Elucidation of remdesivir cytotoxicity pathways through genome-wide CRISPR-Cas9 screening and transcriptomics
et al., bioRxiv, doi:10.1101/2020.08.27.270819, Aug 2020
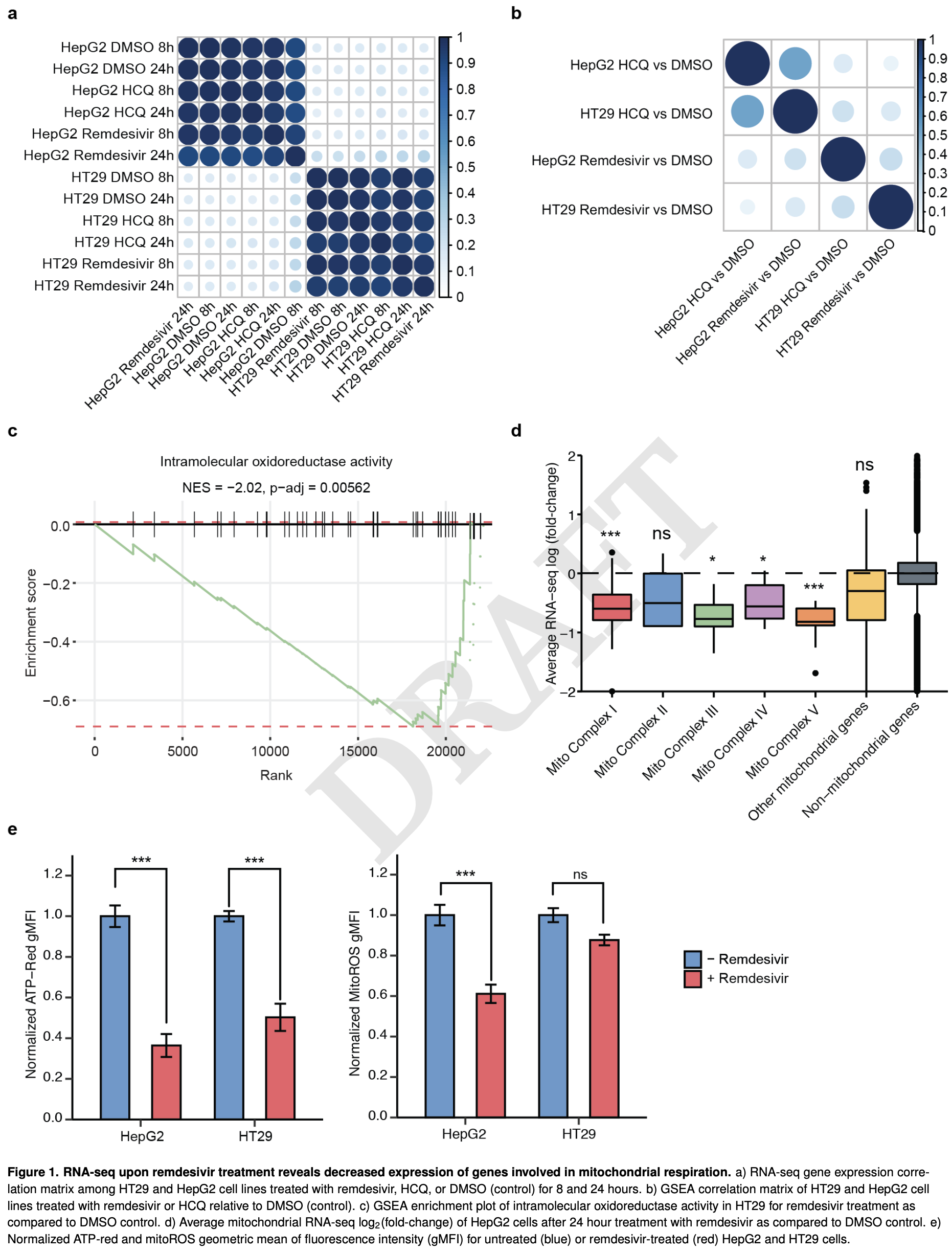
In vitro study analyzing the cytotoxicity pathways of remdesivir using genome-wide CRISPR-Cas9 screening and RNA sequencing. Remdesivir treatment significantly repressed nuclear genes encoding mitochondrial respiratory complexes, leading to decreased ATP production and mitochondrial oxidation.
