Onvansertib and vilazodone inhibit SARS-CoV-2 replication via suppression of METTL3 RNA-m6A enzymatic activity
et al., Antiviral Research, doi:10.1016/j.antiviral.2026.106376, Feb 2026
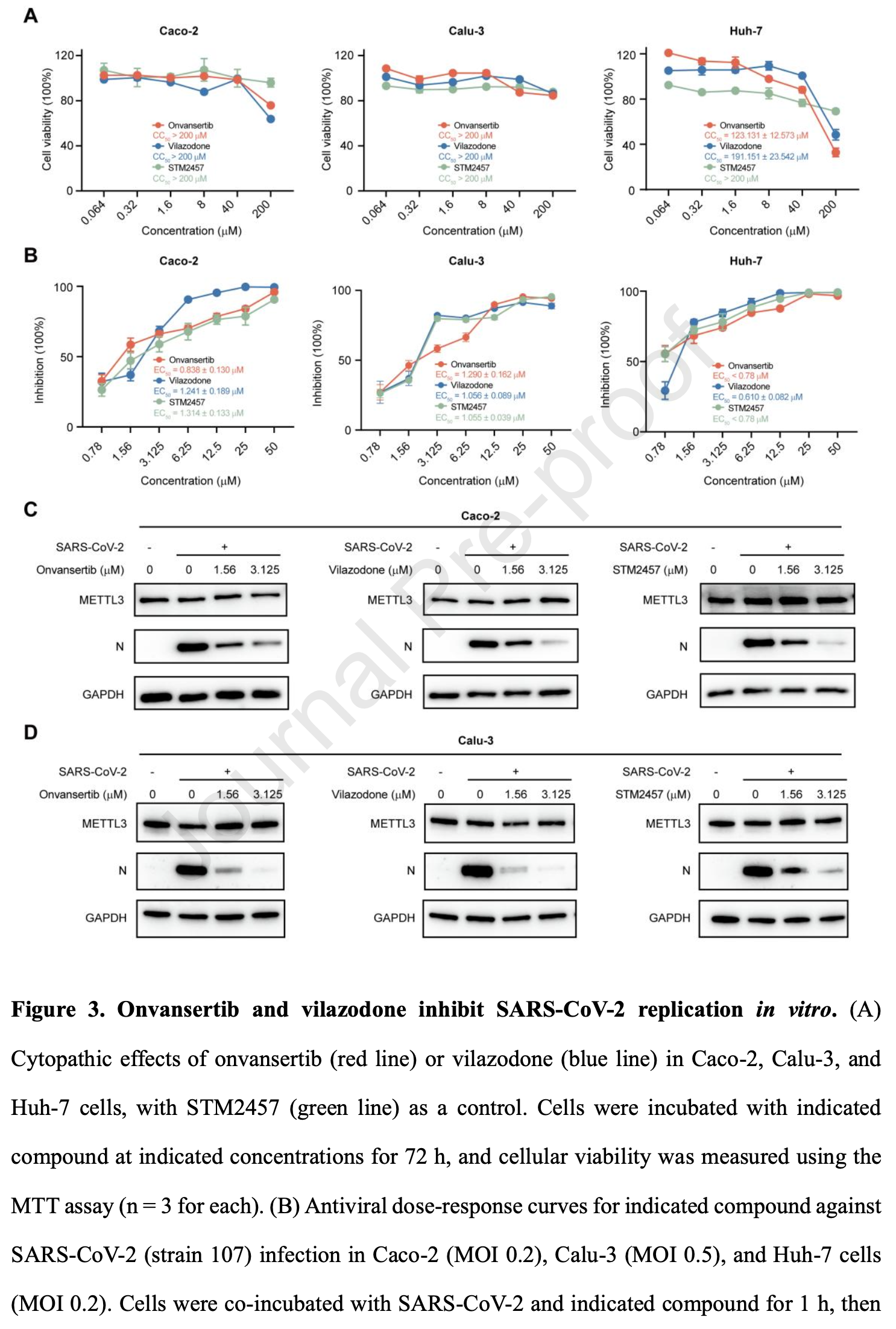
In vitro study showing that onvansertib and vilazodone inhibit SARS-CoV-2 replication by targeting the host RNA methyltransferase METTL3.
Zhang et al., 19 Feb 2026, USA, peer-reviewed, 9 authors.
Contact: zhengyt@mail.kiz.ac.cn, yangwenlan@imu.edu.cn.
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
Onvansertib and vilazodone inhibit SARS-CoV-2 replication via suppression of METTL3 RNA-m6A enzymatic activity
Antiviral Research, doi:10.1016/j.antiviral.2026.106376
The continuous emergence of new SARS-CoV-2 variants limits the effectiveness of current vaccines and monoclonal antibodies, highlighting the urgent need for complementary antiviral strategies. Targeting host proteins that are exploited by the virus represents a promising approach to achieve broad-spectrum inhibition. Given the recently established proviral role of METTL3-dependent RNA m 6 A modification in SARS-CoV-2 replication, we conducted a structure-based virtual screening to repurpose compounds with acceptable safety profiles.
Subsequent biochemical validation identified onvansertib and vilazodone as direct METTL3 binders and potent inhibitors of its methyltransferase activity. Notably, both compounds effectively suppressed the replication of ancestral SARS-CoV-2 and Omicron BA.2 variant in vitro. Mechanistically, each compound potentially enhanced the innate antiviral response of the host and repressed the expression of viral entry-associated host factors during infection; this observation is consistent with the established proviral role of METTL3. In summary, these findings not only substantiate METTL3 as a viable antiviral target but also lay the foundation for further research on the therapeutic potential of onvansertib and vilazodone against COVID-19.
