Repurposing the natural compounds as potential therapeutic agents for COVID-19 based on the molecular docking study of the main protease and the receptor-binding domain of spike protein
, V., Journal of Molecular Modeling, doi:10.1007/s00894-022-05138-3, May 2022
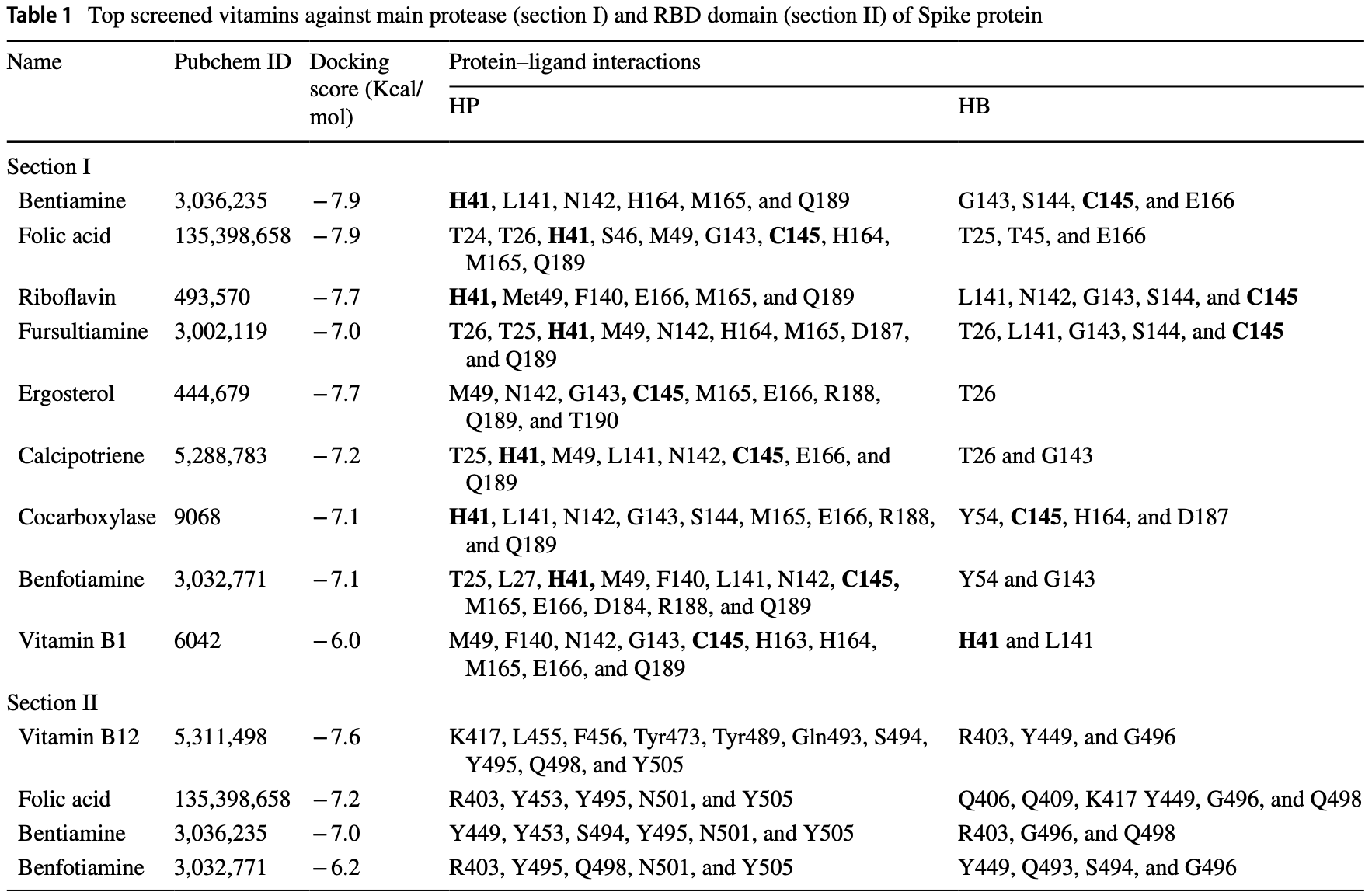
In silico study identifying folic acid, among other compounds, as a potential inhibitor of the SARS-CoV-2 Mpro and spike proteins.
