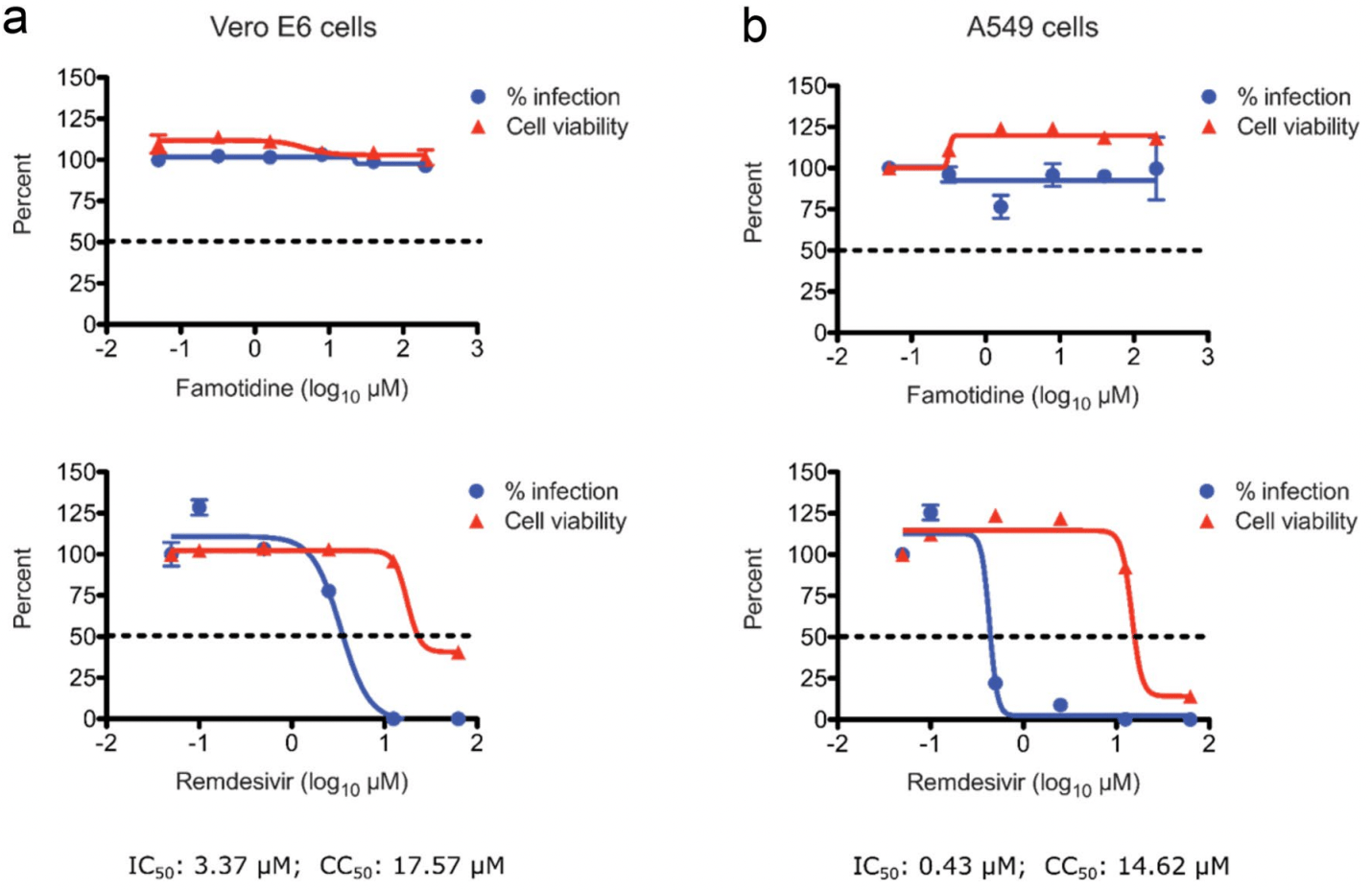
The in-vitro effect of famotidine on SARS-CoV-2 proteases and virus replication
et al., Scientific Reports, doi:10.1038/s41598-021-84782-w, Mar 2021
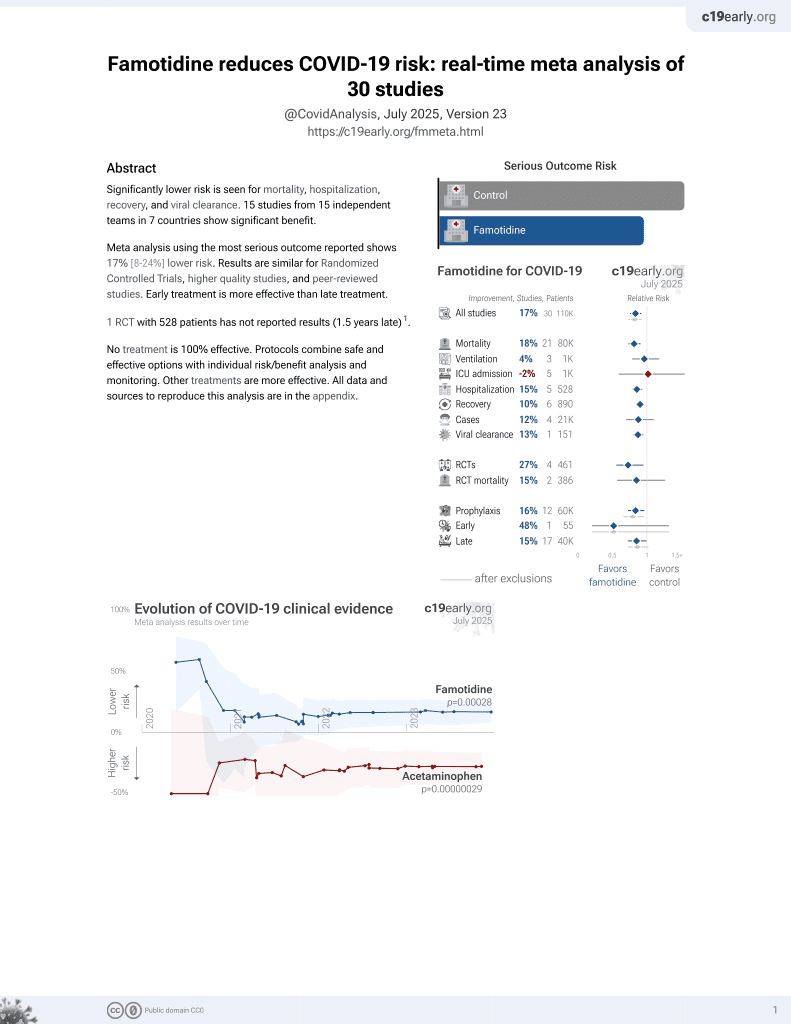
Famotidine for COVID-19
28th treatment shown to reduce risk in
October 2021, now with p = 0.00072 from 29 studies, recognized in 2 countries.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
In vitro study showing that famotidine does not bind or inhibit 3CLpro and PLpro, and no direct antiviral activity was observed with concentrations up to 200µM in Vero E6 and A549 cells.
2 preclinical studies support the efficacy of famotidine for COVID-19:
Loffredo et al., 8 Mar 2021, peer-reviewed, 12 authors.
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
The in-vitro effect of famotidine on SARS-CoV-2 proteases and virus replication
Scientific Reports, doi:10.1038/s41598-021-84782-w
The lack of coronavirus-specific antiviral drugs has instigated multiple drug repurposing studies to redirect previously approved medicines for the treatment of SARS-CoV-2, the coronavirus behind the ongoing COVID-19 pandemic. A recent, large-scale, retrospective clinical study showed that famotidine, when administered at a high dose to hospitalized COVID-19 patients, reduced the rates of intubation and mortality. A separate, patient-reported study associated famotidine use with improvements in mild to moderate symptoms such as cough and shortness of breath. While a prospective, multi-center clinical study is ongoing, two parallel in silico studies have proposed one of the two SARS-CoV-2 proteases, 3CL pro or PL pro , as potential molecular targets of famotidine activity; however, this remains to be experimentally validated. In this report, we systematically analyzed the effect of famotidine on viral proteases and virus replication. Leveraging a series of biophysical and enzymatic assays, we show that famotidine neither binds with nor inhibits the functions of 3CL pro and PL pro . Similarly, no direct antiviral activity of famotidine was observed at concentrations of up to 200 µM, when tested against SARS-CoV-2 in two different cell lines, including a human cell line originating from lungs, a primary target of COVID-19. These results rule out famotidine as a direct-acting inhibitor of SARS-CoV-2 replication and warrant further investigation of its molecular mechanism of action in the context of COVID-19.
Author contributions
Competing interests The authors declare no competing interests.
References
Baez-Santos, Mielech, Deng, Baker, Mesecar, Catalytic function and substrate specificity of the papainlike protease domain of nsp3 from the Middle East respiratory syndrome coronavirus, J. Virol, doi:10.1128/JVI.01294-14
Barretto, The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity, J. Virol, doi:10.1128/JVI.79.24.15189-15198.2005
Beigel, Remdesivir for the treatment of Covid-19 -preliminary report, N. Engl. J. Med, doi:10.1056/NEJMoa2007764
Bhatraju, Covid-19 in critically Ill patients in the seattle region-case series, N. Engl. J. Med, doi:10.1056/NEJMoa2004500
Boulware, A randomized trial of hydroxychloroquine as postexposure prophylaxis for covid-19, N. Engl. J. Med, doi:10.1056/NEJMoa2016638
Cantu, Korek, Central nervous system reactions to histamine-2 receptor blockers, Ann. Intern. Med, doi:10.7326/0003-4819-114-12-1027
Cao, A trial of lopinavir-ritonavir in adults hospitalized with severe covid-19, N. Engl. J. Med, doi:10.1056/NEJMoa2001282
Carsana, Pulmonary post-mortem findings in a series of COVID-19 cases from northern Italy: a two-centre descriptive study, Lancet Infect. Dis, doi:10.1016/S1473-3099(20)30434-5
Corman, Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR, Euro Surveill, doi:10.2807/1560-7917
Freedberg, hospitalized COVID-19 patients: a propensity score matched retrospective cohort study, doi:10.1053/j.gastro.2020.05.053