Onvansertib and vilazodone could enhance host innate immune response activation and suppress proviral host factor expression during SARS-CoV-2 infection Given that both onvansertib and vilazodone inhibit SARS-CoV-2 replication, we next assessed the impact of each compound on host cells using RNA-seq. In infected Caco-2 cells treated with either compound, we identified thousands of differentially expressed genes (DEGs) compared to DMSO controls (Supplementary Fig. 6B ). Onvansertib treatment resulted in more DEGs (3, 683 up; 4, 562 down) than vilazodone (1,361 up; 1,683 down). For comparison, reanalysis of a public RNA-seq data (GSE167075) revealed METTL3 depletion altered 3,845 genes during SARS-CoV-2 infection. Despite the difference in numbers, all three conditions exhibited a consistent bias towards decreasing gene expression (Supplementary Fig. 6B ), suggesting that the down-regulated genes may be crucial for viral infection. To pinpoint the most critical shared responses, we performed an overlay analysis. It identified 155 genes that were consistently up-regulated across all three conditions (Supplementary Fig. 6C ). Gene Ontology (GO) analysis indicated their involvement in diverse cellular processes, including cell division, immune response and apoptotic process (Supplementary Fig. 6D ). Additionally, Gene Set Enrichment Analysis (GSEA) revealed a significant up-regulation of the RIG-I-like receptor signaling pathway (Supplementary Fig. 6G ), confirming..
References
Ahn, Barzi, Ridinger, Samuelsz, Subramanian et al., Onvansertib in combination with FOLFIRI and Bevacizumab in second-line treatment of KRAS-mutant metastatic colorectal cancer: a phase 1b clinical study, Clin Cancer Res
Aly, Scott, Calderon, Haghighi, N 6 -adenosine methylation of SARS-CoV-2 5'-UTR regulates translation, bioRxiv, doi:10.1101/2022.10.17.512569
Arantes, Gomes, Ito, Sarafim, Graf et al., Spatiotemporal dynamics and epidemiological impact of SARS-CoV-2 XBB lineage dissemination in Brazil in 2023, Microbiol Spectr
Banks, Beard, Cao, Cho, Damm et al., Integrated modeling program, applied chemical theory (IMPACT), J Comput Chem
Batista-Roche, Gomez-Gil, Lund, Berlanga-Robles, Garcia-Gasca, Global m 6 A RNA methylation in SARS-CoV-2 positive nasopharyngeal samples in a Mexican population: a first approximation study, Epigenomes
Bedi, Huang, Eberle, Wiedmer, Sledz et al., Small-molecule inhibitors of METTL3, the major human epitranscriptomic writer, ChemMedChem
Beria, Bossi, Brasca, Caruso, Ceccarelli et al., NMS-P937, a 4,5-dihydro-1H-pyrazolo[4,3-h]quinazoline derivative as potent and selective Polo-like kinase 1 inhibitor, Bioorg Med Chem Lett
Beumer, Geurts, Lamers, Puschhof, Zhang et al., A CRISPR/Cas9 genetically engineered organoid biobank reveals essential host factors for coronaviruses, Nat Commun
Bhattacharya, Chatterjee, Lee, Dhama, Chakraborty, Antibody evasion associated with the RBD significant mutations in several emerging SARS-CoV-2 variants and its subvariants, Drug Resist Updat
Bolger, Lohse, Usadel, Trimmomatic: a flexible trimmer for Illumina sequence data, Bioinformatics
Bowers, Chow, Xu, Dror, Eastwood et al., Scalable algorithms for molecular dynamics simulations on commodity clusters, doi:10.1145/1188455.1188544
Burgess, Depledge, Thompson, Srinivas, Grande et al., Targeting the m 6 A RNA modification pathway blocks SARS-CoV-2 and HCoV-OC43 replication, Genes Dev
Campos, Maricato, Braconi, Antoneli, Janini et al., Direct RNA sequencing reveals SARS-CoV-2 m 6 A sites and possible differential DRACH motif methylation among variants, Viruses
Chen, Foloppe, Drug-like bioactive structures and conformational coverage with the LigPrep/ConfGen suite: comparison to programs MOE and catalyst, J Chem Inf Model
Croucher, Ridinger, Becker, Lin, Silberman et al., Spliceosome mutations are associated with clinical response in a phase 1b/2 study of the PLK1 inhibitor onvansertib in combination with decitabine in relapsed or refractory acute myeloid leukemia, Ann Hematol
Dorgham, Elliott, Holley, Mansfield, m 6 A regulates breast cancer proliferation and migration through stage-dependent changes in epithelial to mesenchymal transition gene expression, Front Oncol
Finkel, Mizrahi, Nachshon, Weingarten-Gabbay, Morgenstern et al., The coding capacity of SARS-CoV-2, Nature
Friesner, Banks, Murphy, Halgren, Klicic et al., Glide: a new approach for rapid, accurate docking and scoring. 1. method and assessment of docking accuracy, J Med Chem
Frostell, Vinterback, Sjobom, Protein-ligand interactions using SPR systems, Methods Mol Biol
Hoffmann, Kleine-Weber, Pohlmann, A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells, Mol Cell
Hoffmann, Pohlmann, Novel SARS-CoV-2 receptors: ASGR1 and KREMEN1, Cell Res
J O U R N A L P R E -P R O O F Daniloski, Jordan, Wessels, Hoagland, Kasela et al., Identification of required host factors for SARS-CoV-2 infection in human cells, Cell
J O U R N A L P R E -P R O O F Sendinc, Shi, RNA m 6 A methylation across the transcriptome, Mol Cell
Jafari, Almqvist, Axelsson, Ignatushchenko, Lundback et al., The cellular thermal shift assay for evaluating drug target interactions in cells, Nat Protoc
Jiang, Chen, Xiong, Xu, Gong et al., Knockdown of m 6 A methyltransferase METTL3 in gastric cancer cells results in suppression of cell proliferation, Oncol Lett
Johnsson, Lofas, Lindquist, Immobilization of proteins to a carboxymethyldextranmodified gold surface for biospecific interaction analysis in surface plasmon resonance sensors, Anal Biochem
Kim, Paggi, Park, Bennett, Salzberg, Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype, Nat Biotechnol