Gérard, Zhou, Wu, Kamo, Choi, Kim show increased risk of acute kidney injury, Leo, Briciu, Muntean, Petrov, Arch show increased risk of liver injury, Negru, Cheng, Mohammed, Kwok, Zhu show increased risk of cardiac disorders, and Kwok, Merches, Akinci, Tang, Bagheri show increased risk of mitochondrial toxicity with remdesivir.
1.
Gérard et al., Remdesivir and Acute Renal Failure: A Potential Safety Signal From Disproportionality Analysis of the WHO Safety Database, Clinical Pharmacology & Therapeutics, doi:10.1002/cpt.2145.
2.
Zhou et al., Acute Kidney Injury and Drugs Prescribed for COVID-19 in Diabetes Patients: A Real-World Disproportionality Analysis, Frontiers in Pharmacology, doi:10.3389/fphar.2022.833679.
3.
Wu et al., Acute Kidney Injury Associated With Remdesivir: A Comprehensive Pharmacovigilance Analysis of COVID-19 Reports in FAERS, Frontiers in Pharmacology, doi:10.3389/fphar.2022.692828.
4.
Kamo et al., Association of Antiviral Drugs for the Treatment of COVID-19 With Acute Renal Failure, In Vivo, doi:10.21873/invivo.13637.
5.
Choi et al., Comparative effectiveness of combination therapy with nirmatrelvir–ritonavir and remdesivir versus monotherapy with remdesivir or nirmatrelvir–ritonavir in patients hospitalised with COVID-19: a target trial emulation study, The Lancet Infectious Diseases, doi:10.1016/S1473-3099(24)00353-0.
6.
Kim et al., Investigating the Safety Profile of Fast‐Track COVID‐19 Drugs Using the FDA Adverse Event Reporting System Database: A Comparative Observational Study, Pharmacoepidemiology and Drug Safety, doi:10.1002/pds.70043.
7.
Leo et al., Hepatocellular liver injury in hospitalized patients affected by COVID-19: Presence of different risk factors at different time points, Digestive and Liver Disease, doi:10.1016/j.dld.2021.12.014.
8.
Briciu et al., Evolving Clinical Manifestations and Outcomes in COVID-19 Patients: A Comparative Analysis of SARS-CoV-2 Variant Waves in a Romanian Hospital Setting, Pathogens, doi:10.3390/pathogens12121453.
9.
Muntean et al., Effects of COVID-19 on the Liver and Mortality in Patients with SARS-CoV-2 Pneumonia Caused by Delta and Non-Delta Variants: An Analysis in a Single Centre, Pharmaceuticals, doi:10.3390/ph17010003.
10.
Petrov et al., The Effect of Potentially Hepatotoxic Medicinal Products on Alanine Transaminase Levels in COVID-19 Patients: A Case–Control Study, Safety and Risk of Pharmacotherapy, doi:10.30895/2312-7821-2025-458.
11.
Arch et al., Evaluation of the effectiveness of remdesivir in treating severe COVID-19 using data from the ISARIC WHO Clinical Characterisation Protocol UK: a prospective, national cohort study, medRxiv, doi:10.1101/2021.06.18.21259072.
12.
Negru et al., Comparative Pharmacovigilance Analysis of Approved and Repurposed Antivirals for COVID-19: Insights from EudraVigilance Data, Biomedicines, doi:10.3390/biomedicines13061387.
13.
Cheng et al., Cardiovascular Safety of COVID-19 Treatments: A Disproportionality Analysis of Adverse Event Reports from the WHO VigiBase, Infectious Diseases and Therapy, doi:10.1007/s40121-025-01225-z.
14.
