Evaluation of SARS-CoV-2 isolation in cell culture from nasal/nasopharyngeal swabs or saliva specimens of patients with COVID-19
et al., Research Square, doi:10.21203/rs.3.rs-2676422/v1, Mar 2023
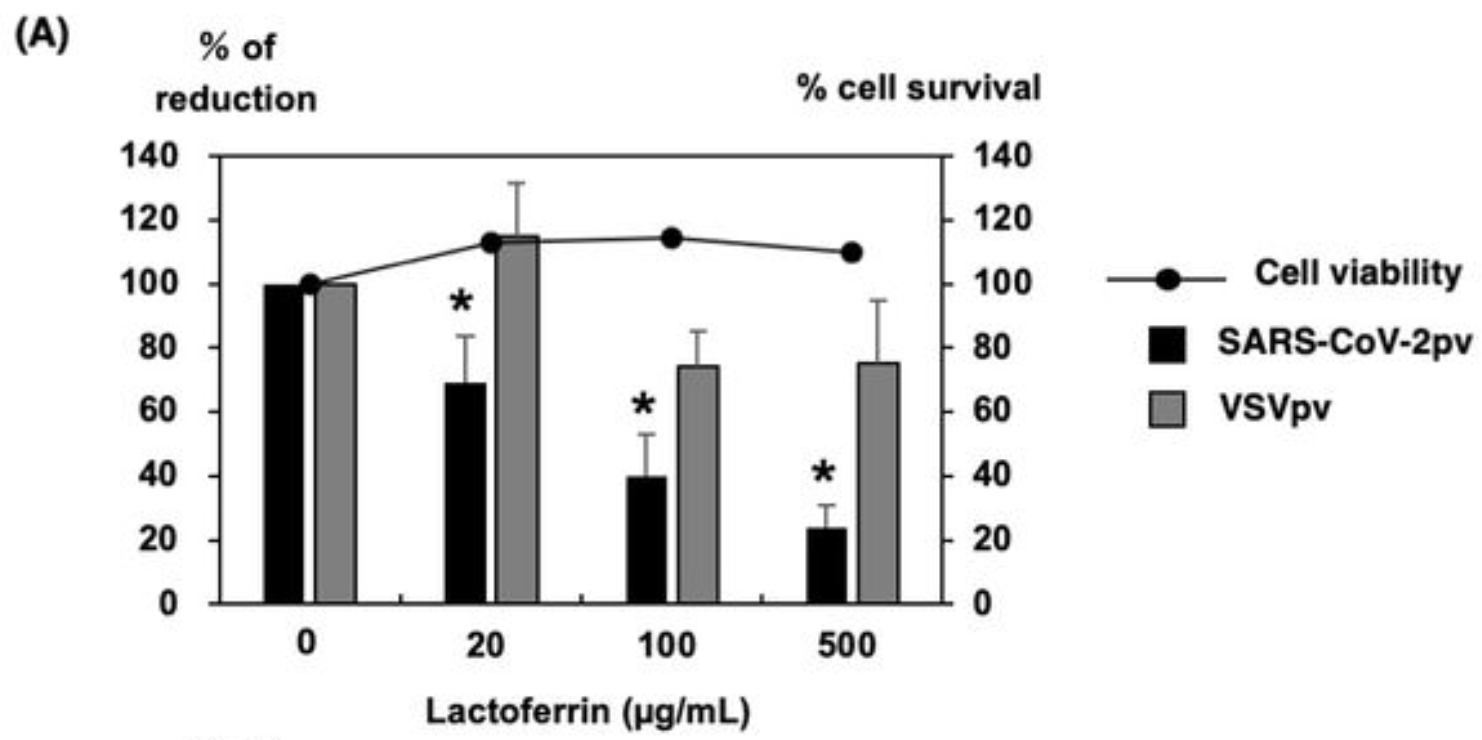
Analysis of nasal/nasopharyngeal swabs and saliva specimens, showing significantly lower isolation efficiency of SARS-CoV-2 in saliva specimens. In vitro study showing that lactoferrin and amylase, components of saliva, were found to inhibit SARS-CoV-2 pseudotyped virus infection.