14 preclinical studies support the efficacy of vitamin B9 for COVID-19:
Vitamin B9 has been identified by the European Food Safety Authority (EFSA) as having sufficient evidence for a causal relationship between intake and optimal immune system function12-14.
Vitamin B9 inhibits SARS-CoV-2 in silico3-11, reduces spike protein binding ability11, binds with the spike protein receptor binding domain for alpha and omicron variants2, inhibits the SARS-CoV-2 nucleocapsid protein3, inhibits 3CLpro and PLpro in enzymatic assays2, significantly reduces infection for alpha and omicron SARS-CoV-2 pseudoviruses2, and inhibits ACE2 expression and SARS-CoV-2 infection in a mouse model11.
1.
Wu et al., Biomarkers Prediction and Immune Landscape in Covid-19 and “Brain Fog”, Elsevier BV, doi:10.2139/ssrn.4897774.
2.
Pennisi et al., An Integrated In Silico and In Vitro Approach for the Identification of Natural Products Active against SARS-CoV-2, Biomolecules, doi:10.3390/biom14010043.
3.
Chen et al., Folic acid: a potential inhibitor against SARS-CoV-2 nucleocapsid protein, Pharmaceutical Biology, doi:10.1080/13880209.2022.2063341.
4.
Eskandari, V., Repurposing the natural compounds as potential therapeutic agents for COVID-19 based on the molecular docking study of the main protease and the receptor-binding domain of spike protein, Journal of Molecular Modeling, doi:10.1007/s00894-022-05138-3.
5.
Pandya et al., Unravelling Vitamin B12 as a potential inhibitor against SARS-CoV-2: A computational approach, Informatics in Medicine Unlocked, doi:10.1016/j.imu.2022.100951.
6.
Serseg et al., Hispidin and Lepidine E: Two Natural Compounds and Folic Acid as Potential Inhibitors of 2019-novel Coronavirus Main Protease (2019-nCoVMpro), Molecular Docking and SAR Study, Current Computer-Aided Drug Design, doi:10.2174/1573409916666200422075440.
7.
Hosseini et al., Computational molecular docking and virtual screening revealed promising SARS-CoV-2 drugs, Precision Clinical Medicine, doi:10.1093/pcmedi/pbab001.
8.
Ugurel et al., Evaluation of the potency of FDA-approved drugs on wild type and mutant SARS-CoV-2 helicase (Nsp13), International Journal of Biological Macromolecules, doi:10.1016/j.ijbiomac.2020.09.138.
9.
Kumar et al., In silico virtual screening-based study of nutraceuticals predicts the therapeutic potentials of folic acid and its derivatives against COVID-19, VirusDisease, doi:10.1007/s13337-020-00643-6.
10.
Moatasim et al., Potent Antiviral Activity of Vitamin B12 against Severe Acute Respiratory Syndrome Coronavirus 2, Middle East Respiratory Syndrome Coronavirus, and Human Coronavirus 229E, Microorganisms, doi:10.3390/microorganisms11112777.
11.
Zhang et al., Folic acid restricts SARS-CoV-2 invasion by methylating ACE2, Frontiers in Microbiology, doi:10.3389/fmicb.2022.980903.
12.
Galmés et al., Suboptimal Consumption of Relevant Immune System Micronutrients Is Associated with a Worse Impact of COVID-19 in Spanish Populations, Nutrients, doi:10.3390/nu14112254.
13.
Galmés (B) et al., Current State of Evidence: Influence of Nutritional and Nutrigenetic Factors on Immunity in the COVID-19 Pandemic Framework, Nutrients, doi:10.3390/nu12092738.