Mohammed et al., Bradycardia associated with remdesivir treatment in coronavirus disease 2019 patients: A propensity score-matched analysis, Medicine, doi:10.1097/MD.0000000000044501.
15.
Kwok et al., Remdesivir induces persistent mitochondrial and structural damage in human induced pluripotent stem cell-derived cardiomyocytes, Cardiovascular Research, doi:10.1093/cvr/cvab311.
16.
Zhu et al., Cardiovascular Risks of COVID-19 Therapeutics: Integrated Analysis of FAERS, Electronic Health Records, and Transcriptomics, Pharmaceuticals, doi:10.3390/ph19040574.
17.
Merches et al., The potential of remdesivir to affect function, metabolism and proliferation of cardiac and kidney cells in vitro, Archives of Toxicology, doi:10.1007/s00204-022-03306-1.
18.
Akinci et al., Elucidation of remdesivir cytotoxicity pathways through genome-wide CRISPR-Cas9 screening and transcriptomics, bioRxiv, doi:10.1101/2020.08.27.270819.
Akinci et al., 28 Aug 2020, USA, preprint, 20 authors.
Contact: rsherwood@rics.bwh.harvard.edu (corresponding author).
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
Abstract: ## Elucidation of remdesivir cytotoxicity pathways through genome-wide CRISPR-Cas9 screening and transcriptomics
Ersin Akinci 1,2,* , Minsun Cha 1,* , Lin Lin 3,* , Grace Yeo 4,5,* , Marisa C. Hamilton 1,* , Callie J. Donahue 6,# , Heysol C. Bermudez-Cabrera 1,# , Larissa C. Zanetti 1,7,# , Maggie Chen 1,8,# , Sammy A. Barkal 1,# , Benyapa Khowpinitchai 1,# , Nam Chu 1,9 , Minja Velimirovic 1,10 , Rikita Jodhani 1 , James D. Fife 1 , Miha Sovrovic 1 , Philip A. Cole 1,9 , Robert A. Davey 6 , Christopher A. Cassa 1 , and Richard I. Sherwood 1,3,
1 Division of Genetics, Department of Medicine, Brigham and Women's Hospital and Harvard Medical School, Boston, MA 02115 2 Department of Agricultural Biotechnology, Faculty of Agriculture, Akdeniz University, Antalya, 07070, Turkey 3
Hubrecht Institute, 3584 CT Utrecht, the Netherlands
4 Department of Electrical Engineering and Computer Science, Massachusetts Institute of Technology, Cambridge, MA 02139, USA 5
Computational and Systems Biology, Massachusetts Institute of Technology, Cambridge, MA 02139, USA
6 Department of Microbiology, National Emerging Infectious Disease Laboratories, Boston University Medical Campus, Boston, MA 02118, USA 7 Hospital Israelita Albert Einstein, São Paulo, SP 05652-900, Brazil
8 Department of Chemistry and Chemical Biology, Harvard University, Cambridge, MA 02138
9 Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115
Centre Hospitalier Universitaire de Québec Research Center-Université Laval, Québec, Québec G1V 4G2, Canada
10
The adenosine analogue remdesivir has emerged as a frontline antiviral treatment for SARS-CoV-2, with preliminary evidence that it reduces the duration and severity of illness 1 . Prior clinical studies have identified adverse events 1,2 , and remdesivir has been shown to inhibit mitochondrial RNA polymerase in biochemical experiments 7 , yet little is known about the specific genetic pathways involved in cellular remdesivir metabolism and cytotoxicity. Through genome-wide CRISPRCas9 screening and RNA sequencing, we show that remdesivir treatment leads to a repression of mitochondrial respiratory activity, and we identify five genes whose loss significantly reduces remdesivir cytotoxicity. In particular, we show that loss of the mitochondrial nucleoside transporter SLC29A3 mitigates remdesivir toxicity without a commensurate decrease in SARS-CoV-2 antiviral potency and that the mitochondrial adenylate kinase AK2 is a remdesivir kinase required for remdesivir efficacy and toxicity. This work elucidates the cellular mechanisms of remdesivir metabolism and provides a candidate gene target to reduce remdesivir cytotoxicity.