Freitas, Characterization and noncovalent inhibition of the deubiquitinase and deISGylase activity of SARS-CoV-2 papain-like protease, ACS Infect. Dis, doi:10.1021/acsinfecdis.0c00168
Harrison, Coronavirus puts drug repurposing on the fast track, Nat. Biotechnol, doi:10.1038/d41587-020-00003-1
Hu, Mast cell-induced lung injury in mice infected with H5N1 influenza virus, J. Virol, doi:10.1128/JVI.06053-11
Jacobs, Discovery, synthesis, and structure-based optimization of a series of N-(tert-butyl)-2-(N-arylamido)-2-(pyridin-3-yl) acetamides (ML188) as potent noncovalent small molecule inhibitors of the severe acute respiratory syndrome coronavirus (SARS-CoV) 3CL protease, J. Med. Chem, doi:10.1021/jm301580n
Jacobs, Probe Reports from the NIH Molecular Libraries Program
Janowitz, Famotidine use and quantitative symptom tracking for COVID-19 in non-hospitalised patients: a case series, doi:10.1136/gutjnl-2020-321852
Kallal, Lee, Thrombotic thrombo-cytopenic purpura associated with histamine H2-receptor antagonist therapy, West. J. Med
Keithley, Histamine H2-receptor antagonists, Nurs Clin North Am
Kirch, Halabi, Linde, Santos, Ohnhaus, Negative effects of famotidine on cardiac performance assessed by noninvasive hemodynamic measurements, Gastroenterology, doi:10.1016/0016-5085(89)90503-9
Kung, Jura, Structural basis for the non-catalytic functions of protein kinases, Structure, doi:10.1016/j.str.2015.10.020
Lee, Wang, Famotidine-induced retinopathy, Eye (Lond), doi:10.1038/sj.eye.6701839
Lindner, The papain-like protease from the severe acute respiratory syndrome coronavirus is a deubiquitinating enzyme, J. Virol, doi:10.1128/JVI.79.24.15199-15208.2005
Lu, Structure-based drug design and structural biology study of novel nonpeptide inhibitors of severe acute respiratory syndrome coronavirus main protease, J. Med. Chem, doi:10.1021/jm060207o
Ma, Expression of SARS-CoV-2 receptor ACE2 and TMPRSS2 in human primary conjunctival and pterygium cell lines and in mouse cornea, Eye, doi:10.1038/s41433-020-0939-4
Marshall, Portales-Cervantes, Leong, Mast cell responses to viruses and pathogen products, Int. J. Mol. Sci, doi:10.3390/ijms20174241
Muramatsu, SARS-CoV 3CL protease cleaves its C-terminal autoprocessing site by novel subsite cooperativity, Proc. Natl. Acad. Sci, doi:10.1073/pnas.1601327113
Ogando, SARS-coronavirus-2 replication in Vero E6 cells: replication kinetics, rapid adaptation and cytopathology, J. Gen. Virol, doi:10.1099/jgv.0.001453
Palacio-Rodriguez, Lans, Cavasotto, Cossio, Exponential consensus ranking improves the outcome in docking and receptor ensemble docking, Sci. Rep, doi:10.1038/s41598-019-41594-3
Ratia, A noncovalent class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication, Proc. Natl. Acad. Sci, doi:10.1073/pnas.0805240105
Shaffer, 15 drugs being tested to treat COVID-19 and how they would work, Nat. Med, doi:10.1038/d41591-020-00019-9
Sun, Li, Yates, Fernig, SimpleDSFviewer: a tool to analyze and view differential scanning fluorimetry data for characterizing protein thermal stability and interactions, Protein Sci, doi:10.1002/pro.3703
Tay, Poh, Renia, Macary, Ng, The trinity of COVID-19: immunity, inflammation and intervention, Nat. Rev. Immunol, doi:10.1038/s41577-020-0311-8
Thangam, The role of histamine and histamine receptors in mast cell-mediated allergy and inflammation: the hunt for new therapeutic targets, Front. Immunol, doi:10.3389/fimmu.2018.01873
Tian, Clinical characteristics and risk factors associated with COVID-19 disease severity in patients with cancer in Wuhan, China: a multicentre, retrospective, cohort study, Lancet Oncol, doi:10.1016/S1470-2045(20)30309-0
Tomar, Ligand-induced dimerization of middle east respiratory syndrome (MERS) coronavirus nsp5 protease (3CLpro): implications for nsp5 regulation and the development of antivirals, J. Biol. Chem, doi:10.1074/jbc.M115.651463