18 preclinical studies support the efficacy of lactoferrin for COVID-19:
1.
da Silva et al., Immunomodulatory effect of bovine lactoferrin during SARS-CoV-2 infection, Frontiers in Immunology, doi:10.3389/fimmu.2024.1456634.
2.
Cutone et al., Lactoferrin binding to Sars-CoV-2 Spike glycoprotein protects host from infection, inflammation and iron dysregulation., Research Square, doi:10.21203/rs.3.rs-1605740/v1.
3.
Miotto et al., Molecular Mechanisms Behind Anti SARS-CoV-2 Action of Lactoferrin, Frontiers in Molecular Biosciences, doi:10.3389/fmolb.2021.607443.
4.
Babulic et al., Lactoferrin Binds through Its N-Terminus to the Receptor-Binding Domain of the SARS-CoV-2 Spike Protein, Pharmaceuticals, doi:10.3390/ph17081021.
5.
Yathindranath et al., Lipid Nanoparticle-Based Inhibitors for SARS-CoV-2 Host Cell Infection, International Journal of Nanomedicine, doi:10.2147/IJN.S448005.
6.
Alves et al., Inhibition of SARS-CoV-2 Infection in Vero Cells by Bovine Lactoferrin under Different Iron-Saturation States, Pharmaceuticals, doi:10.3390/ph16101352.
7.
Kobayashi-Sakamoto et al., Bovine lactoferrin suppresses the cathepsin-dependent pathway of SARS-CoV-2 entry in vitro, International Dairy Journal, doi:10.1016/j.idairyj.2023.105805.
8.
Andreu et al., Liposomal Lactoferrin Exerts Antiviral Activity against HCoV-229E and SARS-CoV-2 Pseudoviruses In Vitro, Viruses, doi:10.3390/v15040972.
9.
Yazawa et al., Evaluation of SARS-CoV-2 isolation in cell culture from nasal/nasopharyngeal swabs or saliva specimens of patients with COVID-19, Research Square, doi:10.21203/rs.3.rs-2676422/v1.
10.
Piacentini et al., Lactoferrin Inhibition of the Complex Formation between ACE2 Receptor and SARS CoV-2 Recognition Binding Domain, International Journal of Molecular Sciences, doi:10.3390/ijms23105436.
11.
Ostrov et al., Highly Specific Sigma Receptor Ligands Exhibit Anti-Viral Properties in SARS-CoV-2 Infected Cells, Pathogens, doi:10.3390/pathogens10111514.
12.
Mirabelli et al., Morphological cell profiling of SARS-CoV-2 infection identifies drug repurposing candidates for COVID-19, Proceedings of the National Academy of Sciences, doi:10.1073/pnas.2105815118.
13.
Salaris et al., Protective Effects of Lactoferrin against SARS-CoV-2 Infection In Vitro, Nutrients, doi:10.3390/nu13020328.
Yazawa et al., 31 Mar 2023, preprint, 8 authors.
Contact: toyamaeiken3@juno.ocn.ne.jp.
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
Evaluation of SARS-CoV-2 isolation in cell culture from nasal/nasopharyngeal swabs or saliva specimens of patients with COVID-19
doi:10.21203/rs.3.rs-2676422/v1
It has been revealed that SARS-CoV-2 can be e ciently isolated from clinical specimens such as nasal/nasopharyngeal swabs or saliva in cultured cells. In this study, we examined the e ciency of viral isolation including SARS-CoV-2 mutant strains between nasal/nasopharyngeal swab or saliva specimens. Furthermore, we also examined the comparison of viral isolation rates by sample species using simulated specimens for COVID-19. As a result, it was found that the isolation e ciency of SARS-CoV-2 in the saliva specimens was signi cantly lower than that in the nasal/nasopharyngeal swab specimens. In order to determine which component of saliva is responsible for the lower isolation rate of saliva specimens, we tested the abilities of lactoferrin, amylase, cathelicidin, and mucin, which are considered to be abundant in saliva, to inhibit the infection of SARS-CoV-2 pseudotyped viruses (SARS-CoV-2pv). Lactoferrin and amylase were found to inhibit SARS-CoV-2pv infection. In conclusion, even if the same number of viral genome copies was detected by the real-time RT-PCR test, infection of SARS-CoV-2 present in saliva is thought to be inhibited by inhibitory factors such as lactoferrin and amylase, compared to nasal/nasopharyngeal swab specimens.