DRAFT Corresponding author: R.I.S. * These authors contributed equally to this work # These authors contributed equally to this work dosing or treatment earlier in disease progression as has been shown for other antiviral drugs such as oseltamivir (Tamiflu) 1,5,7,9-11 . One possible mode of toxicity induced by nucleoside analogues is mitochondrial toxicity, as mitochondrial polymerases lack the selectivity of mammalian polymerases to exclude nucleoside analogues. HIV antiviral nucleotide analogues and Hepatitis C virus (HCV) nucleoside analogs induce mitochondrial toxicity with varying levels of severity 12,13 , and remdesivir has been shown to inhibit mitochondrial RNA polymerase in biochemical experiments 14 , albeit at 100-fold lower rates than RdRp.
CRISPR..
DOI record:
{
"DOI": "10.1101/2020.08.27.270819",
"URL": "http://dx.doi.org/10.1101/2020.08.27.270819",
"abstract": "<jats:p>\n The adenosine analogue remdesivir has emerged as a frontline antiviral treatment for SARS-CoV-2, with preliminary evidence that it reduces the duration and severity of illness\n <jats:sup>1</jats:sup>\n . Prior clinical studies have identified adverse events\n <jats:sup>1,2</jats:sup>\n , and remdesivir has been shown to inhibit mitochondrial RNA polymerase in biochemical experiments\n <jats:sup>7</jats:sup>\n , yet little is known about the specific genetic pathways involved in cellular remdesivir metabolism and cytotoxicity. Through genome-wide CRISPR-Cas9 screening and RNA sequencing, we show that remdesivir treatment leads to a repression of mitochondrial respiratory activity, and we identify five genes whose loss significantly reduces remdesivir cytotoxicity. In particular, we show that loss of the mitochondrial nucleoside transporter\n <jats:italic>SLC29A3</jats:italic>\n mitigates remdesivir toxicity without a commensurate decrease in SARS-CoV-2 antiviral potency and that the mitochondrial adenylate kinase\n <jats:italic>AK2</jats:italic>\n is a remdesivir kinase required for remdesivir efficacy and toxicity. This work elucidates the cellular mechanisms of remdesivir metabolism and provides a candidate gene target to reduce remdesivir cytotoxicity.\n </jats:p>",
"accepted": {
"date-parts": [
[
2020,
8,
28
]
]
},
"author": [
{
"affiliation": [],
"family": "Akinci",
"given": "Ersin",
"sequence": "first"
},
{
"affiliation": [],
"family": "Cha",
"given": "Minsun",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Lin",
"given": "Lin",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Yeo",
"given": "Grace",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Hamilton",
"given": "Marisa C.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Donahue",
"given": "Callie J.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Bermudez-Cabrera",
"given": "Heysol C.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Zanetti",
"given": "Larissa C.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Chen",
"given": "Maggie",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Barkal",
"given": "Sammy A.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Khowpinitchai",
"given": "Benyapa",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Chu",
"given": "Nam",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Velimirovic",
"given": "Minja",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Jodhani",
"given": "Rikita",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Fife",
"given": "James D.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Sovrovic",
"given": "Miha",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Cole",
"given": "Philip A.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Davey",
"given": "Robert A.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Cassa",
"given": "Christopher A.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Sherwood",
"given": "Richard I.",
"sequence": "additional"
}
],
"container-title": [],
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2020,
8,
29
]
],
"date-time": "2020-08-29T00:10:42Z",
"timestamp": 1598659842000
},
"deposited": {
"date-parts": [
[
2026,
1,
15
]
],
"date-time": "2026-01-15T00:46:55Z",
"timestamp": 1768438015000
},
"group-title": "Genetics",
"indexed": {
"date-parts": [
[
2026,
4,
12
]
],
"date-time": "2026-04-12T13:59:07Z",
"timestamp": 1776002347170,
"version": "3.50.1"
},
"institution": [
{
"name": "bioRxiv"
}
],
"is-referenced-by-count": 19,
"issued": {
"date-parts": [
[
2020,
8,
28
]
]
},
"license": [
{
"URL": "http://creativecommons.org/licenses/by-nc-nd/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2020,
8,
28
]
],
"date-time": "2020-08-28T00:00:00Z",
"timestamp": 1598572800000
}
}
],
"link": [
{
"URL": "https://syndication.highwire.org/content/doi/10.1101/2020.08.27.270819",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "54368",
"original-title": [],
"posted": {
"date-parts": [
[
2020,
8,
28
]
]
},
"prefix": "10.64898",
"published": {
"date-parts": [
[
2020,
8,
28
]
]
},
"publisher": "openRxiv",
"reference": [
{
"key": "2024080413273807000_2020.08.27.270819v1.1",
"unstructured": "Beigel, J. H. et al. Remdesivir for the Treatment of Covid-19 — Preliminary Report. N. Engl. J. Med. 0, null (2020)."