Van Hemert, SARS-coronavirus replication/transcription complexes are membrane-protected and need a host factor for activity in vitro, PLoS Pathog, doi:10.1371/journal.ppat.1000054
Wu, Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods, Acta Pharm. Sin. B, doi:10.1016/j.apsb.2020.02.008
Xie, A nanoluciferase SARS-CoV-2 for rapid neutralization testing and screening of anti-infective drugs for COVID-19, doi:10.1101/2020.06.22.165712
Yang, Clinical characteristics, outcomes, and risk factors for mortality in patients with cancer and COVID-19 in Hubei, China: a multicentre, retrospective, cohort study, Lancet Oncol, doi:10.1016/S1470-2045(20)30310-7
Ye, Wang, Mao, The pathogenesis and treatment of the `Cytokine Storm' in COVID-19, J. Infect, doi:10.1016/j.jinf.2020.03.037
Zarnegar, Influenza infection in mice induces accumulation of lung mast cells through the recruitment and maturation of mast cell progenitors, Front. Immunol, doi:10.3389/fimmu.2017.00310
Zhang, Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved alpha-ketoamide inhibitors, Science, doi:10.1126/science.abb3405
DOI record:
{
"DOI": "10.1038/s41598-021-84782-w",
"ISSN": [
"2045-2322"
],
"URL": "http://dx.doi.org/10.1038/s41598-021-84782-w",
"abstract": "<jats:title>Abstract</jats:title><jats:p>The lack of coronavirus-specific antiviral drugs has instigated multiple drug repurposing studies to redirect previously approved medicines for the treatment of SARS-CoV-2, the coronavirus behind the ongoing COVID-19 pandemic. A recent, large-scale, retrospective clinical study showed that famotidine, when administered at a high dose to hospitalized COVID-19 patients, reduced the rates of intubation and mortality. A separate, patient-reported study associated famotidine use with improvements in mild to moderate symptoms such as cough and shortness of breath. While a prospective, multi-center clinical study is ongoing, two parallel in silico studies have proposed one of the two SARS-CoV-2 proteases, 3CL<jats:sup>pro</jats:sup> or PL<jats:sup>pro</jats:sup>, as potential molecular targets of famotidine activity; however, this remains to be experimentally validated. In this report, we systematically analyzed the effect of famotidine on viral proteases and virus replication. Leveraging a series of biophysical and enzymatic assays, we show that famotidine neither binds with nor inhibits the functions of 3CL<jats:sup>pro</jats:sup> and PL<jats:sup>pro</jats:sup>. Similarly, no direct antiviral activity of famotidine was observed at concentrations of up to 200 µM, when tested against SARS-CoV-2 in two different cell lines, including a human cell line originating from lungs, a primary target of COVID-19. These results rule out famotidine as a direct-acting inhibitor of SARS-CoV-2 replication and warrant further investigation of its molecular mechanism of action in the context of COVID-19.</jats:p>",
"alternative-id": [
"84782"
],
"article-number": "5433",
"assertion": [
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Received",
"name": "received",
"order": 1,
"value": "2 November 2020"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "1 February 2021"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "First Online",
"name": "first_online",
"order": 3,
"value": "8 March 2021"
},
{
"group": {
"label": "Competing interests",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 1,
"value": "The authors declare no competing interests."
}
],
"author": [
{
"ORCID": "http://orcid.org/0000-0002-5003-2081",
"affiliation": [],
"authenticated-orcid": false,
"family": "Loffredo",
"given": "Madeline",
"sequence": "first"
},
{
"affiliation": [],
"family": "Lucero",
"given": "Hector",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Chen",
"given": "Da-Yuan",
"sequence": "additional"
},
{
"affiliation": [],
"family": "O’Connell",
"given": "Aoife",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Bergqvist",
