Molecular Mechanisms Behind Anti SARS-CoV-2 Action of Lactoferrin
et al., Frontiers in Molecular Biosciences, doi:10.3389/fmolb.2021.607443, Feb 2021
In silico study of possible molecular mechanisms for anti-SARS-CoV-2 action of lactoferrin.
18 preclinical studies support the efficacy of lactoferrin for COVID-19:
1.
da Silva et al., Immunomodulatory effect of bovine lactoferrin during SARS-CoV-2 infection, Frontiers in Immunology, doi:10.3389/fimmu.2024.1456634.
2.
Cutone et al., Lactoferrin binding to Sars-CoV-2 Spike glycoprotein protects host from infection, inflammation and iron dysregulation., Research Square, doi:10.21203/rs.3.rs-1605740/v1.
3.
Miotto et al., Molecular Mechanisms Behind Anti SARS-CoV-2 Action of Lactoferrin, Frontiers in Molecular Biosciences, doi:10.3389/fmolb.2021.607443.
4.
Babulic et al., Lactoferrin Binds through Its N-Terminus to the Receptor-Binding Domain of the SARS-CoV-2 Spike Protein, Pharmaceuticals, doi:10.3390/ph17081021.
5.
Yathindranath et al., Lipid Nanoparticle-Based Inhibitors for SARS-CoV-2 Host Cell Infection, International Journal of Nanomedicine, doi:10.2147/IJN.S448005.
6.
Alves et al., Inhibition of SARS-CoV-2 Infection in Vero Cells by Bovine Lactoferrin under Different Iron-Saturation States, Pharmaceuticals, doi:10.3390/ph16101352.
7.
Kobayashi-Sakamoto et al., Bovine lactoferrin suppresses the cathepsin-dependent pathway of SARS-CoV-2 entry in vitro, International Dairy Journal, doi:10.1016/j.idairyj.2023.105805.
8.
Andreu et al., Liposomal Lactoferrin Exerts Antiviral Activity against HCoV-229E and SARS-CoV-2 Pseudoviruses In Vitro, Viruses, doi:10.3390/v15040972.
9.
Yazawa et al., Evaluation of SARS-CoV-2 isolation in cell culture from nasal/nasopharyngeal swabs or saliva specimens of patients with COVID-19, Research Square, doi:10.21203/rs.3.rs-2676422/v1.
10.
Piacentini et al., Lactoferrin Inhibition of the Complex Formation between ACE2 Receptor and SARS CoV-2 Recognition Binding Domain, International Journal of Molecular Sciences, doi:10.3390/ijms23105436.
11.
Ostrov et al., Highly Specific Sigma Receptor Ligands Exhibit Anti-Viral Properties in SARS-CoV-2 Infected Cells, Pathogens, doi:10.3390/pathogens10111514.
12.
Mirabelli et al., Morphological cell profiling of SARS-CoV-2 infection identifies drug repurposing candidates for COVID-19, Proceedings of the National Academy of Sciences, doi:10.1073/pnas.2105815118.
13.
Salaris et al., Protective Effects of Lactoferrin against SARS-CoV-2 Infection In Vitro, Nutrients, doi:10.3390/nu13020328.
Miotto et al., 15 Feb 2021, peer-reviewed, 6 authors.
Contact: mattia.miotto@roma1.infn.it.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
Molecular Mechanisms Behind Anti SARS-CoV-2 Action of Lactoferrin
Frontiers in Molecular Biosciences, doi:10.3389/fmolb.2021.607443
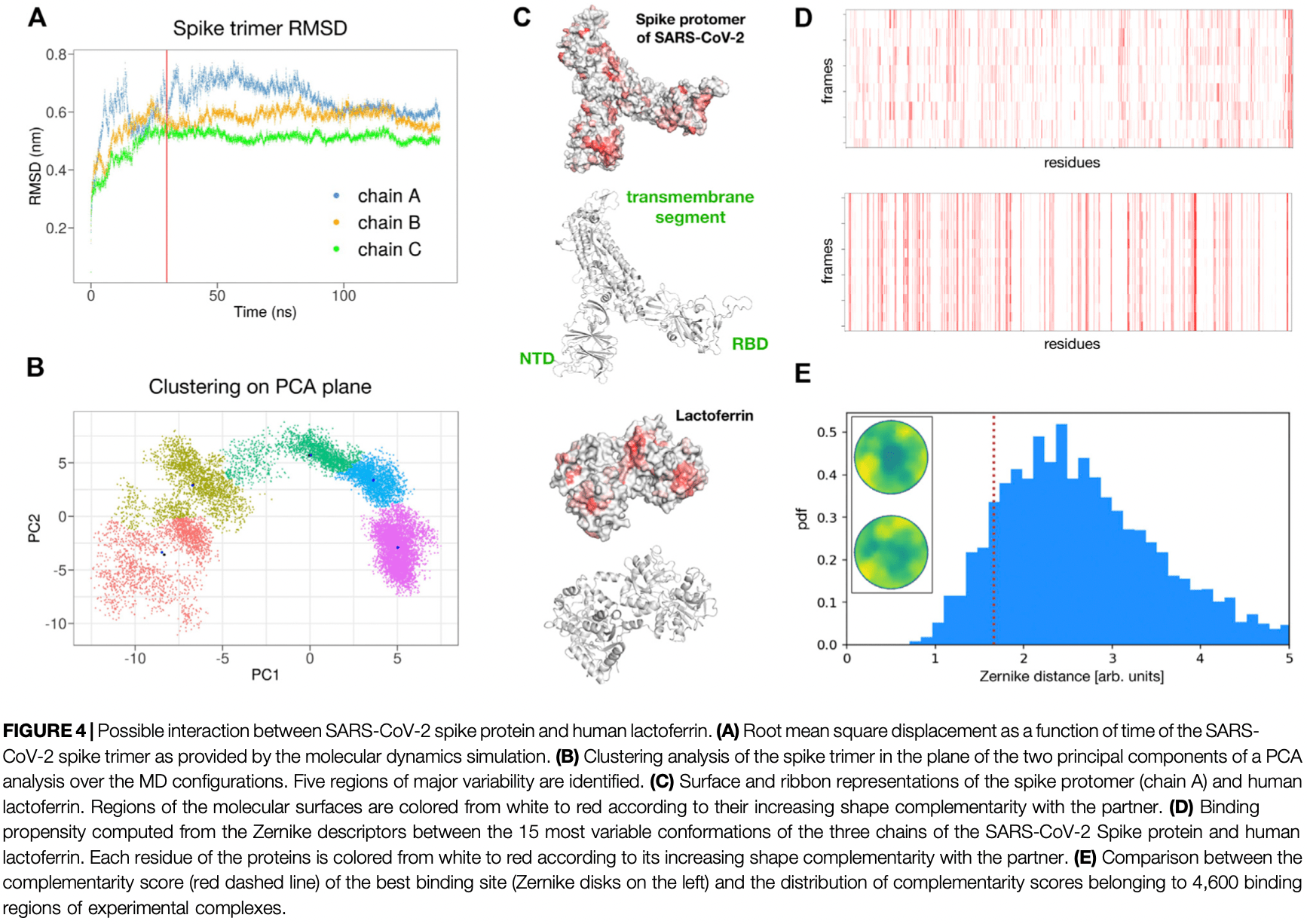
Despite the huge effort to contain the infection, the novel SARS-CoV-2 coronavirus has rapidly become pandemic, mainly due to its extremely high human-to-human transmission capability, and a surprisingly high viral charge of symptom-less people. While the seek for a vaccine is still ongoing, promising results have been obtained with antiviral compounds. In particular, lactoferrin is regarded to have beneficial effects both in preventing and soothing the infection. Here, we explore the possible molecular mechanisms with which lactoferrin interferes with SARS-CoV-2 cell invasion, preventing attachment and/or entry of the virus. To this aim, we search for possible interactions lactoferrin may have with virus structural proteins and host receptors. Representing the molecular iso-electron surface of proteins in terms of 2D-Zernike descriptors, we 1) identified putative regions on the lactoferrin surface able to bind sialic acid present on the host cell membrane, sheltering the cell from the virus attachment; 2) showed that no significant shape complementarity is present between lactoferrin and the ACE2 receptor, while 3) two high complementarity regions are found on the N-and C-terminal domains of the SARS-CoV-2 spike protein, hinting at a possible competition between lactoferrin and ACE2 for the binding to the spike protein.
AUTHOR CONTRIBUTIONS EM, AB, and GR conceived research; MM and LB performed molecular dynamics simulations. MM, LR, and EM performed calculations and computational analysis. All authors analyzed results; all authors wrote and revised the paper.
Conflict of Interest: The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. Copyright © 2021 Miotto, Di Rienzo, Bò, Boffi, Ruocco and Milanetti. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
References
Andersen, Osbakk, Vorland, Traavik, Gutteberg, Lactoferrin and cyclic lactoferricin inhibit the entry of human cytomegalovirus into human fibroblasts, Antiviral. Res, doi:10.1016/s0166-3542(01)00146-2
Baker, Baker, Lactoferrin, Cell. Mol. Life Sci, doi:10.1007/s00018-005-5368-9
Berlutti, Pantanella, Natalizi, Frioni, Paesano et al., Antiviral properties of lactoferrin--a natural immunity molecule, Molecules, doi:10.3390/molecules16086992
Berman, Bourne, Westbrook, Zardecki, The protein data bank