},
{
"DOI": "10.1021/acscentsci.0c00489",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.2"
},
{
"DOI": "10.1038/nature17180",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.3"
},
{
"DOI": "10.1128/mBio.00221-18",
"doi-asserted-by": "crossref",
"key": "2024080413273807000_2020.08.27.270819v1.4",
"unstructured": "Agostini, M. L. et al. Coronavirus Susceptibility to the Antiviral Remdesivir (GS-5734) Is Mediated by the Viral Polymerase and the Proofreading Exoribonuclease. mBio 9, (2018)."
},
{
"DOI": "10.1073/pnas.1922083117",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.5"
},
{
"DOI": "10.1038/s41422-020-0282-0",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.6"
},
{
"DOI": "10.1038/s41586-020-2423-5",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.7"
},
{
"DOI": "10.1016/j.celrep.2020.107940",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.8"
},
{
"key": "2024080413273807000_2020.08.27.270819v1.9",
"unstructured": "Scientists to Stop COVID-19."
},
{
"DOI": "10.1056/NEJMc2022236",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.10"
},
{
"key": "2024080413273807000_2020.08.27.270819v1.11",
"unstructured": "Organization, W. H. WHO guidelines for pharmacological management of pandemic (H1N1) 2009 influenza and other influenza viruses. (2009)."
},
{
"DOI": "10.3389/fmolb.2017.00074",
"doi-asserted-by": "crossref",
"key": "2024080413273807000_2020.08.27.270819v1.12",
"unstructured": "Young, M. J. Off-Target Effects of Drugs that Disrupt Human Mitochondrial DNA Maintenance. Front. Mol. Biosci. 4, (2017)."
},
{
"DOI": "10.1128/AAC.01922-15",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.13"
},
{
"DOI": "10.3390/v11040326",
"doi-asserted-by": "crossref",
"key": "2024080413273807000_2020.08.27.270819v1.14",
"unstructured": "Tchesnokov, E. P. , Feng, J. Y. , Porter, D. P. & Götte, M. Mechanism of Inhibition of Ebola Virus RNA-Dependent RNA Polymerase by Remdesivir. Viruses 11, (2019)."
},
{
"DOI": "10.1038/nrd.2017.111",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.15"
},
{
"DOI": "10.1016/j.cell.2017.10.049",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.16"
},
{
"DOI": "10.1038/nrg3899",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.17"
},
{
"DOI": "10.1093/cid/ciaa237",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.18"
},
{
"DOI": "10.1038/sj.onc.1206622",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.19"
},
{
"DOI": "10.1073/pnas.0506580102",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.20"
},
{
"DOI": "10.1002/anie.201510003",
"article-title": "A Multisite-Binding Switchable Fluorescent Probe for Monitoring Mitochondrial ATP Level Fluctuation in Live Cells",
"doi-asserted-by": "crossref",
"first-page": "1773",
"journal-title": "Angew. Chem. Int. Ed Engl",
"key": "2024080413273807000_2020.08.27.270819v1.21",
"volume": "55",
"year": "2016"
},
{
"DOI": "10.3389/fgene.2019.00131",
"doi-asserted-by": "crossref",
"key": "2024080413273807000_2020.08.27.270819v1.22",
"unstructured": "Iannetti, E. F. et al. Live-Imaging Readouts and Cell Models for Phenotypic Profiling of Mitochondrial Function. Front. Genet. 10, (2019)."