"given": "Simon",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Munawar",
"given": "Ahmad",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Bandara",
"given": "Asanga",
"sequence": "additional"
},
{
"affiliation": [],
"family": "De Graef",
"given": "Steff",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-1360-0852",
"affiliation": [],
"authenticated-orcid": false,
"family": "Weeks",
"given": "Stephen D.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Douam",
"given": "Florian",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Saeed",
"given": "Mohsan",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0001-9617-9460",
"affiliation": [],
"authenticated-orcid": false,
"family": "Munawar",
"given": "Ali H.",
"sequence": "additional"
}
],
"container-title": [
"Scientific Reports"
],
"content-domain": {
"crossmark-restriction": false,
"domain": [
"link.springer.com"
]
},
"created": {
"date-parts": [
[
2021,
3,
8
]
],
"date-time": "2021-03-08T11:03:05Z",
"timestamp": 1615201385000
},
"deposited": {
"date-parts": [
[
2021,
12,
2
]
],
"date-time": "2021-12-02T16:12:26Z",
"timestamp": 1638461546000
},
"indexed": {
"date-parts": [
[
2021,
12,
9
]
],
"date-time": "2021-12-09T08:11:14Z",
"timestamp": 1639037474248
},
"is-referenced-by-count": 6,
"issn-type": [
{
"type": "electronic",
"value": "2045-2322"
}
],
"issue": "1",
"issued": {
"date-parts": [
[
2021,
3,
8
]
]
},
"journal-issue": {
"issue": "1",
"published-print": {
"date-parts": [
[
2021,
12
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2021,
3,
8
]
],
"date-time": "2021-03-08T00:00:00Z",
"timestamp": 1615161600000
}
},
{
"URL": "https://creativecommons.org/licenses/by/4.0",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2021,
3,
8
]
],
"date-time": "2021-03-08T00:00:00Z",
"timestamp": 1615161600000
}
}
],
"link": [
{
"URL": "http://www.nature.com/articles/s41598-021-84782-w.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "http://www.nature.com/articles/s41598-021-84782-w",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "http://www.nature.com/articles/s41598-021-84782-w.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "297",
"original-title": [],
"prefix": "10.1038",
"published": {
"date-parts": [
[
2021,
3,
8
]
]
},
"published-online": {
"date-parts": [
[
2021,
3,
8
]
]
},
"published-print": {
"date-parts": [
[
2021,
12
]
]
},
"publisher": "Springer Science and Business Media LLC",
"reference": [
{
"DOI": "10.1038/d41587-020-00003-1",
"author": "C Harrison",
"doi-asserted-by": "publisher",
"first-page": "379",
"journal-title": "Nat. Biotechnol.",
"key": "84782_CR1",
"unstructured": "Harrison, C. Coronavirus puts drug repurposing on the fast track. Nat. Biotechnol. 38, 379–381. https://doi.org/10.1038/d41587-020-00003-1 (2020).",
"volume": "38",
"year": "2020"
},
{
"DOI": "10.1056/NEJMoa2007764",
"author": "JH Beigel",
"doi-asserted-by": "publisher",
"journal-title": "N. Engl. J. Med.",
"key": "84782_CR2",
"unstructured": "Beigel, J. H. et al. Remdesivir for the treatment of Covid-19 - preliminary report. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2007764 (2020).",
"year": "2020"
},
{
"DOI": "10.1056/NEJMoa2016638",
"author": "DR Boulware",
"doi-asserted-by": "publisher",
"journal-title": "N. Engl. J. Med.",
"key": "84782_CR3",
"unstructured": "Boulware, D. R. et al. A randomized trial of hydroxychloroquine as postexposure prophylaxis for covid-19. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2016638 (2020).",
"year": "2020"
},
{
"DOI": "10.1056/NEJMoa2001282",
"author": "B Cao",
"doi-asserted-by": "publisher",
"first-page": "1787",
"journal-title": "N. Engl. J. Med.",
"key": "84782_CR4",
"unstructured": "Cao, B. et al. A trial of lopinavir-ritonavir in adults hospitalized with severe covid-19. N. Engl. J. Med. 382, 1787–1799. https://doi.org/10.1056/NEJMoa2001282 (2020).",
"volume": "382",