Bianchi, Benvenuto, Giovanetti, Angeletti, Ciccozzi et al., Sars-CoV-2 envelope and membrane proteins: structural differences linked to virus characteristics?, BioMed. Res. Int, doi:10.1155/2020/4389089
Bishop, Gagneux, Evolution of carbohydrate antigens--microbial forces shaping host glycomes?, Glycobiology, doi:10.1093/glycob/cwm005
Brooks, Brooks, Mackerell, Nilsson, Petrella et al., CHARMM: the biomolecular simulation program, J. Comput. Chem, doi:10.1002/jcc.21287
Bussi, Donadio, Parrinello, Canonical sampling through velocity rescaling, J. Chem. Phys, doi:10.1063/1.2408420
Caccavo, Pellegrino, Altamura, Rigon, Amati et al., Antimicrobial and immunoregulatory functions of lactoferrin and its potential therapeutic application, J. Endotoxin. Res, doi:10.1179/096805102125001000
Campione, Cosio, Rosa, Lanna, Di Girolamo et al., Lactoferrin as protective natural barrier of respiratory and intestinal mucosa against coronavirus infection and inflammation, Ijms, doi:10.3390/ijms21144903
Casalino, Gaieb, Goldsmith, Hjorth, Dommer et al., Beyond shielding: The roles of glycans in the sars-cov-2 spike protein, ACS Cent. Sci, doi:10.1021/acscentsci.0c01056
Chang, Ng, Sun, Lactoferrin as potential preventative and adjunct treatment for COVID-19, Int. J. Antimicrob. Agents, doi:10.1016/j.ijantimicag.2020.106118
Cheatham, Miller, Fox, Darden, Kollman, Molecular dynamics simulations on solvated biomolecular systems: The particle mesh ewald method leads to stable trajectories of DNA, RNA, and proteins, J. Am. Chem. Soc, doi:10.1021/ja00119a045
Chen, Li, Weng, ZDOCK: an initial-stage protein-docking algorithm, Proteins, doi:10.1002/prot.10389
Daberdaku, Ferrari, Antibody interface prediction with 3d zernike descriptors and svm, Bioinformatics, doi:10.1093/bioinformatics/bty918
De Carvalho, Da Rocha Matos, Caetano, De Sousa Junior, Da Costa Campos et al., In vitro inhibition of SARS-CoV-2 infection by bovine lactoferrin, bioRxiv, doi:10.1101/2020.05.13.093781
Di Rienzo, Milanetti, Alba, 'abramo, Quantitative characterization of binding pockets and binding complementarity by means of zernike descriptors, J Chem. Inf. Model, doi:10.1021/acs.jcim.9b01066
Di Rienzo, Milanetti, Lepore, Olimpieri, Tramontano, Superposition-free comparison and clustering of antibody binding sites: implications for the prediction of the nature of their antigen, Sci. Rep, doi:10.1038/srep45053
Drobni, Näslund, Evander, Lactoferrin inhibits human papillomavirus binding and uptake in vitro, Antiviral. Res, doi:10.1016/j.antiviral.2004.05.005
Drosten, Günther, Preiser, Van Der Werf, Brodt et al., Identification of a novel coronavirus in patients with severe acute respiratory syndrome, N. Engl. J. Med, doi:10.1056/NEJMoa030747
Gainza, Sverrisson, Monti, Rodolà, Boscaini et al., Deciphering interaction fingerprints from protein molecular surfaces using geometric deep learning, Nat. Methods, doi:10.1038/s41592-019-0666-6
Giansanti, Panella, Leboffe, Antonini, Lactoferrin from milk: Nutraceutical and pharmacological properties, Pharmaceuticals, doi:10.3390/ph9040061
Graham, Baric, Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission, J. Virol, doi:10.1128/JVI.01394-09
Groves, The isolation of a red protein from milk2, J. Am. Chem. Soc, doi:10.1021/ja01498a029
Harmsen, Swart, De Béthune, De Clercq, The, Antiviral effects of plasma and milk proteins: Lactoferrin shows potent activity against both human immunodeficiency virus and human cytomegalovirus replication in vitro, J. Infect. Dis, doi:10.1093/infdis/172.2.380
Hess, Bekker, Berendsen, Fraaije, LINCS: a linear constraint solver for molecular simulations, J. Comput. Chem, doi:10.1002/(sici)1096-987x(199709)18:12<1463::aid-jcc4>3.0.co;2-h
Huang, Wang, Li, Ren, Zhao et al., Clinical features of patients infected with 2019 novel coronavirus in wuhan, china, Lancet, doi:10.1016/s0140-6736(20)30183-5
Hulswit, Lang, Bakkers, Li, Li et al., Human coronaviruses oc43 and hku1 bind to 9-o-acetylated sialic acids via a conserved receptor-binding site in spike protein domain a, Proc. Natl. Acad. Sci, doi:10.1073/pnas.1809667116
Jorgensen, Chandrasekhar, Madura, Impey, Klein, Comparison of simple potential functions for simulating liquid water, J. Chem. Phys, doi:10.1063/1.445869
Kihara, Sael, Chikhi, Esquivel-Rodriguez, Molecular surface representation using 3d zernike descriptors for protein shape comparison and docking, Curr. Protein Pept. Sci, doi:10.2174/138920311796957612
Ksiazek, Erdman, Goldsmith, Zaki, Peret et al., A novel coronavirus associated with severe acute respiratory syndrome, N. Engl. J. Med, doi:10.1056/NEJMoa030781
Kuo, Godeke, Raamsman, Masters, Rottier, Retargeting of coronavirus by substitution of the spike glycoprotein ectodomain: crossing the host cell species barrier, J. Virol, doi:10.1128/jvi.74.3.1393-1406.2000
Lang, Yang, Deng, Liu, Yang et al., Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans, PLoS One, doi:10.1371/journal.pone.0023710
Langford-Smith, Day, Bishop, Clark, Complementing the sugar code: Role of GAGs and sialic acid in complement regulation, Front Immunol, doi:10.3389/fimmu.2015.00025
Li, Li, Farzan, Harrison, Structure of sars coronavirus spike receptor-binding domain complexed with receptor, Science, doi:10.1126/science.1116480