},
{
"DOI": "10.1016/j.tem.2017.07.003",
"article-title": "Surviving Stress: Modulation of ATF4-Mediated Stress Responses in Normal and Malignant Cells",
"doi-asserted-by": "crossref",
"first-page": "794",
"journal-title": "Trends Endocrinol. Metab. TEM",
"key": "2024080413273807000_2020.08.27.270819v1.23",
"volume": "28",
"year": "2017"
},
{
"DOI": "10.1016/j.celrep.2017.04.042",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.24"
},
{
"DOI": "10.1083/jcb.201702058",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.25"
},
{
"DOI": "10.1038/s41418-019-0387-5",
"article-title": "The antimalarial drug amodiaquine stabilizes p53 through ribosome biogenesis stress, independently of its autophagy-inhibitory activity",
"doi-asserted-by": "crossref",
"first-page": "773",
"journal-title": "Cell Death Differ",
"key": "2024080413273807000_2020.08.27.270819v1.26",
"volume": "27",
"year": "2020"
},
{
"DOI": "10.1016/j.jmb.2018.08.019",
"article-title": "Ursodeoxycholic Acid Improves Mitochondrial Function and Redistributes Drp1 in Fibroblasts from Patients with Either Sporadic or Familial Alzheimer’s Disease",
"doi-asserted-by": "crossref",
"first-page": "3942",
"journal-title": "J. Mol. Biol",
"key": "2024080413273807000_2020.08.27.270819v1.27",
"volume": "430",
"year": "2018"
},
{
"DOI": "10.1093/bja/aev400",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.28"
},
{
"DOI": "10.1016/j.celrep.2018.11.043",
"article-title": "Dual Inhibition of the Lactate Transporters MCT1 and MCT4 Is Synthetic Lethal with Metformin due to NAD+ Depletion in Cancer Cells",
"doi-asserted-by": "crossref",
"first-page": "3047",
"journal-title": "Cell Rep",
"key": "2024080413273807000_2020.08.27.270819v1.29",
"volume": "25",
"year": "2018"
},
{
"DOI": "10.1038/nbt.3437",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.30"
},
{
"DOI": "10.1038/s41596-018-0113-7",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.31"
},
{
"DOI": "10.1186/s13059-014-0554-4",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.32"
},
{
"DOI": "10.1021/acs.jmedchem.7b00734",
"article-title": "The ProTide Prodrug Technology: From the Concept to the Clinic",
"doi-asserted-by": "crossref",
"first-page": "2211",
"journal-title": "J. Med. Chem",
"key": "2024080413273807000_2020.08.27.270819v1.33",
"volume": "61",
"year": "2018"
},
{
"DOI": "10.1152/physrev.00023.2017",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.34"
},
{
"DOI": "10.1038/ng.278",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.35"
},
{
"DOI": "10.1038/ng.265",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.36"
},
{
"DOI": "10.1152/ajpgi.90672.2008",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.37"
},
{
"DOI": "10.1016/j.ebiom.2015.07.008",
"article-title": "Discovery of Genetic Variants of the Kinases That Activate Tenofovir in a Compartment-specific Manner",
"doi-asserted-by": "crossref",
"first-page": "1145",
"journal-title": "EBioMedicine",
"key": "2024080413273807000_2020.08.27.270819v1.38",
"volume": "2",
"year": "2015"
},
{
"DOI": "10.1128/AAC.39.10.2304",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.39"
},
{
"DOI": "10.1126/scitranslmed.abb5883",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.40"
},
{
"DOI": "10.1128/AAC.01719-19",
"doi-asserted-by": "crossref",
"key": "2024080413273807000_2020.08.27.270819v1.41",
"unstructured": "Sticher, Z. M. et al. Analysis of the Potential for N4-Hydroxycytidine To Inhibit Mitochondrial Replication and Function. Antimicrob. Agents Chemother. 64, (2020)."