Li, Hui, Bray, Gonzalez, Zeller et al., METTL3 regulates viral m 6 A RNA modification and host cell innate immune responses during SARS-CoV-2 infection, Cell Rep
Liu, Xu, Li, Ye, Zhou et al., The m 6 A methylome of SARS-CoV-2 in host cells, Cell Res
Lu, Wu, Ghoreishi, Chen, Wang et al., OPLS4: improving force field accuracy on challenging regimes of chemical space, J Chem Theory Comput
Lyne, Lamb, Saeh, Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring, J Med Chem
Mu, Zhang, Yang, Zhang, Gao et al., METTL3-mediated mRNA N 6 -methyladenosine is required for oocyte and follicle development in mice, Cell Death Dis
Nie, Li, Wu, Zhao, Hao et al., Quantification of SARS-CoV-2 neutralizing antibody by a pseudotyped virus-based assay, Nat Protoc
Ouyang, Hong, Fu, Li, Zeng et al., METTL3 depletion contributes to tumour progression and drug resistance via N 6 methyladenosine-dependent mechanism in HR+HER2-breast cancer, Breast Cancer Res
Pang, Lu, Zhao, Shen, Fan et al., A variant-proof SARS-CoV-2 vaccine targeting HR1 domain in S2 subunit of spike protein, Cell Res
Planas, Staropoli, Michel, Lemoine, Donati et al., Distinct evolution of SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineages combining increased fitness and antibody evasion, Nat Commun
Planas, Veyer, Baidaliuk, Staropoli, Guivel-Benhassine et al., Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization, Nature
Putri, Anders, Pyl, Pimanda, Zanini, Analysing high-throughput sequencing data in Python with HTSeq 2.0, Bioinformatics
Qiao, Li, Zeng, Liu, Luo et al., SARS-CoV-2 M pro inhibitors with antiviral activity in a transgenic mouse model, Science
Ramirez, Caballero, Is it reliable to take the molecular docking top scoring position as the best solution without considering available structural data?, Molecules
Rossler, Netzl, Knabl, Bante, Wilks et al., Characterizing SARS-CoV-2 neutralization profiles after bivalent boosting using antigenic cartography, Nat Commun
Sastry, Adzhigirey, Day, Annabhimoju, Sherman, Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments, J Comput Aided Mol Des
Sherman, Hao, Qiu, Jiao, Baseler et al., DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update), Nucleic Acids Res
Su, Chhabra, Singh, Ndiaye, Ahmad, PLK1 inhibition-based combination therapies for cancer management, Transl Oncol
Tamura, Ito, Uriu, Zahradnik, Kida et al., Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants, Nat Commun
V'kovski, Kratzel, Steiner, Stalder, Thiel, Coronavirus biology and replication: implications for SARS-CoV-2, Nat Rev Microbiol
Vaid, Mendez, Thombare, Burgos-Panadero, Robinot et al., Global loss of cellular m 6 A RNA methylation following infection with different SARS-CoV-2 variants, Genome Res
Wang, Doxtader, Nam, Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases, Mol Cell
Wang, Feng, Wang, Wang, Zhang, DEGseq: an R package for identifying differentially expressed genes from RNA-seq data, Bioinformatics
Wang, Wang, Wu, Li, Bray et al., PCIF1-mediated deposition of 5'-cap N 6 ,2'-O-dimethyladenosine in ACE2 and TMPRSS2 mRNA regulates susceptibility to SARS-CoV-2 infection, Proc Natl Acad Sci U S A
Wang, Zhou, N 6 -methyladenosine and its implications in viruses, Genomics Proteomics Bioinformatics
Wei, Patil, Collings, Alfajaro, Liang et al., Pharmacological disruption of mSWI/SNF complex activity restricts SARS-CoV-2 infection, Nat Genet
Wishart, Feunang, Guo, Lo, Marcu et al., DrugBank 5.0: a major update to the DrugBank database for 2018, Nucleic Acids Res
Xuan, Chen, Chen, Pang, Huang et al., RMBase v3.0: decode the landscape, mechanisms and functions of RNA modifications, Nucleic Acids Res
Yankova, Blackaby, Albertella, Rak, De Braekeleer et al., Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia, Nature
Yu, Jacobson, Friesner, What role do surfaces play in GB models? A newgeneration of surface-generalized born model based on a novel gaussian surface for biomolecules, J Comput Chem
Zeidan, Ridinger, Lin, Becker, Schiller et al., A phase Ib study of onvansertib, a novel oral PLK1 inhibitor, in combination therapy for patients with relapsed or refractory acute myeloid leukemia, Clin Cancer Res
Zhang, Hao, Ma, Zhang, Hu et al., Methyltransferase-like 3 modulates severe acute respiratory syndrome coronavirus-2 RNA N 6methyladenosine modification and replication, mBio
Zhang, Peng, Wang, N 6 -methyladenosine modification-a key player in viral infection, Cell Mol Biol Lett
Zheng, Xue, Yang, Zhang, Chen et al., Revealing vilazodone's binding mechanism underlying its partial agonism to the 5-HT(1A) receptor in the treatment of major depressive disorder, Phys Chem Chem Phys
DOI record:
{
"DOI": "10.1016/j.antiviral.2026.106376",
"ISSN": [
"0166-3542"
],
"URL": "http://dx.doi.org/10.1016/j.antiviral.2026.106376",
"alternative-id": [
"S0166354226000355"
],
"article-number": "106376",
"assertion": [
{
"label": "This article is maintained by",
"name": "publisher",
"value": "Elsevier"
},
{
"label": "Article Title",
"name": "articletitle",
"value": "Onvansertib and vilazodone inhibit SARS-CoV-2 replication via suppression of METTL3 RNA-m6A enzymatic activity"
},
{
"label": "Journal Title",
"name": "journaltitle",
"value": "Antiviral Research"
},
{
"label": "CrossRef DOI link to publisher maintained version",
"name": "articlelink",
"value": "https://doi.org/10.1016/j.antiviral.2026.106376"
},
{
"label": "Content Type",
"name": "content_type",
"value": "article"
},
{
"label": "Copyright",
"name": "copyright",
"value": "© 2026 The Authors. Published by Elsevier B.V."