Declarations Ethical approval This study was performed in accordance with the Helsinki Declaration and was approved by the ethical review board of the Toyama Institute of Health (approval No.: R2-1). The need of informed consent was also waived by the ethical review board of the Toyama Institute of Health (approval No.: R2-1).
Con ict of interest We have no potential con icts of interest in relation to this work.
References
Dale, Localized antimicrobial peptide expression in human gingiva, J Periodontal Res, doi:10.1034/j.1600-0765.2001.360503.x
Griesemer, Evaluation of Specimen Types and Saliva Stabilization Solutions for SARS-CoV-2 Testing, J Clin Microbiol, doi:10.1128/JCM.01418-20
Hu, Meng, Zhang, Xiang, Wang, The in vitro antiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate coreceptor, Emerg Microbes Infect, doi:10.1080/22221751.2021.1888660
Igarashi, Viral isolation analysis of SARS-CoV-2 from clinical specimens of COVID-19 patients, J Infect Chemother, doi:10.1016/j.jiac.2021.10.028
Kim, Duration of Culturable SARS-CoV-2 in Hospitalized Patients with Covid-19, N Engl J Med, doi:10.1056/NEJMc2027040
Kobayashi-Sakamoto, Bovine lactoferrin increases the poly(I:C)-induced antiviral response in vitro, Biochem Cell Biol, doi:10.1139/bcb-2021-0342
Lai, Identi ed human breast milk compositions effectively inhibit SARS-CoV-2 and variants infection and replication, iScience, doi:10.1016/j.isci.2022.104136
Lynge Pedersen, Belstrom, The role of natural salivary defences in maintaining a healthy oral microbiota, J Dent, doi:10.1016/j.jdent.2018.08.010
Matsuyama, Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells, Proc Natl Acad Sci U S A, doi:10.1073/pnas.2002589117
Scola, Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards, Eur J Clin Microbiol Infect Dis, doi:10.1007/s10096-020-03913-9
Singanayagam, Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to, Euro Surveill, doi:10.2807/1560-7917.ES.2020.25.32.2001483
Suzuki, Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant, Nature, doi:10.1038/s41586-022-04462-1
Tani, Evaluation of SARS-CoV-2 neutralizing antibodies using a vesicular stomatitis virus possessing SARS-CoV-2 spike protein, Virol J, doi:10.1186/s12985-021-01490-7
Tenovuo, Lehtonen, Aaltonen, Vilja, Tuohimaa, Antimicrobial factors in whole saliva of human infants, Infect Immun, doi:10.1128/iai.51.1.49-53.1986
Yamaguchi, Deguchi, Miyazaki, The effects of exercise in forest and urban environments on sympathetic nervous activity of normal young adults, J Int Med Res, doi:10.1177/147323000603400204
Young, Viral Dynamics and Immune Correlates of Coronavirus Disease 2019 (COVID-19) Severity, Clin Infect Dis, doi:10.1093/cid/ciaa1280
Zhang, An updated review of SARS-CoV-2 detection methods in the context of a novel coronavirus pandemic, Bioeng Transl Med, doi:10.1002/btm2.10356
DOI record:
{
"DOI": "10.21203/rs.3.rs-2676422/v1",
"URL": "http://dx.doi.org/10.21203/rs.3.rs-2676422/v1",
"abstract": "<jats:title>Abstract</jats:title>\n <jats:p>It has been revealed that SARS-CoV-2 can be efficiently isolated from clinical specimens such as nasal/nasopharyngeal swabs or saliva in cultured cells. In this study, we examined the efficiency of viral isolation including SARS-CoV-2 mutant strains between nasal/nasopharyngeal swab or saliva specimens. Furthermore, we also examined the comparison of viral isolation rates by sample species using simulated specimens for COVID-19. As a result, it was found that the isolation efficiency of SARS-CoV-2 in the saliva specimens was significantly lower than that in the nasal/nasopharyngeal swab specimens. In order to determine which component of saliva is responsible for the lower isolation rate of saliva specimens, we tested the abilities of lactoferrin, amylase, cathelicidin, and mucin, which are considered to be abundant in saliva, to inhibit the infection of SARS-CoV-2 pseudotyped viruses (SARS-CoV-2pv). Lactoferrin and amylase were found to inhibit SARS-CoV-2pv infection. In conclusion, even if the same number of viral genome copies was detected by the real-time RT-PCR test, infection of SARS-CoV-2 present in saliva is thought to be inhibited by inhibitory factors such as lactoferrin and amylase, compared to nasal/nasopharyngeal swab specimens.</jats:p>",