14.
EFSA, Scientific Opinion on the substantiation of health claims related to folate and blood formation (ID 79), homocysteine metabolism (ID 80), energy-yielding metabolism (ID 90), function of the immune system (ID 91), function of blood vessels (ID 94, 175, 192), cell division (ID 193), and maternal tissue growth during pregnancy (ID 2882) pursuant to Article 13(1) of Regulation (EC) No 1924/2006, EFSA Journal, doi:10.2903/j.efsa.2009.1213.
Eskandari et al., 16 May 2022, peer-reviewed, 1 author.
Contact: veskandari@znu.ac.ir.
In silico studies are an important part of preclinical research, however results may be very different in vivo.
Repurposing the natural compounds as potential therapeutic agents for COVID-19 based on the molecular docking study of the main protease and the receptor-binding domain of spike protein
Journal of Molecular Modeling, doi:10.1007/s00894-022-05138-3
Severe acute respiratory syndrome coronavirus (SARS-CoV-2) enters the cell by interacting with the human angiotensinconverting enzyme 2 (ACE2) receptor through the receptor-binding domain (RBD) of spike (S) protein. In the cell, the viral 3-chymotrypsin-like cysteine protease (3CLpro) enzyme is essential for its life cycle and controls coronavirus replication. Therefore, the S-RBD and 3CLpro are hot targets for drug discovery against SARS-CoV-2. This study was to identify repurposing drugs using in silico screening, docking, and molecular dynamics simulation. The study identified bentiamine, folic acid, benfotiamine, and vitamin B12 against the RBD of S protein and bentiamine, folic acid, fursultiamine, and riboflavin to 3CLpro. The strong and stable binding of these safe and cheap vitamins at the important residues (R403, K417, Y449, Y453, N501, and Y505) in the S-protein-ACE2 interface and 3CLpro binding site residues especially active site residues (His 41 and Cys 145), indicating that they could be valuable repurpose drugs for inhibiting SARS-CoV-2 entry into the host and replication.
Supplementary Information The online version contains supplementary material available at https:// doi. org/ 10. 1007/ s00894-022-05138-3.
Author contribution This manuscript has been done by myself, and there was no coauthor.
Declarations Ethics approval This study does not require ethics approval. Consent to participate This study does not require participant approval. Consent for publication This study does not require publication approval.
Competing interests The author declares no competing interests. Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Abraham, Gready, Optimization of parameters for molecular dynamics simulation using smooth particle-mesh Ewald in GROMACS 4.5, J Comput Chem
Bae, Kamynina, Farinola, Caudill, Stover et al., Provision of folic acid for reducing arsenic toxicity in arsenic-exposed children and adults, Cochrane Database Syst Rev
Ben-Eltriki, Guns, Calcitriol in combination therapy for prostate cancer: pharmacokinetic and pharmacodynamic interactions, J Cancer
Berendsen, Van Der Spoel, Van Drunen, GROMACS: a message-passing parallel molecular dynamics implementation, Comput Phys Commun
Berman, Westbrook, Feng, Gilliland, Bhat et al., The Protein Data Bank, Nucleic Acids Res
Bolton, Wang, Thiessen, Bryant, PubChem: integrated platform of small molecules and biological activities, Annual reports in computational chemistry
Chen, Yiu, Wong, Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL (pro)) structure: virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates, F1000Res
Chikhale, Gurav, Patil, Sinha, Prasad et al., SARS-CoV-2 host entry and replication inhibitors from Indian ginseng: an in-silico approach, J Biomol Struct Dyn
Corrêa, Barros, Fernandes, Sokovic, Bracht et al., A natural food ingredient based on ergosterol: optimization of the extraction from Agaricus blazei, evaluation of bioactive properties and incorporation in yogurts, Food Funct
Dallakyan, Olson, Small-molecule library screening by docking with PyRx, Methods Mol Biol