}
],
"author": [
{
"ORCID": "https://orcid.org/0000-0001-9650-6089",
"affiliation": [],
"authenticated-orcid": false,
"family": "Zhang",
"given": "Ting",
"sequence": "first"
},
{
"ORCID": "https://orcid.org/0000-0002-6716-2456",
"affiliation": [],
"authenticated-orcid": false,
"family": "Wang",
"given": "Zi-Ling",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Li",
"given": "Xi-Ya",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Luo",
"given": "Rong-Hua",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0009-0003-6156-9591",
"affiliation": [],
"authenticated-orcid": false,
"family": "Zhang",
"given": "Lu-Xi",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0009-0004-3758-9207",
"affiliation": [],
"authenticated-orcid": false,
"family": "Zheng",
"given": "Hong-Yi",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Xiong",
"given": "Si-Dong",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Yang",
"given": "Wen-Lan",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0001-5469-0324",
"affiliation": [],
"authenticated-orcid": false,
"family": "Zheng",
"given": "Yong-Tang",
"sequence": "additional"
}
],
"container-title": "Antiviral Research",
"container-title-short": "Antiviral Research",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"elsevier.com",
"sciencedirect.com"
]
},
"created": {
"date-parts": [
[
2026,
2,
19
]
],
"date-time": "2026-02-19T08:02:54Z",
"timestamp": 1771488174000
},
"deposited": {
"date-parts": [
[
2026,
2,
21
]
],
"date-time": "2026-02-21T07:16:13Z",
"timestamp": 1771658173000
},
"funder": [
{
"DOI": "10.13039/501100001809",
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100001809",
"id-type": "DOI"
}
],
"name": "National Natural Science Foundation of China"
}
],
"indexed": {
"date-parts": [
[
2026,
2,
21
]
],
"date-time": "2026-02-21T08:05:55Z",
"timestamp": 1771661155817,
"version": "3.50.1"
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2026,
2
]
]
},
"language": "en",
"license": [
{
"URL": "https://www.elsevier.com/tdm/userlicense/1.0/",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
2,
1
]
],
"date-time": "2026-02-01T00:00:00Z",
"timestamp": 1769904000000
}
},
{
"URL": "https://www.elsevier.com/legal/tdmrep-license",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
2,
1
]
],
"date-time": "2026-02-01T00:00:00Z",
"timestamp": 1769904000000
}
},
{
"URL": "http://creativecommons.org/licenses/by-nc-nd/4.0/",
"content-version": "vor",
"delay-in-days": 20,
"start": {
"date-parts": [
[
2026,
2,
21
]
],
"date-time": "2026-02-21T00:00:00Z",
"timestamp": 1771632000000
}
}
],
"link": [
{
"URL": "https://api.elsevier.com/content/article/PII:S0166354226000355?httpAccept=text/xml",
"content-type": "text/xml",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://api.elsevier.com/content/article/PII:S0166354226000355?httpAccept=text/plain",
"content-type": "text/plain",
"content-version": "vor",
"intended-application": "text-mining"
}
],
"member": "78",
"original-title": [],
"page": "106376",
"prefix": "10.1016",
"published": {
"date-parts": [
[
2026,
2
]
]
},
"published-print": {
"date-parts": [
[
2026,
2
]
]
},
"publisher": "Elsevier BV",
"reference": [
{
"DOI": "10.1158/1078-0432.CCR-23-3053",
"article-title": "Onvansertib in combination with FOLFIRI and Bevacizumab in second-line treatment of KRAS-mutant metastatic colorectal cancer: a phase 1b clinical study",
"author": "Ahn",
"doi-asserted-by": "crossref",
"first-page": "2039",
"journal-title": "Clin Cancer Res.",
"key": "10.1016/j.antiviral.2026.106376_bib1",
"volume": "30",
"year": "2024"
},
{
"article-title": "N6-adenosine methylation of SARS-CoV-2 5’-UTR regulates translation",
"author": "Aly",
"journal-title": "bioRxiv",
"key": "10.1016/j.antiviral.2026.106376_bib2",
"year": "2022"
},
{
"DOI": "10.1128/spectrum.03831-23",
"article-title": "Spatiotemporal dynamics and epidemiological impact of SARS-CoV-2 XBB lineage dissemination in Brazil in 2023",
"author": "Arantes",
"doi-asserted-by": "crossref",
"journal-title": "Microbiol Spectr",
"key": "10.1016/j.antiviral.2026.106376_bib3",
"volume": "12",
"year": "2024"
},
{
"DOI": "10.1038/s41564-021-00958-0",
"article-title": "Cellular host factors for SARS-CoV-2 infection",
"author": "Baggen",
"doi-asserted-by": "crossref",
"first-page": "1219",
"journal-title": "Nat Microbiol",
"key": "10.1016/j.antiviral.2026.106376_bib4",
"volume": "6",
"year": "2021"
},
{
"DOI": "10.1002/jcc.20292",
"article-title": "Integrated modeling program, applied chemical theory (IMPACT)",
"author": "Banks",
"doi-asserted-by": "crossref",
"first-page": "1752",
"journal-title": "J Comput Chem",
"key": "10.1016/j.antiviral.2026.106376_bib5",
"volume": "26",
"year": "2005"
},
{
"DOI": "10.3390/epigenomes6030016",