"accepted": {
"date-parts": [
[
2023,
3,
10
]
]
},
"author": [
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Yazawa",
"given": "Shunsuke",
"sequence": "first"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Yamazaki",
"given": "Emiko",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Saga",
"given": "Yumiko",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Itamochi",
"given": "Masae",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Inasaki",
"given": "Noriko",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Shimada",
"given": "Takahisa",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Oishi",
"given": "Kazunori",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Toyama Institute of Health"
}
],
"family": "Tani",
"given": "Hideki",
"sequence": "additional"
}
],
"container-title": [],
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2023,
3,
31
]
],
"date-time": "2023-03-31T19:45:56Z",
"timestamp": 1680291956000
},
"deposited": {
"date-parts": [
[
2023,
3,
31
]
],
"date-time": "2023-03-31T19:46:00Z",
"timestamp": 1680291960000
},
"group-title": "In Review",
"indexed": {
"date-parts": [
[
2023,
4,
1
]
],
"date-time": "2023-04-01T04:47:28Z",
"timestamp": 1680324448422
},
"institution": [
{
"name": "Research Square"
}
],
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2023,
3,
31
]
]
},
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "unspecified",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
3,
31
]
],
"date-time": "2023-03-31T00:00:00Z",
"timestamp": 1680220800000
}
}
],
"link": [
{
"URL": "https://www.researchsquare.com/article/rs-2676422/v1",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.researchsquare.com/article/rs-2676422/v1.html",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "8761",
"original-title": [],
"posted": {
"date-parts": [
[
2023,
3,
31
]
]
},
"prefix": "10.21203",
"published": {
"date-parts": [
[
2023,
3,
31
]
]
},
"publisher": "Research Square Platform LLC",
"reference": [
{
"DOI": "10.1002/btm2.10356",
"author": "Zhang Y",
"doi-asserted-by": "publisher",
"key": "ref1",
"unstructured": "Zhang, Y. et al. An updated review of SARS-CoV-2 detection methods in the context of a novel coronavirus pandemic. Bioeng Transl Med, e10356, doi:10.1002/btm2.10356 (2022).",
"year": "2022"
},
{
"DOI": "10.1128/JCM.01418-20",
"article-title": "Evaluation of Specimen Types and Saliva Stabilization Solutions for SARS-CoV-2 Testing",
"author": "Griesemer SB",
"doi-asserted-by": "publisher",
"journal-title": "J Clin Microbiol",
"key": "ref2",
"unstructured": "Griesemer, S. B. et al. Evaluation of Specimen Types and Saliva Stabilization Solutions for SARS-CoV-2 Testing. J Clin Microbiol 59, doi:10.1128/JCM.01418-20 (2021).",
"volume": "59",
"year": "2021"
},
{
"DOI": "10.1016/j.jiac.2021.10.028",
"article-title": "Viral isolation analysis of SARS-CoV-2 from clinical specimens of COVID-19 patients",
"author": "Igarashi E",
"doi-asserted-by": "publisher",
"first-page": "347",
"journal-title": "J Infect Chemother",
"key": "ref3",
"unstructured": "Igarashi, E. et al. Viral isolation analysis of SARS-CoV-2 from clinical specimens of COVID-19 patients. J Infect Chemother 28, 347–351, doi:10.1016/j.jiac.2021.10.028 (2022).",