Li, Structural analysis of major species barriers between humans and palm civets for severe acute respiratory syndrome coronavirus infections, J. Virol, doi:10.1128/JVI.00442-08
Li, Zhang, Sui, Kuhn, Moore et al., Receptor and viral determinants of sars-coronavirus adaptation to human ace2, EMBO. J, doi:10.1038/sj.emboj.7600640
Liu, Chopra, Li, Wolfert, Tompkins et al., SARS-CoV-2 spike protein binds heparan sulfate in a length-and sequencedependent manner, bioRxiv, doi:10.1101/2020.05.10.087288
Marchetti, Trybala, Superti, Johansson, Bergström, Inhibition of herpes simplex virus infection by lactoferrin is dependent on interference with the virus binding to glycosaminoglycans, Virology, doi:10.1016/j.virol.2003.09.029
Mccann, Lee, Wan, Roginski, Coventry, The effect of bovine lactoferrin and lactoferricin b on the ability of feline calicivirus (a norovirus surrogate) and poliovirus to infect cell cultures, J. Appl. Microbiol, doi:10.1046/j.1365-2672.2003.02071.x
Milanetti, Miotto, Di Rienzo, Monti, Gosti et al., In-silico evidence for two receptors based strategy of sars-cov-2, doi:10.1101/2020.03.24.006197
Milanetti, Miotto, Rienzo, Monti, Gosti et al., 2d zernike polynomial expansion: finding the protein-protein binding regions, Comput. Struct. Biotechnol. J, doi:10.1016/j.csbj.2020.11.051
Miotto, Rienzo, Gosti, Milanetti, Ruocco, Does blood type affect the covid-19 infection pattern?
Niaz, Saeed, Ahmed, Imran, Maan et al., Lactoferrin (LF): a natural antimicrobial protein, International Journal of Food Properties, doi:10.1080/10942912.2019.1666137
Park, Walls, Wang, Sauer, Li et al., Structures of MERS-CoV spike glycoprotein in complex with sialoside attachment receptors, Nat. Struct. Mol. Biol, doi:10.1038/s41594-019-0334-7
Parrinello, Rahman, Crystal structure and pair potentials: A molecular-dynamics study, Phys. Rev. Lett, doi:10.1103/physrevlett.45.1196
Prompetchara, Ketloy, Palaga, Immune responses in covid-19 and potential vaccines: Lessons learned from sars and mers epidemic. Asian Pac, J. Allergy Immunol, doi:10.12932/AP-200220-0772
Puddu, Borghi, Gessani, Valenti, Belardelli et al., Antiviral effect of bovine lactoferrin saturated with metal ions on early steps of human immunodeficiency virus type 1 infection, Int. J. Biochem. Cell Biol, doi:10.1016/s1357-2725(98)00066-1
Raman, Tharakaraman, Sasisekharan, Sasisekharan, Glycan-protein interactions in viral pathogenesis, Curr. Opin. Struct. Biol, doi:10.1016/j.sbi.2016.10.003
Redwan, Uversky, El-Fakharany, Al-Mehdar, Potential lactoferrin activity against pathogenic viruses, C R Biol, doi:10.1016/j.crvi.2014.08.003
Richards, Areas, volumes, packing and protein structure, Annu. Rev. Biophys. Bioeng, doi:10.1146/annurev.bb.06.060177.001055
Robson, Bioinformatics studies on a function of the sars-cov-2 spike glycoprotein as the binding of host sialic acid glycans, Comput. Biol. Med, doi:10.1016/j.compbiomed.2020.103849
Roy, Kucukural, Zhang, I-TASSER: a unified platform for automated protein structure and function prediction, Nat. Protoc, doi:10.1038/nprot.2010.5
Schwegmann-Weßels, Herrler, Sialic acids as receptor determinants for coronaviruses, Glycoconj J, doi:10.1007/s10719-006-5437-9
Seah, Su, Lingam, Revisiting the dangers of the coronavirus in the ophthalmology practice, Eye (Lond), doi:10.1038/s41433-020-0790-7
Senior, Evans, Jumper, Kirkpatrick, Sifre et al., Improved protein structure prediction using potentials from deep learning, Nature, doi:10.1038/s41586-019-1923-7
Sorensen, Sorensen, The proteins in whey. Compte rendu des Travaux du Laboratoire de Carlsberg, Ser. Chim, doi:10.3168/jds.s0022-0302(40)95543-6
Su, Wong, Shi, Liu, Lai et al., Epidemiology, genetic recombination, and pathogenesis of coronaviruses, Trends Microbiol, doi:10.1016/j.tim.2016.03.003
Superti, Siciliano, Rega, Giansanti, Valenti et al., Involvement of bovine lactoferrin metal saturation, sialic acid and protein fragments in the inhibition of rotavirus infection, Biochim. Biophys. Acta, doi:10.1016/s0304-4165(01)00178-7
Thomas, The structure of the membrane protein of SARS-CoV-2 resembles the sugar transporter SemiSWEET, Pathog Immun, doi:10.20411/pai.v5i1.377
Tortorici, Walls, Lang, Wang, Li et al., Structural basis for human coronavirus attachment to sialic acid receptors, Nat. Struct. Mol. Biol, doi:10.1016/s0166-3542(01)00195-4
Van Der Spoel, Lindahl, Hess, Groenhof, Mark et al., GROMACS: fast, flexible, and free, J. Comput. Chem, doi:10.1002/jcc.20291
Vandelli, Monti, Milanetti, Ponti, Tartaglia, Structural analysis of sars-cov-2 and prediction of the human interactome, Nucleic Acids Res, doi:10.1093/nar/gkaa864
Varki, Biological roles of glycans, Glycobiology, doi:10.1093/glycob/cww086
Venkatraman, Yang, Sael, Kihara, Protein-protein docking using region-based 3d zernike descriptors, BMC bioinformatics, doi:10.1186/1471-2105-10-407
Waarts, Aneke, Smit, Kimata, Bittman et al., Antiviral activity of human lactoferrin: inhibition of alphavirus interaction with heparan sulfate, Virology, doi:10.1016/j.virol.2005.01.010