"article-title": "Global m6A RNA methylation in SARS-CoV-2 positive nasopharyngeal samples in a Mexican population: a first approximation study",
"author": "Batista-Roche",
"doi-asserted-by": "crossref",
"first-page": "16",
"journal-title": "Epigenomes",
"key": "10.1016/j.antiviral.2026.106376_bib6",
"volume": "6",
"year": "2022"
},
{
"DOI": "10.1002/cmdc.202000011",
"article-title": "Small-molecule inhibitors of METTL3, the major human epitranscriptomic writer",
"author": "Bedi",
"doi-asserted-by": "crossref",
"first-page": "744",
"journal-title": "ChemMedChem",
"key": "10.1016/j.antiviral.2026.106376_bib7",
"volume": "15",
"year": "2020"
},
{
"DOI": "10.1016/j.bmcl.2011.03.054",
"article-title": "NMS-P937, a 4,5-dihydro-1H-pyrazolo[4,3-h]quinazoline derivative as potent and selective Polo-like kinase 1 inhibitor",
"author": "Beria",
"doi-asserted-by": "crossref",
"first-page": "2969",
"journal-title": "Bioorg Med Chem Lett",
"key": "10.1016/j.antiviral.2026.106376_bib8",
"volume": "21",
"year": "2011"
},
{
"DOI": "10.1038/s41467-021-25729-7",
"article-title": "A CRISPR/Cas9 genetically engineered organoid biobank reveals essential host factors for coronaviruses",
"author": "Beumer",
"doi-asserted-by": "crossref",
"first-page": "5498",
"journal-title": "Nat Commun",
"key": "10.1016/j.antiviral.2026.106376_bib9",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1016/j.drup.2023.101008",
"article-title": "Antibody evasion associated with the RBD significant mutations in several emerging SARS-CoV-2 variants and its subvariants",
"author": "Bhattacharya",
"doi-asserted-by": "crossref",
"first-page": "101008",
"journal-title": "Drug Resist Updat",
"key": "10.1016/j.antiviral.2026.106376_bib10",
"volume": "71",
"year": "2023"
},
{
"DOI": "10.1145/1188455.1188544",
"doi-asserted-by": "crossref",
"key": "10.1016/j.antiviral.2026.106376_bib11",
"unstructured": "Bowers, K.J., Chow, D.E., Xu, H., Dror, R.O., Eastwood, M.P., Gregersen, B.A., Klepeis, J.L., Kolossvary, I., Moraes, M.A., Sacerdoti, F.D., Salmon J.K., Shan Y.B., Shaw D.E., 2006. Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proceedings of the ACM/IEEE Conference on Supercomputing (SC06). https://dl.acm.org/doi/pdf/10.1145/1188455.1188544"
},
{
"DOI": "10.1093/bioinformatics/btu170",
"article-title": "Trimmomatic: a flexible trimmer for Illumina sequence data",
"author": "Bolger",
"doi-asserted-by": "crossref",
"first-page": "2114",
"journal-title": "Bioinformatics",
"key": "10.1016/j.antiviral.2026.106376_bib12",
"volume": "30",
"year": "2014"
},
{
"DOI": "10.1101/gad.348320.121",
"article-title": "Targeting the m6A RNA modification pathway blocks SARS-CoV-2 and HCoV-OC43 replication",
"author": "Burgess",
"doi-asserted-by": "crossref",
"first-page": "1005",
"journal-title": "Genes Dev",
"key": "10.1016/j.antiviral.2026.106376_bib13",
"volume": "35",
"year": "2021"
},
{
"DOI": "10.3390/v13112108",
"article-title": "Direct RNA sequencing reveals SARS-CoV-2 m6A sites and possible differential DRACH motif methylation among variants",
"author": "Campos",
"doi-asserted-by": "crossref",
"first-page": "2108",
"journal-title": "Viruses",
"key": "10.1016/j.antiviral.2026.106376_bib14",
"volume": "13",
"year": "2021"
},
{
"DOI": "10.1021/ci100026x",
"article-title": "Drug-like bioactive structures and conformational coverage with the LigPrep/ConfGen suite: comparison to programs MOE and catalyst",
"author": "Chen",
"doi-asserted-by": "crossref",
"first-page": "822",
"journal-title": "J Chem Inf Model",
"key": "10.1016/j.antiviral.2026.106376_bib15",
"volume": "50",
"year": "2010"
},
{
"DOI": "10.1007/s00277-023-05442-9",
"article-title": "Spliceosome mutations are associated with clinical response in a phase 1b/2 study of the PLK1 inhibitor onvansertib in combination with decitabine in relapsed or refractory acute myeloid leukemia",
"author": "Croucher",
"doi-asserted-by": "crossref",
"first-page": "3049",
"journal-title": "Ann Hematol",
"key": "10.1016/j.antiviral.2026.106376_bib16",
"volume": "102",
"year": "2023"
},
{
"DOI": "10.1016/j.cell.2020.10.030",
"article-title": "Identification of required host factors for SARS-CoV-2 infection in human cells",
"author": "Daniloski",
"doi-asserted-by": "crossref",
"first-page": "92",
"journal-title": "Cell",
"key": "10.1016/j.antiviral.2026.106376_bib17",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.3389/fonc.2023.1268977",
"article-title": "m6A regulates breast cancer proliferation and migration through stage-dependent changes in epithelial to mesenchymal transition gene expression",
"author": "Dorgham",
"doi-asserted-by": "crossref",
"first-page": "1268977",
"journal-title": "Front Oncol",
"key": "10.1016/j.antiviral.2026.106376_bib18",
"volume": "13",
"year": "2023"
},
{
"DOI": "10.1038/s41586-020-2739-1",
"article-title": "The coding capacity of SARS-CoV-2",