"year": "2020"
},
{
"author": "JK Keithley",
"first-page": "361",
"journal-title": "Nurs Clin North Am",
"key": "84782_CR5",
"unstructured": "Keithley, J. K. Histamine H2-receptor antagonists. Nurs Clin North Am 26, 361–373 (1991).",
"volume": "26",
"year": "1991"
},
{
"DOI": "10.1053/j.gastro.2020.05.053",
"author": "DE Freedberg",
"doi-asserted-by": "publisher",
"journal-title": "Gastroenterology",
"key": "84782_CR6",
"unstructured": "Freedberg, D. E. et al. Famotidine use is associated with improved clinical outcomes in hospitalized COVID-19 patients: a propensity score matched retrospective cohort study. Gastroenterology https://doi.org/10.1053/j.gastro.2020.05.053 (2020).",
"year": "2020"
},
{
"DOI": "10.1136/gutjnl-2020-321852",
"author": "T Janowitz",
"doi-asserted-by": "publisher",
"journal-title": "Gut",
"key": "84782_CR7",
"unstructured": "Janowitz, T. et al. Famotidine use and quantitative symptom tracking for COVID-19 in non-hospitalised patients: a case series. Gut https://doi.org/10.1136/gutjnl-2020-321852 (2020).",
"year": "2020"
},
{
"DOI": "10.1016/j.apsb.2020.02.008",
"author": "C Wu",
"doi-asserted-by": "publisher",
"journal-title": "Acta Pharm. Sin. B",
"key": "84782_CR8",
"unstructured": "Wu, C. et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm. Sin. B https://doi.org/10.1016/j.apsb.2020.02.008 (2020).",
"year": "2020"
},
{
"DOI": "10.1038/d41591-020-00019-9",
"author": "L Shaffer",
"doi-asserted-by": "publisher",
"journal-title": "Nat. Med.",
"key": "84782_CR9",
"unstructured": "Shaffer, L. 15 drugs being tested to treat COVID-19 and how they would work. Nat. Med. https://doi.org/10.1038/d41591-020-00019-9 (2020).",
"year": "2020"
},
{
"DOI": "10.1126/science.abb3405",
"author": "L Zhang",
"doi-asserted-by": "publisher",
"first-page": "409",
"journal-title": "Science",
"key": "84782_CR10",
"unstructured": "Zhang, L. et al. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved alpha-ketoamide inhibitors. Science 368, 409–412. https://doi.org/10.1126/science.abb3405 (2020).",
"volume": "368",
"year": "2020"
},
{
"DOI": "10.1073/pnas.1601327113",
"author": "T Muramatsu",
"doi-asserted-by": "publisher",
"first-page": "12997",
"journal-title": "Proc. Natl. Acad. Sci. USA",
"key": "84782_CR11",
"unstructured": "Muramatsu, T. et al. SARS-CoV 3CL protease cleaves its C-terminal autoprocessing site by novel subsite cooperativity. Proc. Natl. Acad. Sci. USA 113, 12997–13002. https://doi.org/10.1073/pnas.1601327113 (2016).",
"volume": "113",
"year": "2016"
},
{
"DOI": "10.1074/jbc.M115.651463",
"author": "S Tomar",
"doi-asserted-by": "publisher",
"first-page": "19403",
"journal-title": "J. Biol. Chem.",
"key": "84782_CR12",
"unstructured": "Tomar, S. et al. Ligand-induced dimerization of middle east respiratory syndrome (MERS) coronavirus nsp5 protease (3CLpro): implications for nsp5 regulation and the development of antivirals. J. Biol. Chem. 290, 19403–19422. https://doi.org/10.1074/jbc.M115.651463 (2015).",
"volume": "290",
"year": "2015"
},
{
"DOI": "10.1128/JVI.01294-14",
"author": "YM Baez-Santos",
"doi-asserted-by": "publisher",
"first-page": "12511",
"journal-title": "J. Virol.",
"key": "84782_CR13",
"unstructured": "Baez-Santos, Y. M., Mielech, A. M., Deng, X., Baker, S. & Mesecar, A. D. Catalytic function and substrate specificity of the papain-like protease domain of nsp3 from the Middle East respiratory syndrome coronavirus. J. Virol. 88, 12511–12527. https://doi.org/10.1128/JVI.01294-14 (2014).",
"volume": "88",
"year": "2014"
},
{
"DOI": "10.1021/jm301580n",
"author": "J Jacobs",
"doi-asserted-by": "publisher",
"first-page": "534",
"journal-title": "J. Med. Chem.",
"key": "84782_CR14",
"unstructured": "Jacobs, J. et al. Discovery, synthesis, and structure-based optimization of a series of N-(tert-butyl)-2-(N-arylamido)-2-(pyridin-3-yl) acetamides (ML188) as potent noncovalent small molecule inhibitors of the severe acute respiratory syndrome coronavirus (SARS-CoV) 3CL protease. J. Med. Chem. 56, 534–546. https://doi.org/10.1021/jm301580n (2013).",