Fehr, Perlman, Coronaviruses: an overview of their replication and pathogenesis, Methods Mol Biol
Gahlawat, Kumar, Kumar, Sandhu, Singh et al., Structure-based virtual screening to discover potential lead molecules for the SARS-CoV-2 main protease, J Chem Inf Model
Hanwell, Curtis, Lonie, Vandermeersch, Zurek et al., Avogadro: an advanced semantic chemical editor, visualization, and analysis platform, J Cheminform
Heywood, Wood, Majeed, Tumorigenic and toxic effect of O, S-dibenzoyl thiamine hydrochloride in prolonged dietary administration to rats, Toxicol Lett
Khan, Zia, Ashraf, Uddin, Ul-Haq, Identification of chymotrypsin-like protease inhibitors of SARS-CoV-2 via integrated computational approach, J Biomol Struct Dyn
Laskowski, Swindells, LigPlot+: multiple ligandprotein interaction diagrams for drug discovery, J Chem Inf Model
Martoňák, Laio, Parrinello, Predicting crystal structures: the Parrinello-Rahman method revisited, Phys Rev Lett
Morris, Huey, Lindstrom, Sanner, Belew et al., AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility, J Comput Chem
Naqvi, Fatima, Mohammad, Fatima, Singh et al., Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: structural genomics approach, Biochim Biophys Acta (BBA)-Mol Basis Dis
Ravichandran, Coyle, Klenow, Tang, Grubbs et al., Antibody signature induced by SARS-CoV-2 spike protein immunogens in rabbits, Sci Transl Med
Satake, Amano, Okamoto, Calcipotriol and betamethasone dipropionate synergistically enhances the balance between regulatory and proinflammatory T cells in a murine psoriasis model, Sci Rep
Schmid, Eichenberger, Choutko, Riniker, Winger et al., Testing of the GROMOS force-field versions: 54A7 and 54B7, Eur Biophys J
Schoeman, Fielding, Coronavirus envelope protein: current knowledge, Virol J
Schrödinger, None, PyMOL. The PyMOL Molecular Graphics System
Schüttelkopf, Van Aalten, PRODRG: a tool for high-throughput crystallography of protein-ligand complexes, Acta Crystallogr Sect D
Turner, XMGRACE, Version 5.1. 19
Unni, Aouti, Thiyagarajan, Padmanabhan, Identification of a repurposed drug as an inhibitor of spike protein of human coronavirus SARS-CoV-2 by computational methods, J Biosci
Visualizer, Version 4.5. Softw Vis Anal Protein Struct
Vogel, Dali-Youcef, Kaltenbach, Andres, Homocysteine, vitamin B12, folate and cognitive functions: a systematic and critical review of the literature, Int J Clin Pract
Wu, Zhao, Yu, Chen, Song, A new coronavirus associated with human respiratory disease in China, Nature
Yang, Wang, COVID-19: a new challenge for human beings, Cell Mol Immunol
Yoshida, Hishiyama, Igarashi, A novel method for determining total vitamin B1 in processed food enriched with dibenzoyl thiamine, J Jpn Soc Food Sci Technol
DOI record:
{
"DOI": "10.1007/s00894-022-05138-3",
"ISSN": [
"1610-2940",
"0948-5023"
],
"URL": "http://dx.doi.org/10.1007/s00894-022-05138-3",
"alternative-id": [
"5138"
],
"article-number": "153",
"assertion": [
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Received",
"name": "received",
"order": 1,
"value": "23 December 2020"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "26 April 2022"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "First Online",
"name": "first_online",
"order": 3,
"value": "16 May 2022"
},
{
"group": {
"label": "Declarations",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 1
},
{
"group": {
"label": "Ethics approval",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 2,
"value": "This study does not require ethics approval."
},
{
"group": {
"label": "Consent to participate",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 3,
"value": "This study does not require participant approval."
},
{
"group": {
"label": "Consent for publication",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 4,
"value": "This study does not require publication approval."
},
{
"group": {
"label": "Competing interests",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 5,
"value": "The author declares no competing interests."