"volume": "28",
"year": "2022"
},
{
"DOI": "10.1034/j.1600-0765.2001.360503.x",
"article-title": "Localized antimicrobial peptide expression in human gingiva",
"author": "Dale BA",
"doi-asserted-by": "publisher",
"first-page": "285",
"journal-title": "J Periodontal Res",
"key": "ref4",
"unstructured": "Dale, B. A. et al. Localized antimicrobial peptide expression in human gingiva. J Periodontal Res 36, 285–294, doi:10.1034/j.1600-0765.2001.360503.x (2001).",
"volume": "36",
"year": "2001"
},
{
"DOI": "10.1016/j.jdent.2018.08.010",
"article-title": "The role of natural salivary defences in maintaining a healthy oral microbiota",
"author": "Lynge Pedersen AM",
"doi-asserted-by": "publisher",
"first-page": "S3-S12",
"issue": "1",
"journal-title": "J Dent",
"key": "ref5",
"unstructured": "Lynge Pedersen, A. M. & Belstrom, D. The role of natural salivary defences in maintaining a healthy oral microbiota. J Dent 80 Suppl 1, S3-S12, doi:10.1016/j.jdent.2018.08.010 (2019).",
"volume": "80",
"year": "2019"
},
{
"DOI": "10.1093/cid/ciaa1280",
"article-title": "Viral Dynamics and Immune Correlates of Coronavirus Disease 2019 (COVID-19) Severity",
"author": "Young BE",
"doi-asserted-by": "publisher",
"first-page": "e2932-e2942",
"journal-title": "Clin Infect Dis",
"key": "ref6",
"unstructured": "Young, B. E. et al. Viral Dynamics and Immune Correlates of Coronavirus Disease 2019 (COVID-19) Severity. Clin Infect Dis 73, e2932-e2942, doi:10.1093/cid/ciaa1280 (2021).",
"volume": "73",
"year": "2021"
},
{
"DOI": "10.2807/1560-7917.ES.2020.25.32.2001483",
"article-title": "Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020",
"author": "Singanayagam A",
"doi-asserted-by": "publisher",
"journal-title": "Euro Surveill",
"key": "ref7",
"unstructured": "Singanayagam, A. et al. Duration of infectiousness and correlation with RT-PCR cycle threshold values in cases of COVID-19, England, January to May 2020. Euro Surveill 25, doi:10.2807/1560-7917.ES.2020.25.32.2001483 (2020).",
"volume": "25",
"year": "2020"
},
{
"DOI": "10.1007/s10096-020-03913-9",
"article-title": "Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards",
"author": "Scola B",
"doi-asserted-by": "publisher",
"first-page": "1059",
"journal-title": "Eur J Clin Microbiol Infect Dis",
"key": "ref8",
"unstructured": "La Scola, B. et al. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur J Clin Microbiol Infect Dis 39, 1059–1061, doi:10.1007/s10096-020-03913-9 (2020).",
"volume": "39",
"year": "2020"
},
{
"DOI": "10.1056/NEJMc2027040",
"article-title": "Duration of Culturable SARS-CoV-2 in Hospitalized Patients with Covid-19",
"author": "Kim MC",
"doi-asserted-by": "publisher",
"first-page": "671",
"journal-title": "N Engl J Med",
"key": "ref9",
"unstructured": "Kim, M. C. et al. Duration of Culturable SARS-CoV-2 in Hospitalized Patients with Covid-19. N Engl J Med 384, 671–673, doi:10.1056/NEJMc2027040 (2021).",
"volume": "384",
"year": "2021"
},
{
"DOI": "10.1038/s41586-022-04462-1",
"article-title": "Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant",
"author": "Suzuki R",
"doi-asserted-by": "publisher",
"first-page": "700",
"journal-title": "Nature",
"key": "ref10",
"unstructured": "Suzuki, R. et al. Attenuated fusogenicity and pathogenicity of SARS-CoV-2 Omicron variant. Nature 603, 700–705, doi:10.1038/s41586-022-04462-1 (2022).",
"volume": "603",
"year": "2022"
},
{
"DOI": "10.1080/22221751.2021.1888660",
"article-title": "The in vitro antiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate co-receptor",
"author": "Hu Y",
"doi-asserted-by": "publisher",
"first-page": "317",
"journal-title": "Emerg Microbes Infect",
"key": "ref11",