"volume": "56",
"year": "2013"
},
{
"key": "84782_CR15",
"unstructured": "Jacobs, J. et al. Probe Reports from the NIH Molecular Libraries Program (2010)."
},
{
"DOI": "10.1073/pnas.0805240105",
"author": "K Ratia",
"doi-asserted-by": "publisher",
"first-page": "16119",
"journal-title": "Proc. Natl. Acad. Sci. USA",
"key": "84782_CR16",
"unstructured": "Ratia, K. et al. A noncovalent class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication. Proc. Natl. Acad. Sci. USA 105, 16119–16124. https://doi.org/10.1073/pnas.0805240105 (2008).",
"volume": "105",
"year": "2008"
},
{
"DOI": "10.1021/jm060207o",
"author": "IL Lu",
"doi-asserted-by": "publisher",
"first-page": "5154",
"journal-title": "J. Med. Chem.",
"key": "84782_CR17",
"unstructured": "Lu, I. L. et al. Structure-based drug design and structural biology study of novel nonpeptide inhibitors of severe acute respiratory syndrome coronavirus main protease. J. Med. Chem. 49, 5154–5161. https://doi.org/10.1021/jm060207o (2006).",
"volume": "49",
"year": "2006"
},
{
"DOI": "10.1128/JVI.79.24.15199-15208.2005",
"author": "HA Lindner",
"doi-asserted-by": "publisher",
"first-page": "15199",
"journal-title": "J. Virol.",
"key": "84782_CR18",
"unstructured": "Lindner, H. A. et al. The papain-like protease from the severe acute respiratory syndrome coronavirus is a deubiquitinating enzyme. J. Virol. 79, 15199–15208. https://doi.org/10.1128/JVI.79.24.15199-15208.2005 (2005).",
"volume": "79",
"year": "2005"
},
{
"DOI": "10.1128/JVI.79.24.15189-15198.2005",
"author": "N Barretto",
"doi-asserted-by": "publisher",
"first-page": "15189",
"journal-title": "J. Virol.",
"key": "84782_CR19",
"unstructured": "Barretto, N. et al. The papain-like protease of severe acute respiratory syndrome coronavirus has deubiquitinating activity. J. Virol. 79, 15189–15198. https://doi.org/10.1128/JVI.79.24.15189-15198.2005 (2005).",
"volume": "79",
"year": "2005"
},
{
"DOI": "10.1371/journal.ppat.1000054",
"author": "MJ van Hemert",
"doi-asserted-by": "publisher",
"first-page": "e1000054",
"journal-title": "PLoS Pathog.",
"key": "84782_CR20",
"unstructured": "van Hemert, M. J. et al. SARS-coronavirus replication/transcription complexes are membrane-protected and need a host factor for activity in vitro. PLoS Pathog. 4, e1000054. https://doi.org/10.1371/journal.ppat.1000054 (2008).",
"volume": "4",
"year": "2008"
},
{
"DOI": "10.1021/acsinfecdis.0c00168",
"author": "BT Freitas",
"doi-asserted-by": "publisher",
"journal-title": "ACS Infect. Dis.",
"key": "84782_CR21",
"unstructured": "Freitas, B. T. et al. Characterization and noncovalent inhibition of the deubiquitinase and deISGylase activity of SARS-CoV-2 papain-like protease. ACS Infect. Dis. https://doi.org/10.1021/acsinfecdis.0c00168 (2020).",
"year": "2020"
},
{
"DOI": "10.1016/j.str.2015.10.020",
"author": "JE Kung",
"doi-asserted-by": "publisher",
"first-page": "7",
"journal-title": "Structure",
"key": "84782_CR22",
"unstructured": "Kung, J. E. & Jura, N. Structural basis for the non-catalytic functions of protein kinases. Structure 24, 7–24. https://doi.org/10.1016/j.str.2015.10.020 (2016).",
"volume": "24",
"year": "2016"
},
{
"DOI": "10.1038/s41433-020-0939-4",
"author": "D Ma",
"doi-asserted-by": "publisher",
"first-page": "1212",
"journal-title": "Eye",
"key": "84782_CR23",
"unstructured": "Ma, D. et al. Expression of SARS-CoV-2 receptor ACE2 and TMPRSS2 in human primary conjunctival and pterygium cell lines and in mouse cornea. Eye 34, 1212–1219. https://doi.org/10.1038/s41433-020-0939-4 (2020).",
"volume": "34",
"year": "2020"
},
{
"DOI": "10.1101/2020.06.22.165712",
"author": "X Xie",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "84782_CR24",