},
{
"label": "Free to read",
"name": "free",
"value": "This content has been made available to all."
}
],
"author": [
{
"ORCID": "http://orcid.org/0000-0002-4639-504X",
"affiliation": [],
"authenticated-orcid": false,
"family": "Eskandari",
"given": "Vajiheh",
"sequence": "first"
}
],
"container-title": "Journal of Molecular Modeling",
"container-title-short": "J Mol Model",
"content-domain": {
"crossmark-restriction": false,
"domain": [
"link.springer.com"
]
},
"created": {
"date-parts": [
[
2022,
5,
16
]
],
"date-time": "2022-05-16T22:03:00Z",
"timestamp": 1652738580000
},
"deposited": {
"date-parts": [
[
2023,
2,
5
]
],
"date-time": "2023-02-05T09:58:21Z",
"timestamp": 1675591101000
},
"funder": [
{
"name": "I didnt have funding source"
}
],
"indexed": {
"date-parts": [
[
2023,
3,
30
]
],
"date-time": "2023-03-30T16:09:18Z",
"timestamp": 1680192558927
},
"is-referenced-by-count": 8,
"issue": "6",
"issued": {
"date-parts": [
[
2022,
5,
16
]
]
},
"journal-issue": {
"issue": "6",
"published-print": {
"date-parts": [
[
2022,
6
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://www.springer.com/tdm",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2022,
5,
16
]
],
"date-time": "2022-05-16T00:00:00Z",
"timestamp": 1652659200000
}
},
{
"URL": "https://www.springer.com/tdm",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2022,
5,
16
]
],
"date-time": "2022-05-16T00:00:00Z",
"timestamp": 1652659200000
}
}
],
"link": [
{
"URL": "https://link.springer.com/content/pdf/10.1007/s00894-022-05138-3.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://link.springer.com/article/10.1007/s00894-022-05138-3/fulltext.html",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://link.springer.com/content/pdf/10.1007/s00894-022-05138-3.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "297",
"original-title": [],
"prefix": "10.1007",
"published": {
"date-parts": [
[
2022,
5,
16
]
]
},
"published-online": {
"date-parts": [
[
2022,
5,
16
]
]
},
"published-print": {
"date-parts": [
[
2022,
6
]
]
},
"publisher": "Springer Science and Business Media LLC",
"reference": [
{
"key": "5138_CR1",
"unstructured": "Data obtained from https://covid19.who.int/ (World Health Organization)"
},
{
"DOI": "10.1007/978-1-4939-2438-7_1",
"author": "AR Fehr",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "Methods Mol Biol",
"key": "5138_CR2",
"unstructured": "Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 1282:1–23",
"volume": "1282",
"year": "2015"
},
{
"DOI": "10.1038/s41423-020-0407-x",
"author": "P Yang",
"doi-asserted-by": "publisher",
"first-page": "555",
"journal-title": "Cell Mol Immunol",
"key": "5138_CR3",
"unstructured": "Yang P, Wang X (2020) COVID-19: a new challenge for human beings. Cell Mol Immunol 17:555–557",
"volume": "17",
"year": "2020"
},
{
"DOI": "10.1038/s41586-020-2008-3",
"author": "F Wu",
"doi-asserted-by": "publisher",
"first-page": "265",
"journal-title": "Nature",
"key": "5138_CR4",
"unstructured": "Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG et al (2020) A new coronavirus associated with human respiratory disease in China. Nature 579:265–269",
"volume": "579",
"year": "2020"
},
{
"DOI": "10.12688/f1000research.22457.2",
"author": "YW Chen",
"doi-asserted-by": "publisher",
"first-page": "129",
"journal-title": "F1000Res",
"key": "5138_CR5",