"author": "Finkel",
"doi-asserted-by": "crossref",
"first-page": "125",
"journal-title": "Nature",
"key": "10.1016/j.antiviral.2026.106376_bib19",
"volume": "589",
"year": "2021"
},
{
"DOI": "10.1021/jm0306430",
"article-title": "Glide: a new approach for rapid, accurate docking and scoring. 1. method and assessment of docking accuracy",
"author": "Friesner",
"doi-asserted-by": "crossref",
"first-page": "1739",
"journal-title": "J Med Chem.",
"key": "10.1016/j.antiviral.2026.106376_bib20",
"volume": "47",
"year": "2004"
},
{
"DOI": "10.1007/978-1-62703-398-5_6",
"article-title": "Protein-ligand interactions using SPR systems",
"author": "Frostell",
"doi-asserted-by": "crossref",
"first-page": "139",
"journal-title": "Methods Mol Biol",
"key": "10.1016/j.antiviral.2026.106376_bib21",
"volume": "1008",
"year": "2013"
},
{
"DOI": "10.1016/j.molcel.2020.04.022",
"article-title": "A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells",
"author": "Hoffmann",
"doi-asserted-by": "crossref",
"first-page": "779",
"journal-title": "Mol Cell",
"key": "10.1016/j.antiviral.2026.106376_bib22",
"volume": "78",
"year": "2020"
},
{
"DOI": "10.1038/s41422-021-00603-9",
"article-title": "SARS-CoV-2 receptors: ASGR1 and KREMEN1",
"author": "Hoffmann",
"doi-asserted-by": "crossref",
"first-page": "1",
"journal-title": "Cell Res",
"key": "10.1016/j.antiviral.2026.106376_bib23",
"volume": "32",
"year": "2022"
},
{
"DOI": "10.1038/nprot.2014.138",
"article-title": "The cellular thermal shift assay for evaluating drug target interactions in cells",
"author": "Jafari",
"doi-asserted-by": "crossref",
"first-page": "2100",
"journal-title": "Nat Protoc",
"key": "10.1016/j.antiviral.2026.106376_bib24",
"volume": "9",
"year": "2014"
},
{
"DOI": "10.3892/ol.2020.11794",
"article-title": "Knockdown of m6A methyltransferase METTL3 in gastric cancer cells results in suppression of cell proliferation",
"author": "Jiang",
"doi-asserted-by": "crossref",
"first-page": "2191",
"journal-title": "Oncol Lett",
"key": "10.1016/j.antiviral.2026.106376_bib25",
"volume": "20",
"year": "2020"
},
{
"DOI": "10.1016/0003-2697(91)90424-R",
"article-title": "Immobilization of proteins to a carboxymethyldextran-modified gold surface for biospecific interaction analysis in surface plasmon resonance sensors",
"author": "Johnsson",
"doi-asserted-by": "crossref",
"first-page": "268",
"journal-title": "Anal Biochem.",
"key": "10.1016/j.antiviral.2026.106376_bib26",
"volume": "198",
"year": "1991"
},
{
"DOI": "10.1038/s41587-019-0201-4",
"article-title": "Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype",
"author": "Kim",
"doi-asserted-by": "crossref",
"first-page": "907",
"journal-title": "Nat Biotechnol",
"key": "10.1016/j.antiviral.2026.106376_bib27",
"volume": "37",
"year": "2019"
},
{
"DOI": "10.1016/j.celrep.2021.109091",
"article-title": "METTL3 regulates viral m6A RNA modification and host cell innate immune responses during SARS-CoV-2 infection",
"author": "Li",
"doi-asserted-by": "crossref",
"first-page": "109091",
"journal-title": "Cell Rep",
"key": "10.1016/j.antiviral.2026.106376_bib28",
"volume": "35",
"year": "2021"
},
{
"DOI": "10.1038/s41422-020-00465-7",
"article-title": "The m6A methylome of SARS-CoV-2 in host cells",
"author": "Liu",
"doi-asserted-by": "crossref",
"first-page": "404",
"journal-title": "Cell Res",
"key": "10.1016/j.antiviral.2026.106376_bib29",
"volume": "31",
"year": "2021"
},
{
"DOI": "10.1021/acs.jctc.1c00302",
"article-title": "OPLS4: improving force field accuracy on challenging regimes of chemical space",
"author": "Lu",
"doi-asserted-by": "crossref",
"first-page": "4291",
"journal-title": "J Chem Theory Comput.",
"key": "10.1016/j.antiviral.2026.106376_bib30",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.1021/jm060522a",
"article-title": "Accurate prediction of the relative potencies of members of a series of kinase inhibitors using molecular docking and MM-GBSA scoring",
"author": "Lyne",
"doi-asserted-by": "crossref",
"first-page": "4805",
"journal-title": "J Med Chem.",
"key": "10.1016/j.antiviral.2026.106376_bib31",
"volume": "49",
"year": "2006"
},
{
"DOI": "10.1038/s41419-021-04272-9",
"article-title": "METTL3-mediated mRNA N6-methyladenosine is required for oocyte and follicle development in mice",
"author": "Mu",
"doi-asserted-by": "crossref",
"first-page": "989",
"journal-title": "Cell Death Dis",
"key": "10.1016/j.antiviral.2026.106376_bib32",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1038/s41596-020-0394-5",
"article-title": "Quantification of SARS-CoV-2 neutralizing antibody by a pseudotyped virus-based assay",
"author": "Nie",
"doi-asserted-by": "crossref",
"first-page": "3699",
"journal-title": "Nat Protoc",
"key": "10.1016/j.antiviral.2026.106376_bib33",
"volume": "15",
"year": "2020"
},
{