"unstructured": "Xie, X. et al. A nanoluciferase SARS-CoV-2 for rapid neutralization testing and screening of anti-infective drugs for COVID-19. bioRxiv https://doi.org/10.1101/2020.06.22.165712 (2020).",
"year": "2020"
},
{
"DOI": "10.1038/s41598-019-41594-3",
"author": "K Palacio-Rodriguez",
"doi-asserted-by": "publisher",
"first-page": "5142",
"journal-title": "Sci. Rep.",
"key": "84782_CR25",
"unstructured": "Palacio-Rodriguez, K., Lans, I., Cavasotto, C. N. & Cossio, P. Exponential consensus ranking improves the outcome in docking and receptor ensemble docking. Sci. Rep. 9, 5142. https://doi.org/10.1038/s41598-019-41594-3 (2019).",
"volume": "9",
"year": "2019"
},
{
"DOI": "10.1038/s41577-020-0311-8",
"author": "MZ Tay",
"doi-asserted-by": "publisher",
"first-page": "363",
"journal-title": "Nat. Rev. Immunol.",
"key": "84782_CR26",
"unstructured": "Tay, M. Z., Poh, C. M., Renia, L., MacAry, P. A. & Ng, L. F. P. The trinity of COVID-19: immunity, inflammation and intervention. Nat. Rev. Immunol. 20, 363–374. https://doi.org/10.1038/s41577-020-0311-8 (2020).",
"volume": "20",
"year": "2020"
},
{
"DOI": "10.1016/j.jinf.2020.03.037",
"author": "Q Ye",
"doi-asserted-by": "publisher",
"first-page": "607",
"journal-title": "J. Infect.",
"key": "84782_CR27",
"unstructured": "Ye, Q., Wang, B. & Mao, J. The pathogenesis and treatment of the `Cytokine Storm’ in COVID-19. J. Infect. 80, 607–613. https://doi.org/10.1016/j.jinf.2020.03.037 (2020).",
"volume": "80",
"year": "2020"
},
{
"DOI": "10.1016/S1473-3099(20)30434-5",
"author": "L Carsana",
"doi-asserted-by": "publisher",
"journal-title": "Lancet Infect. Dis.",
"key": "84782_CR28",
"unstructured": "Carsana, L. et al. Pulmonary post-mortem findings in a series of COVID-19 cases from northern Italy: a two-centre descriptive study. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(20)30434-5 (2020).",
"year": "2020"
},
{
"DOI": "10.1056/NEJMoa2004500",
"author": "PK Bhatraju",
"doi-asserted-by": "publisher",
"first-page": "2012",
"journal-title": "N. Engl. J. Med.",
"key": "84782_CR29",
"unstructured": "Bhatraju, P. K. et al. Covid-19 in critically Ill patients in the seattle region—case series. N. Engl. J. Med. 382, 2012–2022. https://doi.org/10.1056/NEJMoa2004500 (2020).",
"volume": "382",
"year": "2020"
},
{
"DOI": "10.1016/S1470-2045(20)30309-0",
"author": "J Tian",
"doi-asserted-by": "publisher",
"journal-title": "Lancet Oncol.",
"key": "84782_CR30",
"unstructured": "Tian, J. et al. Clinical characteristics and risk factors associated with COVID-19 disease severity in patients with cancer in Wuhan, China: a multicentre, retrospective, cohort study. Lancet Oncol. https://doi.org/10.1016/S1470-2045(20)30309-0 (2020).",
"year": "2020"
},
{
"DOI": "10.1016/S1470-2045(20)30310-7",
"author": "K Yang",
"doi-asserted-by": "publisher",
"journal-title": "Lancet Oncol.",
"key": "84782_CR31",
"unstructured": "Yang, K. et al. Clinical characteristics, outcomes, and risk factors for mortality in patients with cancer and COVID-19 in Hubei, China: a multicentre, retrospective, cohort study. Lancet Oncol. https://doi.org/10.1016/S1470-2045(20)30310-7 (2020).",
"year": "2020"
},
{
"DOI": "10.3389/fimmu.2018.01873",
"author": "EB Thangam",
"doi-asserted-by": "publisher",
"first-page": "1873",
"journal-title": "Front. Immunol.",
"key": "84782_CR32",
"unstructured": "Thangam, E. B. et al. The role of histamine and histamine receptors in mast cell-mediated allergy and inflammation: the hunt for new therapeutic targets. Front. Immunol. 9, 1873. https://doi.org/10.3389/fimmu.2018.01873 (2018).",
"volume": "9",
"year": "2018"
},
{
"DOI": "10.3390/ijms20174241",
"author": "JS Marshall",
"doi-asserted-by": "publisher",
"journal-title": "Int. J. Mol. Sci.",
"key": "84782_CR33",
"unstructured": "Marshall, J. S., Portales-Cervantes, L. & Leong, E. Mast cell responses to viruses and pathogen products. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20174241 (2019).",