"unstructured": "Chen YW, Yiu CB, Wong KY (2020) Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL (pro)) structure: virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates. F1000Res 9:129",
"volume": "9",
"year": "2020"
},
{
"DOI": "10.1080/07391102.2020.1751298",
"doi-asserted-by": "crossref",
"key": "5138_CR6",
"unstructured": "Khan SA, Zia K, Ashraf S, Uddin R, Ul-Haq Z (2020) Identification of chymotrypsin-like protease inhibitors of SARS-CoV-2 via integrated computational approach. J Biomol Struct Dyn 1–10"
},
{
"DOI": "10.1016/j.bbadis.2020.165878",
"doi-asserted-by": "crossref",
"key": "5138_CR7",
"unstructured": "Naqvi AAT, Fatima K, Mohammad T, Fatima U, Singh IK, Singh A, et al (2020) Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: structural genomics approach. Biochim Biophys Acta (BBA)-Mol Basis Dis 165878"
},
{
"DOI": "10.1021/acs.jcim.0c00546",
"doi-asserted-by": "crossref",
"key": "5138_CR8",
"unstructured": "Gahlawat A, Kumar N, Kumar R, Sandhu H, Singh IP, Singh S et al (2020) Structure-based virtual screening to discover potential lead molecules for the SARS-CoV-2 main protease. J Chem Inf Model"
},
{
"DOI": "10.1186/s12985-019-1182-0",
"author": "D Schoeman",
"doi-asserted-by": "publisher",
"first-page": "69",
"journal-title": "Virol J",
"key": "5138_CR9",
"unstructured": "Schoeman D, Fielding BC (2019) Coronavirus envelope protein: current knowledge. Virol J 16:69",
"volume": "16",
"year": "2019"
},
{
"DOI": "10.1126/scitranslmed.abc3539",
"doi-asserted-by": "crossref",
"key": "5138_CR10",
"unstructured": "Ravichandran S, Coyle EM, Klenow L, Tang J, Grubbs G, Liu S et al (2020) Antibody signature induced by SARS-CoV-2 spike protein immunogens in rabbits. Sci Transl Med. 12"
},
{
"DOI": "10.1007/s12038-020-00102-w",
"doi-asserted-by": "crossref",
"key": "5138_CR11",
"unstructured": "Unni S, Aouti S, Thiyagarajan S, Padmanabhan B (2020) Identification of a repurposed drug as an inhibitor of spike protein of human coronavirus SARS-CoV-2 by computational methods. J Biosci 45"
},
{
"DOI": "10.1080/07391102.2020.1778539",
"doi-asserted-by": "crossref",
"key": "5138_CR12",
"unstructured": "Chikhale RV, Gurav SS, Patil RB, Sinha SK, Prasad SK, Shakya A et al (2020) SARS-CoV-2 host entry and replication inhibitors from Indian ginseng: an in-silico approach. J Biomol Struct Dyn. 1–12"
},
{
"DOI": "10.1093/nar/28.1.235",
"author": "HM Berman",
"doi-asserted-by": "publisher",
"first-page": "235",
"journal-title": "Nucleic Acids Res",
"key": "5138_CR13",
"unstructured": "Berman HM, Westbrook J, Feng Z, Gilliland G, Bhat TN, Weissig H et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242",
"volume": "28",
"year": "2000"
},
{
"key": "5138_CR14",
"unstructured": "Visualizer ADS (2017) Version 4.5. Softw Vis Anal Protein Struct"
},
{
"DOI": "10.1002/jcc.21256",
"author": "GM Morris",
"doi-asserted-by": "publisher",
"first-page": "2785",
"journal-title": "J Comput Chem",
"key": "5138_CR15",
"unstructured": "Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS et al (2009) AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 30:2785–2791",
"volume": "30",
"year": "2009"
},
{
"DOI": "10.1016/S1574-1400(08)00012-1",
"doi-asserted-by": "crossref",
"key": "5138_CR16",
"unstructured": "Bolton EE, Wang Y, Thiessen PA, Bryant SH (2008) PubChem: integrated platform of small molecules and biological activities. In: Annual reports in computational chemistry, Elsevier, Vol. 4, pp. 217–41"