Yan, Zhang, Li, Xia, Guo et al., Structural basis for the recognition of the sars-cov-2 by full-length human ace2, Science, doi:10.1126/science.abb2762
Yi, Kaneko, Yu, Murakami, Hepatitis C virus envelope proteins bind lactoferrin, J. Virol, doi:10.1128/JVI.71.8.5997-6002.1997
Zaki, Van Boheemen, Bestebroer, Osterhaus, Fouchier et al., Isolation of a novel coronavirus from a man with pneumonia in saudi arabia, N Engl J Med, doi:10.1056/NEJMoa1211721
Zhang, Skolnick, Tm-align: a protein structure alignment algorithm based on the tm-score, Nucleic Acids Res, doi:10.1093/nar/gki524
Zhou, Yang, Wang, Hu, Zhang et al., A pneumonia outbreak associated with a new coronavirus of probable bat origin, Nature, doi:10.1038/s41586-020-2012-7
Zhu, Zhang, Wang, Li, Yang et al., A novel coronavirus from patients with pneumonia in china, N. Engl. J. Med, doi:10.1056/NEJMoa2001017
DOI record:
{
"DOI": "10.3389/fmolb.2021.607443",
"ISSN": [
"2296-889X"
],
"URL": "http://dx.doi.org/10.3389/fmolb.2021.607443",
"abstract": "<jats:p>Despite the huge effort to contain the infection, the novel SARS-CoV-2 coronavirus has rapidly become pandemic, mainly due to its extremely high human-to-human transmission capability, and a surprisingly high viral charge of symptom-less people. While the seek for a vaccine is still ongoing, promising results have been obtained with antiviral compounds. In particular, lactoferrin is regarded to have beneficial effects both in preventing and soothing the infection. Here, we explore the possible molecular mechanisms with which lactoferrin interferes with SARS-CoV-2 cell invasion, preventing attachment and/or entry of the virus. To this aim, we search for possible interactions lactoferrin may have with virus structural proteins and host receptors. Representing the molecular iso-electron surface of proteins in terms of 2D-Zernike descriptors, we 1) identified putative regions on the lactoferrin surface able to bind sialic acid present on the host cell membrane, sheltering the cell from the virus attachment; 2) showed that no significant shape complementarity is present between lactoferrin and the ACE2 receptor, while 3) two high complementarity regions are found on the N- and C-terminal domains of the SARS-CoV-2 spike protein, hinting at a possible competition between lactoferrin and ACE2 for the binding to the spike protein.</jats:p>",
"alternative-id": [
"10.3389/fmolb.2021.607443"
],
"author": [
{
"affiliation": [],
"family": "Miotto",
"given": "Mattia",
"sequence": "first"
},
{
"affiliation": [],
"family": "Di Rienzo",
"given": "Lorenzo",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Bò",
"given": "Leonardo",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Boffi",
"given": "Alberto",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Ruocco",
"given": "Giancarlo",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Milanetti",
"given": "Edoardo",
"sequence": "additional"
}
],
"container-title": "Frontiers in Molecular Biosciences",
"container-title-short": "Front. Mol. Biosci.",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"frontiersin.org"
]
},
"created": {
"date-parts": [
[
2021,
2,
15
]
],
"date-time": "2021-02-15T09:27:34Z",
"timestamp": 1613381254000
},
"deposited": {
"date-parts": [
[
2021,
2,
15
]
],
"date-time": "2021-02-15T09:27:37Z",
"timestamp": 1613381257000
},
"funder": [
{
"DOI": "10.13039/501100009531",
"doi-asserted-by": "publisher",
"name": "Istituto Italiano di Tecnologia"
}
],
"indexed": {
"date-parts": [
[
2022,
8,
3
]
],
"date-time": "2022-08-03T11:52:43Z",
"timestamp": 1659527563920
},
"is-referenced-by-count": 23,
"issued": {
"date-parts": [
[
2021,
2,
15
]
]
},
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2021,
2,
15
]
],
"date-time": "2021-02-15T00:00:00Z",
"timestamp": 1613347200000
}
}
],
"link": [
{
"URL": "https://www.frontiersin.org/articles/10.3389/fmolb.2021.607443/full",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "1965",
"original-title": [],
"prefix": "10.3389",
"published": {
"date-parts": [
[
2021,
2,
15
]
]
},
"published-online": {
"date-parts": [
[
2021,
2,
15
]
]
},
"publisher": "Frontiers Media SA",
"reference": [
{
"DOI": "10.1016/s0166-3542(01)00146-2",
"article-title": "Lactoferrin and cyclic lactoferricin inhibit the entry of human cytomegalovirus into human fibroblasts",
"author": "Andersen",
"doi-asserted-by": "publisher",
"first-page": "141",
"journal-title": "Antiviral. Res.",
"key": "B1",
"volume": "51",
"year": "2001"
},
{
"DOI": "10.1007/s00018-005-5368-9",
"article-title": "Lactoferrin",
"author": "Baker",
"doi-asserted-by": "publisher",
"first-page": "2531",
"journal-title": "Cell. Mol. Life Sci.",
"key": "B2",
"volume": "62",
"year": "2005"
},
{
"DOI": "10.3390/molecules16086992",
"article-title": "Antiviral properties of lactoferrin--a natural immunity molecule",
"author": "Berlutti",
"doi-asserted-by": "publisher",
"first-page": "6992",
"journal-title": "Molecules",
"key": "B3",
"volume": "16",
"year": "2011"
},
{
"DOI": "10.1201/9780203911327.ch14",
"article-title": "The protein data bank",
"author": "Berman",
"doi-asserted-by": "crossref",
"first-page": "394",
"key": "B4",
"volume-title": "Protein Structure",
"year": "2003"
},
{
"DOI": "10.1155/2020/4389089",