"year": "2019"
},
{
"DOI": "10.3389/fimmu.2017.00310",
"author": "B Zarnegar",
"doi-asserted-by": "publisher",
"first-page": "310",
"journal-title": "Front. Immunol.",
"key": "84782_CR34",
"unstructured": "Zarnegar, B. et al. Influenza infection in mice induces accumulation of lung mast cells through the recruitment and maturation of mast cell progenitors. Front. Immunol. 8, 310. https://doi.org/10.3389/fimmu.2017.00310 (2017).",
"volume": "8",
"year": "2017"
},
{
"DOI": "10.1128/JVI.06053-11",
"author": "Y Hu",
"doi-asserted-by": "publisher",
"first-page": "3347",
"journal-title": "J. Virol.",
"key": "84782_CR35",
"unstructured": "Hu, Y. et al. Mast cell-induced lung injury in mice infected with H5N1 influenza virus. J. Virol. 86, 3347–3356. https://doi.org/10.1128/JVI.06053-11 (2012).",
"volume": "86",
"year": "2012"
},
{
"DOI": "10.1016/0016-5085(89)90503-9",
"author": "W Kirch",
"doi-asserted-by": "publisher",
"first-page": "1388",
"journal-title": "Gastroenterology",
"key": "84782_CR36",
"unstructured": "Kirch, W., Halabi, A., Linde, M., Santos, S. R. & Ohnhaus, E. E. Negative effects of famotidine on cardiac performance assessed by noninvasive hemodynamic measurements. Gastroenterology 96, 1388–1392. https://doi.org/10.1016/0016-5085(89)90503-9 (1989).",
"volume": "96",
"year": "1989"
},
{
"DOI": "10.1038/sj.eye.6701839",
"author": "YC Lee",
"doi-asserted-by": "publisher",
"first-page": "260",
"journal-title": "Eye (Lond)",
"key": "84782_CR37",
"unstructured": "Lee, Y. C. & Wang, C. C. Famotidine-induced retinopathy. Eye (Lond) 20, 260–263. https://doi.org/10.1038/sj.eye.6701839 (2006).",
"volume": "20",
"year": "2006"
},
{
"author": "SM Kallal",
"first-page": "446",
"journal-title": "West. J. Med.",
"key": "84782_CR38",
"unstructured": "Kallal, S. M. & Lee, M. Thrombotic thrombo-cytopenic purpura associated with histamine H2-receptor antagonist therapy. West. J. Med. 164, 446–448 (1996).",
"volume": "164",
"year": "1996"
},
{
"DOI": "10.7326/0003-4819-114-12-1027",
"author": "TG Cantu",
"doi-asserted-by": "publisher",
"first-page": "1027",
"journal-title": "Ann. Intern. Med.",
"key": "84782_CR39",
"unstructured": "Cantu, T. G. & Korek, J. S. Central nervous system reactions to histamine-2 receptor blockers. Ann. Intern. Med. 114, 1027–1034. https://doi.org/10.7326/0003-4819-114-12-1027 (1991).",
"volume": "114",
"year": "1991"
},
{
"DOI": "10.1002/pro.3703",
"author": "C Sun",
"doi-asserted-by": "publisher",
"first-page": "19",
"journal-title": "Protein Sci.",
"key": "84782_CR40",
"unstructured": "Sun, C., Li, Y., Yates, E. A. & Fernig, D. G. SimpleDSFviewer: a tool to analyze and view differential scanning fluorimetry data for characterizing protein thermal stability and interactions. Protein Sci. 29, 19–27. https://doi.org/10.1002/pro.3703 (2020).",
"volume": "29",
"year": "2020"
},
{
"DOI": "10.1099/jgv.0.001453",
"author": "NS Ogando",
"doi-asserted-by": "publisher",
"journal-title": "J. Gen. Virol.",
"key": "84782_CR41",
"unstructured": "Ogando, N. S. et al. SARS-coronavirus-2 replication in Vero E6 cells: replication kinetics, rapid adaptation and cytopathology. J. Gen. Virol. https://doi.org/10.1099/jgv.0.001453 (2020).",
"year": "2020"
},
{
"DOI": "10.2807/1560-7917",
"author": "VM Corman",
"doi-asserted-by": "publisher",
"journal-title": "Euro Surveill",
"key": "84782_CR42",
"unstructured": "Corman, V. M. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill https://doi.org/10.2807/1560-7917 (2020).",
"year": "2020"
}
],
"reference-count": 42,
"references-count": 42,
"relation": {},
"score": 1,
"short-container-title": [
"Sci Rep"
],
"short-title": [],
"source": "Crossref",
"subject": [
"Multidisciplinary"
],
"subtitle": [],
"title": [
"The in-vitro effect of famotidine on SARS-CoV-2 proteases and virus replication"
],
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.1007/springer_crossmark_policy",
"volume": "11"
}