"DOI": "10.1186/s13058-022-01598-w",
"article-title": "METTL3 depletion contributes to tumour progression and drug resistance via N6 methyladenosine-dependent mechanism in HR+HER2-breast cancer",
"author": "Ouyang",
"doi-asserted-by": "crossref",
"first-page": "19",
"journal-title": "Breast Cancer Res",
"key": "10.1016/j.antiviral.2026.106376_bib34",
"volume": "25",
"year": "2023"
},
{
"DOI": "10.1038/s41422-022-00746-3",
"article-title": "A variant-proof SARS-CoV-2 vaccine targeting HR1 domain in S2 subunit of spike protein",
"author": "Pang",
"doi-asserted-by": "crossref",
"first-page": "1068",
"journal-title": "Cell Res",
"key": "10.1016/j.antiviral.2026.106376_bib35",
"volume": "32",
"year": "2022"
},
{
"DOI": "10.1038/s41467-024-46490-7",
"article-title": "Distinct evolution of SARS-CoV-2 Omicron XBB and BA.2.86/JN.1 lineages combining increased fitness and antibody evasion",
"author": "Planas",
"doi-asserted-by": "crossref",
"first-page": "2254",
"journal-title": "Nat Commun",
"key": "10.1016/j.antiviral.2026.106376_bib36",
"volume": "15",
"year": "2024"
},
{
"DOI": "10.1038/s41586-021-03777-9",
"article-title": "Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization",
"author": "Planas",
"doi-asserted-by": "crossref",
"first-page": "276",
"journal-title": "Nature",
"key": "10.1016/j.antiviral.2026.106376_bib37",
"volume": "596",
"year": "2021"
},
{
"DOI": "10.1093/bioinformatics/btac166",
"article-title": "Analysing high-throughput sequencing data in Python with HTSeq 2.0",
"author": "Putri",
"doi-asserted-by": "crossref",
"first-page": "2943",
"journal-title": "Bioinformatics",
"key": "10.1016/j.antiviral.2026.106376_bib38",
"volume": "38",
"year": "2022"
},
{
"DOI": "10.1126/science.abf1611",
"article-title": "SARS-CoV-2 Mpro inhibitors with antiviral activity in a transgenic mouse model",
"author": "Qiao",
"doi-asserted-by": "crossref",
"first-page": "1374",
"journal-title": "Science",
"key": "10.1016/j.antiviral.2026.106376_bib39",
"volume": "371",
"year": "2021"
},
{
"DOI": "10.3390/molecules23051038",
"article-title": "Is it reliable to take the molecular docking top scoring position as the best solution without considering available structural data?",
"author": "Ramirez",
"doi-asserted-by": "crossref",
"first-page": "1038",
"journal-title": "Molecules",
"key": "10.1016/j.antiviral.2026.106376_bib40",
"volume": "23",
"year": "2018"
},
{
"DOI": "10.1038/s41467-023-41049-4",
"article-title": "Characterizing SARS-CoV-2 neutralization profiles after bivalent boosting using antigenic cartography",
"author": "Rossler",
"doi-asserted-by": "crossref",
"first-page": "5224",
"journal-title": "Nat Commun",
"key": "10.1016/j.antiviral.2026.106376_bib41",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.1007/s10822-013-9644-8",
"article-title": "Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments",
"author": "Sastry",
"doi-asserted-by": "crossref",
"first-page": "221",
"journal-title": "J Comput Aided Mol Des",
"key": "10.1016/j.antiviral.2026.106376_bib42",
"volume": "27",
"year": "2013"
},
{
"DOI": "10.1016/j.molcel.2023.01.006",
"article-title": "RNA m6A methylation across the transcriptome",
"author": "Sendinc",
"doi-asserted-by": "crossref",
"first-page": "428",
"journal-title": "Mol Cell",
"key": "10.1016/j.antiviral.2026.106376_bib43",
"volume": "83",
"year": "2023"
},
{
"DOI": "10.1093/nar/gkac194",
"article-title": "DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update)",
"author": "Sherman",
"doi-asserted-by": "crossref",
"first-page": "216",
"journal-title": "Nucleic Acids Res",
"key": "10.1016/j.antiviral.2026.106376_bib44",
"volume": "50",
"year": "2022"
},
{
"DOI": "10.1016/j.tranon.2021.101332",
"article-title": "PLK1 inhibition-based combination therapies for cancer management",
"author": "Su",
"doi-asserted-by": "crossref",
"journal-title": "Transl Oncol",
"key": "10.1016/j.antiviral.2026.106376_bib45",
"volume": "16",
"year": "2022"
},
{
"DOI": "10.1038/s41467-023-38435-3",
"article-title": "Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants",
"author": "Tamura",
"doi-asserted-by": "crossref",
"first-page": "2800",
"journal-title": "Nat Commun",
"key": "10.1016/j.antiviral.2026.106376_bib46",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.1038/s41579-020-00468-6",
"article-title": "Coronavirus biology and replication: implications for SARS-CoV-2",
"author": "V’Kovski",
"doi-asserted-by": "crossref",
"first-page": "155",
"journal-title": "Nat Rev Microbiol",
"key": "10.1016/j.antiviral.2026.106376_bib47",
"volume": "19",
"year": "2021"
},
{
"DOI": "10.1101/gr.276407.121",
"article-title": "Global loss of cellular m6A RNA methylation following infection with different SARS-CoV-2 variants",
"author": "Vaid",
"doi-asserted-by": "crossref",
"first-page": "299",