},
{
"DOI": "10.1186/1758-2946-4-17",
"author": "MD Hanwell",
"doi-asserted-by": "publisher",
"first-page": "17",
"journal-title": "J Cheminform",
"key": "5138_CR17",
"unstructured": "Hanwell MD, Curtis DE, Lonie DC, Vandermeersch T, Zurek E, Hutchison GR (2012) Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J Cheminform 4:17",
"volume": "4",
"year": "2012"
},
{
"DOI": "10.1007/978-1-4939-2269-7_19",
"author": "S Dallakyan",
"doi-asserted-by": "publisher",
"first-page": "243",
"journal-title": "Methods Mol Biol",
"key": "5138_CR18",
"unstructured": "Dallakyan S, Olson AJ (2015) Small-molecule library screening by docking with PyRx. Methods Mol Biol 1263:243–250",
"volume": "1263",
"year": "2015"
},
{
"DOI": "10.1021/ci200227u",
"author": "RA Laskowski",
"doi-asserted-by": "publisher",
"first-page": "2778",
"journal-title": "J Chem Inf Model",
"key": "5138_CR19",
"unstructured": "Laskowski RA, Swindells MB (2011) LigPlot+: multiple ligand-protein interaction diagrams for drug discovery. J Chem Inf Model 51:2778–2786",
"volume": "51",
"year": "2011"
},
{
"DOI": "10.1016/0010-4655(95)00042-E",
"author": "HJ Berendsen",
"doi-asserted-by": "publisher",
"first-page": "43",
"journal-title": "Comput Phys Commun",
"key": "5138_CR20",
"unstructured": "Berendsen HJ, van der Spoel D, van Drunen R (1995) GROMACS: a message-passing parallel molecular dynamics implementation. Comput Phys Commun 91:43–56",
"volume": "91",
"year": "1995"
},
{
"DOI": "10.1007/s00249-011-0700-9",
"author": "N Schmid",
"doi-asserted-by": "publisher",
"first-page": "843",
"journal-title": "Eur Biophys J",
"key": "5138_CR21",
"unstructured": "Schmid N, Eichenberger AP, Choutko A, Riniker S, Winger M, Mark AE, van Gunsteren Definition WF (2011) Testing of the GROMOS force-field versions: 54A7 and 54B7. Eur Biophys J 40:843–856",
"volume": "40",
"year": "2011"
},
{
"DOI": "10.1107/S0907444904011679",
"author": "AW Schüttelkopf",
"doi-asserted-by": "publisher",
"first-page": "1355",
"journal-title": "Acta Crystallogr Sect D",
"key": "5138_CR22",
"unstructured": "Schüttelkopf AW, van Aalten DMF (2004) PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr Sect D 60:1355–1363",
"volume": "60",
"year": "2004"
},
{
"DOI": "10.1103/PhysRevLett.90.075503",
"author": "R Martoňák",
"doi-asserted-by": "publisher",
"journal-title": "Phys Rev Lett",
"key": "5138_CR23",
"unstructured": "Martoňák R, Laio A, Parrinello M (2003) Predicting crystal structures: the Parrinello-Rahman method revisited. Phys Rev Lett 7:075503",
"volume": "7",
"year": "2003"
},
{
"DOI": "10.1002/jcc.21773",
"author": "MJ Abraham",
"doi-asserted-by": "publisher",
"first-page": "2031",
"journal-title": "J Comput Chem",
"key": "5138_CR24",
"unstructured": "Abraham MJ, Gready JE (2011) Optimization of parameters for molecular dynamics simulation using smooth particle-mesh Ewald in GROMACS 4.5. J Comput Chem 32:2031–2040",
"volume": "32",
"year": "2011"
},
{
"key": "5138_CR25",
"unstructured": "Turner PJ (2005) XMGRACE, Version 5.1. 19. Center for Coastal and Land-Margin Research, Oregon Graduate Institute of Science and Technology, Beaverton, OR"
},
{
"key": "5138_CR26",
"unstructured": "Schrödinger L (2017) PyMOL. The PyMOL Molecular Graphics System, Version. 2"
},
{
"DOI": "10.3136/nskkk.55.421",