"article-title": "Sars-CoV-2 envelope and membrane proteins: structural differences linked to virus characteristics?",
"author": "Bianchi",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "BioMed. Res. Int.",
"key": "B5",
"volume": "2020",
"year": "2020"
},
{
"DOI": "10.1093/glycob/cwm005",
"article-title": "Evolution of carbohydrate antigens--microbial forces shaping host glycomes?",
"author": "Bishop",
"doi-asserted-by": "publisher",
"first-page": "23R",
"journal-title": "Glycobiology",
"key": "B6",
"volume": "17",
"year": "2007"
},
{
"DOI": "10.1002/jcc.21287",
"article-title": "CHARMM: the biomolecular simulation program",
"author": "Brooks",
"doi-asserted-by": "publisher",
"first-page": "1545",
"journal-title": "J. Comput. Chem.",
"key": "B7",
"volume": "30",
"year": "2009"
},
{
"DOI": "10.1063/1.2408420",
"article-title": "Canonical sampling through velocity rescaling",
"author": "Bussi",
"doi-asserted-by": "publisher",
"first-page": "014101",
"journal-title": "J. Chem. Phys.",
"key": "B8",
"volume": "126",
"year": "2007"
},
{
"DOI": "10.1179/096805102125001000",
"article-title": "Antimicrobial and immunoregulatory functions of lactoferrin and its potential therapeutic application",
"author": "Caccavo",
"doi-asserted-by": "publisher",
"first-page": "403",
"journal-title": "J. Endotoxin. Res.",
"key": "B9",
"volume": "8",
"year": "2002"
},
{
"DOI": "10.3390/ijms21144903",
"article-title": "Lactoferrin as protective natural barrier of respiratory and intestinal mucosa against coronavirus infection and inflammation",
"author": "Campione",
"doi-asserted-by": "publisher",
"first-page": "4903",
"journal-title": "Ijms",
"key": "B10",
"volume": "21",
"year": "2020"
},
{
"DOI": "10.1021/acscentsci.0c01056",
"article-title": "Beyond shielding: The roles of glycans in the sars-cov-2 spike protein",
"author": "Casalino",
"doi-asserted-by": "publisher",
"first-page": "1722",
"journal-title": "ACS Cent. Sci.",
"key": "B11",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1016/j.ijantimicag.2020.106118",
"article-title": "Lactoferrin as potential preventative and adjunct treatment for COVID-19",
"author": "Chang",
"doi-asserted-by": "publisher",
"first-page": "106118",
"journal-title": "Int. J. Antimicrob. Agents",
"key": "B12",
"volume": "56",
"year": "2020"
},
{
"DOI": "10.1021/ja00119a045",
"article-title": "Molecular dynamics simulations on solvated biomolecular systems: The particle mesh ewald method leads to stable trajectories of DNA, RNA, and proteins",
"author": "Cheatham",
"doi-asserted-by": "publisher",
"first-page": "4193",
"journal-title": "J. Am. Chem. Soc.",
"key": "B13",
"volume": "117",
"year": "1995"
},
{
"DOI": "10.1002/prot.10389",
"article-title": "ZDOCK: an initial-stage protein-docking algorithm",
"author": "Chen",
"doi-asserted-by": "publisher",
"first-page": "80",
"journal-title": "Proteins",
"key": "B14",
"volume": "52",
"year": "2003"
},
{
"DOI": "10.1093/bioinformatics/bty918",
"article-title": "Antibody interface prediction with 3d zernike descriptors and svm",
"author": "Daberdaku",
"doi-asserted-by": "publisher",
"first-page": "1870",
"journal-title": "Bioinformatics",
"key": "B15",
"volume": "35",
"year": "2019"
},
{
"DOI": "10.1101/2020.05.13.093781",
"article-title": "In vitro inhibition of SARS-CoV-2 infection by bovine lactoferrin [Preprint]",
"author": "de Carvalho",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B16",
"year": "2020"
},
{
"DOI": "10.1021/acs.jcim.9b01066",
"article-title": "Quantitative characterization of binding pockets and binding complementarity by means of zernike descriptors",
"author": "Di Rienzo",
"doi-asserted-by": "publisher",
"first-page": "1390",
"journal-title": "J Chem. Inf. Model.",
"key": "B17",
"volume": "60",
"year": "2020"
},
{
"DOI": "10.1038/srep45053",
"article-title": "Superposition-free comparison and clustering of antibody binding sites: implications for the prediction of the nature of their antigen",
"author": "Di Rienzo",
"doi-asserted-by": "publisher",
"first-page": "45053",
"journal-title": "Sci. Rep.",
"key": "B18",
"volume": "7",
"year": "2017"
},
{
"DOI": "10.1016/j.antiviral.2004.05.005",
"article-title": "Lactoferrin inhibits human papillomavirus binding and uptake in vitro",
"author": "Drobni",
"doi-asserted-by": "publisher",
"first-page": "63",
"journal-title": "Antiviral. Res.",
"key": "B19",
"volume": "64",
"year": "2004"
},
{
"DOI": "10.1056/NEJMoa030747",
"article-title": "Identification of a novel coronavirus in patients with severe acute respiratory syndrome",
"author": "Drosten",
"doi-asserted-by": "publisher",
"first-page": "1967",
"journal-title": "N. Engl. J. Med.",
"key": "B20",
"volume": "348",
"year": "2003"
},
{
"DOI": "10.1038/s41592-019-0666-6",
"article-title": "Deciphering interaction fingerprints from protein molecular surfaces using geometric deep learning",
"author": "Gainza",
"doi-asserted-by": "publisher",
"first-page": "184",
"journal-title": "Nat. Methods.",
"key": "B21",
"volume": "17",
"year": "2020"
},
{
"DOI": "10.3390/ph9040061",
"article-title": "Lactoferrin from milk: Nutraceutical and pharmacological properties",
"author": "Giansanti",
"doi-asserted-by": "publisher",
