and MM/GBSA methods. 5. Improved docking performance using high solute dielectric constant MM/GBSA and MM/PBSA rescoring",
"author": "SUN H.",
"doi-asserted-by": "crossref",
"first-page": "22035",
"issue": "40",
"journal-title": "Physical Chemistry Chemical Physics",
"key": "ref47",
"volume": "16",
"year": "2014"
},
{
"DOI": "10.1039/C4CP01388C",
"article-title": "Assessing the performance of MM/PBSA and MM/GBSA methods. 4. Accuracies of MM/PBSA and MM/GBSA methodologies evaluated by various simulation protocols using PDBbind data set",
"author": "SUN H.",
"doi-asserted-by": "crossref",
"first-page": "16719",
"issue": "31",
"journal-title": "Physical Chemistry Chemical Physics",
"key": "ref48",
"volume": "16",
"year": "2014"
},
{
"article-title": "Protective and antiviral activities of Nigella sativa against avian influenza (H9N2) in turkeys",
"author": "UMAR S.",
"journal-title": "Journal of the Saudi Society of Agricultural Sciences",
"key": "ref49",
"year": "2016"
},
{
"DOI": "10.1002/wcms.52",
"article-title": "Understanding drug-likeness",
"author": "URSU O.",
"doi-asserted-by": "crossref",
"first-page": "760",
"issue": "5",
"journal-title": "WIREs Computational Molecular Science",
"key": "ref50",
"volume": "1",
"year": "2011"
},
{
"DOI": "10.2174/156802608786786624",
"article-title": "Medicinal chemistry and the Molecular Operating Environment (MOE): application of QSAR and molecular docking to drug discovery",
"author": "VILAR S.",
"doi-asserted-by": "crossref",
"first-page": "1555",
"issue": "18",
"journal-title": "Current Topics in Medicinal Chemistry",
"key": "ref51",
"volume": "8",
"year": "2008"
},
{
"DOI": "10.1016/j.drudis.2007.11.007",
"article-title": "Assessing drug-likeness – what are we missing?",
"author": "VISTOLI G.",
"doi-asserted-by": "crossref",
"first-page": "285",
"issue": "7-8",
"journal-title": "Drug Discovery Today",
"key": "ref52",
"volume": "13",
"year": "2008"
},
{
"DOI": "10.1016/S0169-409X(02)00003-0",
"article-title": "Prediction of ‘drug-likeness’",
"author": "WALTERS W.P.",
"doi-asserted-by": "crossref",
"first-page": "255",
"issue": "3",
"journal-title": "Advanced Drug Delivery Reviews",
"key": "ref53",
"volume": "54",
"year": "2002"
},
{
"DOI": "10.1128/JVI.02114-07",
"article-title": "Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design",
"author": "XUE X.",
"doi-asserted-by": "crossref",
"first-page": "2515",
"issue": "5",
"journal-title": "Journal of Virology",
"key": "ref54",
"volume": "82",
"year": "2008"
},
{
"DOI": "10.1126/science.abb3405",
"article-title": "Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors",
"author": "ZHANG L.",
"doi-asserted-by": "crossref",
"first-page": "409",
"issue": "6489",
"journal-title": "Science",
"key": "ref55",
"volume": "368",
"year": "2020"
}
],
"reference-count": 55,
"references-count": 55,
"relation": {},
"resource": {
"primary": {
"URL": "http://www.scielo.br/scielo.php?script=sci_arttext&pid=S1519-69842024000100270&tlng=en"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"General Agricultural and Biological Sciences"
],
"subtitle": [],
"title": "Inhibitory effect of thymoquinone from Nigella sativa against SARS-CoV-2 main protease. An in-silico study",
"type": "journal-article",
"volume": "84"
}