},
{
"DOI": "10.15252/embj.201593118",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.42"
},
{
"DOI": "10.1016/j.cell.2017.02.004",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.43"
},
{
"DOI": "10.1007/s13238-017-0423-5",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.44"
},
{
"DOI": "10.1038/nrm3700",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.45"
},
{
"DOI": "10.1016/j.ejmech.2004.11.015",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.46"
},
{
"DOI": "10.1074/jbc.M110.161802",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.47"
},
{
"DOI": "10.1074/jbc.270.38.22105",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.48"
},
{
"DOI": "10.1093/oxfordjournals.jbchem.a021915",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.49"
},
{
"DOI": "10.1371/journal.pone.0195764",
"article-title": "Discovery of genetic variants of the kinases that activate tenofovir among individuals in the United States, Thailand, and South Africa: HPTN067",
"doi-asserted-by": "crossref",
"first-page": "e0195764",
"journal-title": "PLOS ONE",
"key": "2024080413273807000_2020.08.27.270819v1.50",
"volume": "13",
"year": "2018"
},
{
"DOI": "10.1080/15257770.2016.1210805",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.51"
},
{
"DOI": "10.1016/j.ccr.2012.05.016",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.52"
},
{
"DOI": "10.1039/C2TX20051A",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.53"
},
{
"DOI": "10.1016/S0140-6736(20)31022-9",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.54"
},
{
"DOI": "10.1056/NEJMoa1910993",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.55"
},
{
"DOI": "10.1371/journal.pone.0185056",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.56"
},
{
"article-title": "STAR: ultrafast universal RNA-seq aligner",
"first-page": "15",
"journal-title": "Bioinforma. Oxf. Engl",
"key": "2024080413273807000_2020.08.27.270819v1.57",
"volume": "29",
"year": "2013"
},
{
"article-title": "HTSeq–a Python framework to work with high-throughput sequencing data",
"first-page": "166",
"journal-title": "Bioinforma. Oxf. Engl",
"key": "2024080413273807000_2020.08.27.270819v1.58",
"volume": "31",
"year": "2015"
},
{
"DOI": "10.1038/s41596-018-0103-9",
"article-title": "Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap",
"doi-asserted-by": "crossref",
"first-page": "482",
"journal-title": "Nat. Protoc",
"key": "2024080413273807000_2020.08.27.270819v1.59",
"volume": "14",
"year": "2019"
},
{
"DOI": "10.1093/nar/gky1055",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.60"
},
{
"DOI": "10.1038/nmeth.3047",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.61"
},
{
"DOI": "10.1016/j.cell.2013.12.001",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.62"
},
{
"DOI": "10.1074/jbc.270.38.22105",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.63"
},
{
"DOI": "10.1093/oxfordjournals.jbchem.a021915",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.64"
},
{
"DOI": "10.1093/nar/gkw784",
"doi-asserted-by": "publisher",
"key": "2024080413273807000_2020.08.27.270819v1.65"
},
{
"article-title": "Improved structure, function and compatibility for CellProfiler: modular high-throughput image analysis software",
"first-page": "1179",
"journal-title": "Bioinforma. Oxf. Engl",
"key": "2024080413273807000_2020.08.27.270819v1.66",
"volume": "27",
"year": "2011"
}
],
"reference-count": 66,
"references-count": 66,
"relation": {
"has-review": [
{
"asserted-by": "object",
"id": "10.1162/2e3983f5.83dfeb61",
"id-type": "doi"
},
{
"asserted-by": "object",
"id": "10.1162/2e3983f5.8bc2d5fa",
"id-type": "doi"
}
]
},
"resource": {
"primary": {
"URL": "http://biorxiv.org/lookup/doi/10.1101/2020.08.27.270819"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"subtype": "preprint",
"title": "Elucidation of remdesivir cytotoxicity pathways through genome-wide CRISPR-Cas9 screening and transcriptomics",
"type": "posted-content"
}