"first-page": "61",
"journal-title": "Pharmaceuticals (Basel)",
"key": "B22",
"volume": "9",
"year": "2016"
},
{
"DOI": "10.1128/JVI.01394-09",
"article-title": "Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission",
"author": "Graham",
"doi-asserted-by": "publisher",
"first-page": "3134",
"journal-title": "J. Virol.",
"key": "B23",
"volume": "84",
"year": "2010"
},
{
"DOI": "10.1021/ja01498a029",
"article-title": "The isolation of a red protein from milk2",
"author": "Groves",
"doi-asserted-by": "publisher",
"first-page": "3345",
"journal-title": "J. Am. Chem. Soc.",
"key": "B24",
"volume": "82",
"year": "1960"
},
{
"DOI": "10.1093/infdis/172.2.380",
"article-title": "Antiviral effects of plasma and milk proteins: Lactoferrin shows potent activity against both human immunodeficiency virus and human cytomegalovirus replication in vitro",
"author": "Harmsen",
"doi-asserted-by": "publisher",
"first-page": "380",
"journal-title": "J. Infect. Dis.",
"key": "B25",
"volume": "172",
"year": "1995"
},
{
"DOI": "10.1002/(sici)1096-987x(199709)18:12<1463::aid-jcc4>3.0.co;2-h",
"article-title": "LINCS: a linear constraint solver for molecular simulations",
"author": "Hess",
"doi-asserted-by": "publisher",
"first-page": "1463",
"journal-title": "J. Comput. Chem.",
"key": "B26",
"volume": "18",
"year": "1997"
},
{
"DOI": "10.1016/s0140-6736(20)30183-5",
"article-title": "Clinical features of patients infected with 2019 novel coronavirus in wuhan, china",
"author": "Huang",
"doi-asserted-by": "publisher",
"first-page": "497",
"journal-title": "Lancet",
"key": "B27",
"volume": "395",
"year": "2020"
},
{
"DOI": "10.1073/pnas.1809667116",
"article-title": "Human coronaviruses oc43 and hku1 bind to 9-o-acetylated sialic acids via a conserved receptor-binding site in spike protein domain a",
"author": "Hulswit",
"doi-asserted-by": "publisher",
"first-page": "2681",
"journal-title": "Proc. Natl. Acad. Sci. USA",
"key": "B28",
"volume": "116",
"year": "2019"
},
{
"DOI": "10.1063/1.445869",
"article-title": "Comparison of simple potential functions for simulating liquid water",
"author": "Jorgensen",
"doi-asserted-by": "publisher",
"first-page": "926",
"journal-title": "J. Chem. Phys.",
"key": "B29",
"volume": "79",
"year": "1983"
},
{
"DOI": "10.2174/138920311796957612",
"article-title": "Molecular surface representation using 3d zernike descriptors for protein shape comparison and docking",
"author": "Kihara",
"doi-asserted-by": "publisher",
"first-page": "520",
"journal-title": "Curr. Protein Pept. Sci.",
"key": "B30",
"volume": "12",
"year": "2011"
},
{
"DOI": "10.1056/NEJMoa030781",
"article-title": "A novel coronavirus associated with severe acute respiratory syndrome",
"author": "Ksiazek",
"doi-asserted-by": "publisher",
"first-page": "1953",
"journal-title": "N. Engl. J. Med.",
"key": "B31",
"volume": "348",
"year": "2003"
},
{
"DOI": "10.1128/jvi.74.3.1393-1406.2000",
"article-title": "Retargeting of coronavirus by substitution of the spike glycoprotein ectodomain: crossing the host cell species barrier",
"author": "Kuo",
"doi-asserted-by": "publisher",
"first-page": "1393",
"journal-title": "J. Virol.",
"key": "B32",
"volume": "74",
"year": "2000"
},
{
"DOI": "10.1371/journal.pone.0023710",
"article-title": "Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans",
"author": "Lang",
"doi-asserted-by": "publisher",
"first-page": "e23710",
"journal-title": "PLoS One",
"key": "B33",
"volume": "6",
"year": "2011"
},
{
"DOI": "10.3389/fimmu.2015.00025",
"article-title": "Complementing the sugar code: Role of GAGs and sialic acid in complement regulation",
"author": "Langford-Smith",
"doi-asserted-by": "publisher",
"first-page": "25",
"journal-title": "Front Immunol",
"key": "B34",
"volume": "6",
"year": "2015"
},
{
"DOI": "10.1128/JVI.00442-08",
"article-title": "Structural analysis of major species barriers between humans and palm civets for severe acute respiratory syndrome coronavirus infections",
"author": "Li",
"doi-asserted-by": "publisher",
"first-page": "6984",
"journal-title": "J. Virol.",
"key": "B35",
"volume": "82",
"year": "2008"
},
{
"DOI": "10.1126/science.1116480",
"article-title": "Structure of sars coronavirus spike receptor-binding domain complexed with receptor",
"author": "Li",
"doi-asserted-by": "publisher",
"first-page": "1864",
"journal-title": "Science",
"key": "B36",
"volume": "309",
"year": ""
},
{
"DOI": "10.1038/sj.emboj.7600640",
"article-title": "Receptor and viral determinants of sars-coronavirus adaptation to human ace2",
"author": "Li",
"doi-asserted-by": "publisher",
"first-page": "1634",
"journal-title": "EMBO. J.",
"key": "B37",
"volume": "24",
"year": ""
},
{
"DOI": "10.1101/2020.05.10.087288",
"article-title": "SARS-CoV-2 spike protein binds heparan sulfate in a length- and sequence-dependent manner [Preprint]",
"author": "Liu",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B38",
"year": "2020"
},
{
"DOI": "10.1016/j.virol.2003.09.029",
"article-title": "Inhibition of herpes simplex virus infection by lactoferrin is dependent on interference with the virus binding to glycosaminoglycans",
"author": "Marchetti",
"doi-asserted-by": "publisher",