"unstructured": "Hu, Y., Meng, X., Zhang, F., Xiang, Y. & Wang, J. The in vitro antiviral activity of lactoferrin against common human coronaviruses and SARS-CoV-2 is mediated by targeting the heparan sulfate co-receptor. Emerg Microbes Infect 10, 317–330, doi:10.1080/22221751.2021.1888660 (2021).",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1139/bcb-2021-0342",
"article-title": "Bovine lactoferrin increases the poly(I:C)-induced antiviral response in vitro",
"author": "Kobayashi-Sakamoto M",
"doi-asserted-by": "publisher",
"first-page": "338",
"journal-title": "Biochem Cell Biol",
"key": "ref12",
"unstructured": "Kobayashi-Sakamoto, M. et al. Bovine lactoferrin increases the poly(I:C)-induced antiviral response in vitro. Biochem Cell Biol 100, 338–348, doi:10.1139/bcb-2021-0342 (2022).",
"volume": "100",
"year": "2022"
},
{
"DOI": "10.1128/iai.51.1.49-53.1986",
"article-title": "Antimicrobial factors in whole saliva of human infants",
"author": "Tenovuo J",
"doi-asserted-by": "publisher",
"first-page": "49",
"journal-title": "Infect Immun",
"key": "ref13",
"unstructured": "Tenovuo, J., Lehtonen, O. P., Aaltonen, A. S., Vilja, P. & Tuohimaa, P. Antimicrobial factors in whole saliva of human infants. Infect Immun 51, 49–53, doi:10.1128/iai.51.1.49-53.1986 (1986).",
"volume": "51",
"year": "1986"
},
{
"DOI": "10.1016/j.isci.2022.104136",
"article-title": "Identified human breast milk compositions effectively inhibit SARS-CoV-2 and variants infection and replication",
"author": "Lai X",
"doi-asserted-by": "publisher",
"first-page": "104136",
"journal-title": "iScience",
"key": "ref14",
"unstructured": "Lai, X. et al. Identified human breast milk compositions effectively inhibit SARS-CoV-2 and variants infection and replication. iScience 25, 104136, doi:10.1016/j.isci.2022.104136 (2022).",
"volume": "25",
"year": "2022"
},
{
"DOI": "10.1177/147323000603400204",
"article-title": "The effects of exercise in forest and urban environments on sympathetic nervous activity of normal young adults",
"author": "Yamaguchi M",
"doi-asserted-by": "publisher",
"first-page": "152",
"journal-title": "J Int Med Res",
"key": "ref15",
"unstructured": "Yamaguchi, M., Deguchi, M. & Miyazaki, Y. The effects of exercise in forest and urban environments on sympathetic nervous activity of normal young adults. J Int Med Res 34, 152–159, doi:10.1177/147323000603400204 (2006).",
"volume": "34",
"year": "2006"
},
{
"DOI": "10.1073/pnas.2002589117",
"article-title": "Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells",
"author": "Matsuyama S",
"doi-asserted-by": "publisher",
"first-page": "7001",
"journal-title": "Proc Natl Acad Sci U S A",
"key": "ref16",
"unstructured": "Matsuyama, S. et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc Natl Acad Sci U S A 117, 7001–7003, doi:10.1073/pnas.2002589117 (2020).",
"volume": "117",
"year": "2020"
},
{
"DOI": "10.1186/s12985-021-01490-7",
"article-title": "Evaluation of SARS-CoV-2 neutralizing antibodies using a vesicular stomatitis virus possessing SARS-CoV-2 spike protein",
"author": "Tani H",
"doi-asserted-by": "publisher",
"first-page": "16",
"journal-title": "Virol J",
"key": "ref17",
"unstructured": "Tani, H. et al. Evaluation of SARS-CoV-2 neutralizing antibodies using a vesicular stomatitis virus possessing SARS-CoV-2 spike protein. Virol J 18, 16, doi:10.1186/s12985-021-01490-7 (2021).",
"volume": "18",
"year": "2021"
}
],
"reference-count": 17,
"references-count": 17,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.researchsquare.com/article/rs-2676422/v1"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subtitle": [],
"subtype": "preprint",
"title": "Evaluation of SARS-CoV-2 isolation in cell culture from nasal/nasopharyngeal swabs or saliva specimens of patients with COVID-19",
"type": "posted-content"
}