"doi-asserted-by": "crossref",
"key": "5138_CR27",
"unstructured": "Yoshida M, Hishiyama T, Igarashi T (2008) A novel method for determining total vitamin B1 in processed food enriched with dibenzoyl thiamine. J Jpn Soc Food Sci Technol (Japan)"
},
{
"DOI": "10.1016/0378-4274(85)90184-5",
"author": "R Heywood",
"doi-asserted-by": "publisher",
"first-page": "53",
"journal-title": "Toxicol Lett",
"key": "5138_CR28",
"unstructured": "Heywood R, Wood J, Majeed S (1985) Tumorigenic and toxic effect of O, S-dibenzoyl thiamine hydrochloride in prolonged dietary administration to rats. Toxicol Lett 26:53–58",
"volume": "26",
"year": "1985"
},
{
"DOI": "10.1038/s41598-018-37186-2",
"author": "K Satake",
"doi-asserted-by": "crossref",
"first-page": "1",
"journal-title": "Sci Rep",
"key": "5138_CR29",
"unstructured": "Satake K, Amano T, Okamoto T (2019) Calcipotriol and betamethasone dipropionate synergistically enhances the balance between regulatory and proinflammatory T cells in a murine psoriasis model. Sci Rep 9:1–11",
"volume": "9",
"year": "2019"
},
{
"DOI": "10.7150/jca.13470",
"author": "M Ben-Eltriki",
"doi-asserted-by": "publisher",
"first-page": "391",
"journal-title": "J Cancer",
"key": "5138_CR30",
"unstructured": "Ben-Eltriki M, Deb S, Guns EST (2016) Calcitriol in combination therapy for prostate cancer: pharmacokinetic and pharmacodynamic interactions. J Cancer 7:391",
"volume": "7",
"year": "2016"
},
{
"DOI": "10.1002/14651858.CD012649",
"doi-asserted-by": "crossref",
"key": "5138_CR31",
"unstructured": "Bae S, Kamynina E, Farinola AF, Caudill MA, Stover PJ, Cassano PA, et al (2017) Provision of folic acid for reducing arsenic toxicity in arsenic‐exposed children and adults. Cochrane Database Syst Rev 2017"
},
{
"DOI": "10.1039/C7FO02007D",
"author": "RC Corrêa",
"doi-asserted-by": "publisher",
"first-page": "1465",
"journal-title": "Food Funct",
"key": "5138_CR32",
"unstructured": "Corrêa RC, Barros L, Fernandes Â, Sokovic M, Bracht A, Peralta RM et al (2018) A natural food ingredient based on ergosterol: optimization of the extraction from Agaricus blazei, evaluation of bioactive properties and incorporation in yogurts. Food Funct 9:1465–1474",
"volume": "9",
"year": "2018"
},
{
"DOI": "10.1111/j.1742-1241.2009.02026.x",
"author": "T Vogel",
"doi-asserted-by": "publisher",
"first-page": "1061",
"journal-title": "Int J Clin Pract",
"key": "5138_CR33",
"unstructured": "Vogel T, Dali-Youcef N, Kaltenbach G, Andres E (2009) Homocysteine, vitamin B12, folate and cognitive functions: a systematic and critical review of the literature. Int J Clin Pract 63:1061–1067",
"volume": "63",
"year": "2009"
}
],
"reference-count": 33,
"references-count": 33,
"relation": {},
"resource": {
"primary": {
"URL": "https://link.springer.com/10.1007/s00894-022-05138-3"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"Computational Theory and Mathematics",
"Inorganic Chemistry",
"Organic Chemistry",
"Physical and Theoretical Chemistry",
"Computer Science Applications",
"Catalysis"
],
"subtitle": [],
"title": "Repurposing the natural compounds as potential therapeutic agents for COVID-19 based on the molecular docking study of the main protease and the receptor-binding domain of spike protein",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.1007/springer_crossmark_policy",
"volume": "28"
}