"first-page": "405",
"journal-title": "Virology",
"key": "B39",
"volume": "318",
"year": "2004"
},
{
"DOI": "10.1046/j.1365-2672.2003.02071.x",
"article-title": "The effect of bovine lactoferrin and lactoferricin b on the ability of feline calicivirus (a norovirus surrogate) and poliovirus to infect cell cultures",
"author": "McCann",
"doi-asserted-by": "publisher",
"first-page": "1026",
"journal-title": "J. Appl. Microbiol.",
"key": "B40",
"volume": "95",
"year": "2003"
},
{
"DOI": "10.1101/2020.03.24.006197",
"article-title": "In-silico evidence for two receptors based strategy of sars-cov-2",
"author": "Milanetti",
"doi-asserted-by": "publisher",
"journal-title": "arXiv, arXiv–2003",
"key": "B41",
"year": ""
},
{
"DOI": "10.1016/j.csbj.2020.11.051",
"article-title": "2d zernike polynomial expansion: finding the protein-protein binding regions",
"author": "Milanetti",
"doi-asserted-by": "publisher",
"first-page": "29",
"journal-title": "Comput. Struct. Biotechnol. J.",
"key": "B42",
"volume": "19",
"year": ""
},
{
"article-title": "Does blood type affect the covid-19 infection pattern?",
"author": "Miotto",
"journal-title": "arXiv:2007.06296",
"key": "B43",
"year": "2020"
},
{
"DOI": "10.1080/10942912.2019.1666137",
"article-title": "Lactoferrin (LF): a natural antimicrobial protein",
"author": "Niaz",
"doi-asserted-by": "publisher",
"first-page": "1626",
"journal-title": "International Journal of Food Properties",
"key": "B44",
"volume": "22",
"year": "2019"
},
{
"DOI": "10.1038/s41594-019-0334-7",
"article-title": "Structures of MERS-CoV spike glycoprotein in complex with sialoside attachment receptors",
"author": "Park",
"doi-asserted-by": "publisher",
"first-page": "1151",
"journal-title": "Nat. Struct. Mol. Biol.",
"key": "B45",
"volume": "26",
"year": "2019"
},
{
"DOI": "10.1103/physrevlett.45.1196",
"article-title": "Crystal structure and pair potentials: A molecular-dynamics study",
"author": "Parrinello",
"doi-asserted-by": "publisher",
"first-page": "1196",
"journal-title": "Phys. Rev. Lett.",
"key": "B46",
"volume": "45",
"year": "1980"
},
{
"DOI": "10.12932/AP-200220-0772",
"article-title": "Immune responses in covid-19 and potential vaccines: Lessons learned from sars and mers epidemic",
"author": "Prompetchara",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "Asian Pac. J. Allergy Immunol.",
"key": "B47",
"volume": "38",
"year": "2020"
},
{
"DOI": "10.1016/s1357-2725(98)00066-1",
"article-title": "Antiviral effect of bovine lactoferrin saturated with metal ions on early steps of human immunodeficiency virus type 1 infection",
"author": "Puddu",
"doi-asserted-by": "publisher",
"first-page": "1055",
"journal-title": "Int. J. Biochem. Cell Biol.",
"key": "B48",
"volume": "30",
"year": "1998"
},
{
"DOI": "10.1016/j.sbi.2016.10.003",
"article-title": "Glycan-protein interactions in viral pathogenesis",
"author": "Raman",
"doi-asserted-by": "publisher",
"first-page": "153",
"journal-title": "Curr. Opin. Struct. Biol.",
"key": "B49",
"volume": "40",
"year": "2016"
},
{
"DOI": "10.1016/j.crvi.2014.08.003",
"article-title": "Potential lactoferrin activity against pathogenic viruses",
"author": "Redwan",
"doi-asserted-by": "publisher",
"first-page": "581",
"journal-title": "C R Biol.",
"key": "B50",
"volume": "337",
"year": "2014"
},
{
"DOI": "10.1146/annurev.bb.06.060177.001055",
"article-title": "Areas, volumes, packing and protein structure",
"author": "Richards",
"doi-asserted-by": "publisher",
"first-page": "151",
"journal-title": "Annu. Rev. Biophys. Bioeng.",
"key": "B51",
"volume": "6",
"year": "1977"
},
{
"DOI": "10.1016/j.compbiomed.2020.103849",
"article-title": "Bioinformatics studies on a function of the sars-cov-2 spike glycoprotein as the binding of host sialic acid glycans.",
"author": "Robson",
"doi-asserted-by": "publisher",
"first-page": "103849",
"journal-title": "Comput. Biol. Med",
"key": "B52",
"volume": "122",
"year": "2020"
},
{
"DOI": "10.1038/nprot.2010.5",
"article-title": "I-TASSER: a unified platform for automated protein structure and function prediction",
"author": "Roy",
"doi-asserted-by": "publisher",
"first-page": "725",
"journal-title": "Nat. Protoc.",
"key": "B53",
"volume": "5",
"year": "2010"
},
{
"DOI": "10.1007/s10719-006-5437-9",
"article-title": "Sialic acids as receptor determinants for coronaviruses",
"author": "Schwegmann-Weßels",
"doi-asserted-by": "publisher",
"first-page": "51",
"journal-title": "Glycoconj J",
"key": "B54",
"volume": "23",
"year": "2006"
},
{
"DOI": "10.1038/s41433-020-0790-7",
"article-title": "Revisiting the dangers of the coronavirus in the ophthalmology practice",
"author": "Seah",
"doi-asserted-by": "publisher",
"first-page": "1155",
"journal-title": "Eye (Lond)",
"key": "B55",
"volume": "34",
"year": "2020"
},
{
"DOI": "10.1038/s41586-019-1923-7",
"article-title": "Improved protein structure prediction using potentials from deep learning",
"author": "Senior",
"doi-asserted-by": "publisher",
"first-page": "706",
"journal-title": "Nature",
"key": "B56",
"volume": "577",
"year": "2020"
},
{
"DOI": "10.3168/jds.s0022-0302(40)95543-6",
"article-title": "The proteins in whey. Compte rendu des Travaux du Laboratoire de Carlsberg",