"journal-title": "Genome Res",
"key": "10.1016/j.antiviral.2026.106376_bib48",
"volume": "33",
"year": "2023"
},
{
"DOI": "10.1093/bioinformatics/btp612",
"article-title": "DEGseq: an R package for identifying differentially expressed genes from RNA-seq data",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "136",
"journal-title": "Bioinformatics",
"key": "10.1016/j.antiviral.2026.106376_bib49",
"volume": "26",
"year": "2010"
},
{
"article-title": "PCIF1-mediated deposition of 5’-cap N6,2’-O-dimethyladenosine in ACE2 and TMPRSS2 mRNA regulates susceptibility to SARS-CoV-2 infection",
"author": "Wang",
"journal-title": "Proc Natl Acad Sci U S A.",
"key": "10.1016/j.antiviral.2026.106376_bib50",
"volume": "120",
"year": "2023"
},
{
"DOI": "10.1016/j.molcel.2016.05.041",
"article-title": "Structural basis for cooperative function of Mettl3 and Mettl14 methyltransferases",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "306",
"journal-title": "Mol Cell",
"key": "10.1016/j.antiviral.2026.106376_bib51",
"volume": "63",
"year": "2016"
},
{
"article-title": "N6-methyladenosine and its implications in viruses",
"author": "Wang",
"first-page": "695",
"journal-title": "Genomics Proteomics Bioinformatics",
"key": "10.1016/j.antiviral.2026.106376_bib52",
"volume": "4",
"year": "2022"
},
{
"DOI": "10.1038/s41588-023-01307-z",
"article-title": "Pharmacological disruption of mSWI/SNF complex activity restricts SARS-CoV-2 infection",
"author": "Wei",
"doi-asserted-by": "crossref",
"first-page": "471",
"journal-title": "Nat Genet",
"key": "10.1016/j.antiviral.2026.106376_bib53",
"volume": "55",
"year": "2023"
},
{
"DOI": "10.1093/nar/gkx1037",
"article-title": "DrugBank 5.0: a major update to the DrugBank database for 2018",
"author": "Wishart",
"doi-asserted-by": "crossref",
"first-page": "1074",
"journal-title": "Nucleic Acids Res",
"key": "10.1016/j.antiviral.2026.106376_bib54",
"volume": "46",
"year": "2018"
},
{
"DOI": "10.1093/nar/gkad1070",
"article-title": "RMBase v3.0: decode the landscape, mechanisms and functions of RNA modifications",
"author": "Xuan",
"doi-asserted-by": "crossref",
"first-page": "273",
"journal-title": "Nucleic Acids Res",
"key": "10.1016/j.antiviral.2026.106376_bib55",
"volume": "52",
"year": "2024"
},
{
"DOI": "10.1038/s41586-021-03536-w",
"article-title": "Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia",
"author": "Yankova",
"doi-asserted-by": "crossref",
"first-page": "597",
"journal-title": "Nature",
"key": "10.1016/j.antiviral.2026.106376_bib56",
"volume": "593",
"year": "2021"
},
{
"DOI": "10.1002/jcc.20307",
"article-title": "What role do surfaces play in GB models? A new-generation of surface-generalized born model based on a novel gaussian surface for biomolecules",
"author": "Yu",
"doi-asserted-by": "crossref",
"first-page": "72",
"journal-title": "J Comput Chem",
"key": "10.1016/j.antiviral.2026.106376_bib57",
"volume": "27",
"year": "2006"
},
{
"DOI": "10.1158/1078-0432.CCR-20-2586",
"article-title": "A phase Ib study of onvansertib, a novel oral PLK1 inhibitor, in combination therapy for patients with relapsed or refractory acute myeloid leukemia",
"author": "Zeidan",
"doi-asserted-by": "crossref",
"first-page": "6132",
"journal-title": "Clin Cancer Res.",
"key": "10.1016/j.antiviral.2026.106376_bib58",
"volume": "26",
"year": "2020"
},
{
"DOI": "10.1128/mBio.01067-21",
"article-title": "Methyltransferase-like 3 modulates severe acute respiratory syndrome coronavirus-2 RNA N6-methyladenosine modification and replication",
"author": "Zhang",
"doi-asserted-by": "crossref",
"journal-title": "mBio",
"key": "10.1016/j.antiviral.2026.106376_bib59",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1186/s11658-023-00490-5",
"article-title": "N6-methyladenosine modification-a key player in viral infection",
"author": "Zhang",
"doi-asserted-by": "crossref",
"first-page": "78",
"journal-title": "Cell Mol Biol Lett",
"key": "10.1016/j.antiviral.2026.106376_bib60",
"volume": "28",
"year": "2023"
},
{
"DOI": "10.1039/C7CP05688E",
"article-title": "Revealing vilazodone’s binding mechanism underlying its partial agonism to the 5-HT(1A) receptor in the treatment of major depressive disorder",
"author": "Zheng",
"doi-asserted-by": "crossref",
"first-page": "28885",
"journal-title": "Phys Chem Chem Phys",
"key": "10.1016/j.antiviral.2026.106376_bib61",
"volume": "19",
"year": "2017"
}
],
"reference-count": 61,
"references-count": 61,
"relation": {},
"resource": {
"primary": {
"URL": "https://linkinghub.elsevier.com/retrieve/pii/S0166354226000355"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Onvansertib and vilazodone inhibit SARS-CoV-2 replication via suppression of METTL3 RNA-m6A enzymatic activity",
"type": "journal-article",
"update-policy": "https://doi.org/10.1016/elsevier_cm_policy"
}