"author": "Sorensen",
"doi-asserted-by": "publisher",
"first-page": "55",
"journal-title": "Ser. Chim.",
"key": "B57",
"volume": "23",
"year": "1940"
},
{
"DOI": "10.1002/jcc.20291",
"article-title": "GROMACS: fast, flexible, and free",
"author": "Van Der Spoel",
"doi-asserted-by": "publisher",
"first-page": "1701",
"journal-title": "J. Comput. Chem.",
"key": "B58",
"volume": "26",
"year": "2005"
},
{
"DOI": "10.1016/j.tim.2016.03.003",
"article-title": "Epidemiology, genetic recombination, and pathogenesis of coronaviruses",
"author": "Su",
"doi-asserted-by": "publisher",
"first-page": "490",
"journal-title": "Trends Microbiol.",
"key": "B59",
"volume": "24",
"year": "2016"
},
{
"DOI": "10.1016/s0304-4165(01)00178-7",
"article-title": "Involvement of bovine lactoferrin metal saturation, sialic acid and protein fragments in the inhibition of rotavirus infection",
"author": "Superti",
"doi-asserted-by": "publisher",
"first-page": "107",
"journal-title": "Biochim. Biophys. Acta.",
"key": "B60",
"volume": "1528",
"year": "2001"
},
{
"DOI": "10.20411/pai.v5i1.377",
"article-title": "The structure of the membrane protein of SARS-CoV-2 resembles the sugar transporter SemiSWEET",
"author": "Thomas",
"doi-asserted-by": "publisher",
"first-page": "342",
"journal-title": "Pathog Immun",
"key": "B61",
"volume": "5",
"year": "2020"
},
{
"DOI": "10.1038/s41594-019-0233-y",
"article-title": "Structural basis for human coronavirus attachment to sialic acid receptors",
"author": "Tortorici",
"doi-asserted-by": "publisher",
"first-page": "481",
"journal-title": "Nat. Struct. Mol. Biol.",
"key": "B62",
"volume": "26",
"year": "2019"
},
{
"DOI": "10.1016/s0166-3542(01)00195-4",
"article-title": "Antiviral activities of lactoferrin",
"author": "van der Strate",
"doi-asserted-by": "publisher",
"first-page": "225",
"journal-title": "Antiviral Res",
"key": "B63",
"volume": "52",
"year": "2001"
},
{
"DOI": "10.1093/nar/gkaa864",
"article-title": "Structural analysis of sars-cov-2 and prediction of the human interactome",
"author": "Vandelli",
"doi-asserted-by": "publisher",
"first-page": "11270",
"journal-title": "Nucleic Acids Res",
"key": "B64",
"year": "2020"
},
{
"DOI": "10.1093/glycob/cww086",
"article-title": "Biological roles of glycans",
"author": "Varki",
"doi-asserted-by": "publisher",
"first-page": "3",
"journal-title": "Glycobiology",
"key": "B65",
"volume": "27",
"year": "2016"
},
{
"DOI": "10.1186/1471-2105-10-407",
"article-title": "Protein-protein docking using region-based 3d zernike descriptors",
"author": "Venkatraman",
"doi-asserted-by": "publisher",
"first-page": "407",
"journal-title": "BMC bioinformatics",
"key": "B66",
"volume": "10",
"year": "2009"
},
{
"DOI": "10.1016/j.virol.2005.01.010",
"article-title": "Antiviral activity of human lactoferrin: inhibition of alphavirus interaction with heparan sulfate",
"author": "Waarts",
"doi-asserted-by": "publisher",
"first-page": "284",
"journal-title": "Virology",
"key": "B67",
"volume": "333",
"year": "2005"
},
{
"DOI": "10.1126/science.abb2762",
"article-title": "Structural basis for the recognition of the sars-cov-2 by full-length human ace2",
"author": "Yan",
"doi-asserted-by": "publisher",
"first-page": "1444",
"journal-title": "Science",
"key": "B68",
"volume": "367",
"year": "2020"
},
{
"DOI": "10.1128/JVI.71.8.5997-6002.1997",
"article-title": "Hepatitis C virus envelope proteins bind lactoferrin",
"author": "Yi",
"doi-asserted-by": "publisher",
"first-page": "5997",
"journal-title": "J. Virol.",
"key": "B69",
"volume": "71",
"year": "1997"
},
{
"DOI": "10.1056/NEJMoa1211721",
"article-title": "Isolation of a novel coronavirus from a man with pneumonia in saudi arabia",
"author": "Zaki",
"doi-asserted-by": "publisher",
"first-page": "1814",
"journal-title": "N Engl J Med",
"key": "B70",
"volume": "367",
"year": "2012"
},
{
"DOI": "10.1093/nar/gki524",
"article-title": "Tm-align: a protein structure alignment algorithm based on the tm-score",
"author": "Zhang",
"doi-asserted-by": "publisher",
"first-page": "2302",
"journal-title": "Nucleic Acids Res",
"key": "B71",
"volume": "33",
"year": "2005"
},
{
"DOI": "10.1038/s41586-020-2012-7",
"article-title": "A pneumonia outbreak associated with a new coronavirus of probable bat origin",
"author": "Zhou",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "Nature",
"key": "B72",
"volume": "579",
"year": "2020"
},
{
"DOI": "10.1056/NEJMoa2001017",
"article-title": "A novel coronavirus from patients with pneumonia in china, 2019",
"author": "Zhu",
"doi-asserted-by": "publisher",
"first-page": "727",
"journal-title": "N. Engl. J. Med.",
"key": "B73",
"volume": "382",
"year": "2020"
}
],
"reference-count": 73,
"references-count": 73,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.frontiersin.org/articles/10.3389/fmolb.2021.607443/full"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"Biochemistry, Genetics and Molecular Biology (miscellaneous)",
"Molecular Biology",
"Biochemistry"
],
"subtitle": [],
"title": "Molecular Mechanisms Behind Anti SARS-CoV-2 Action of Lactoferrin",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.3389/crossmark-policy",
"volume": "8"
}