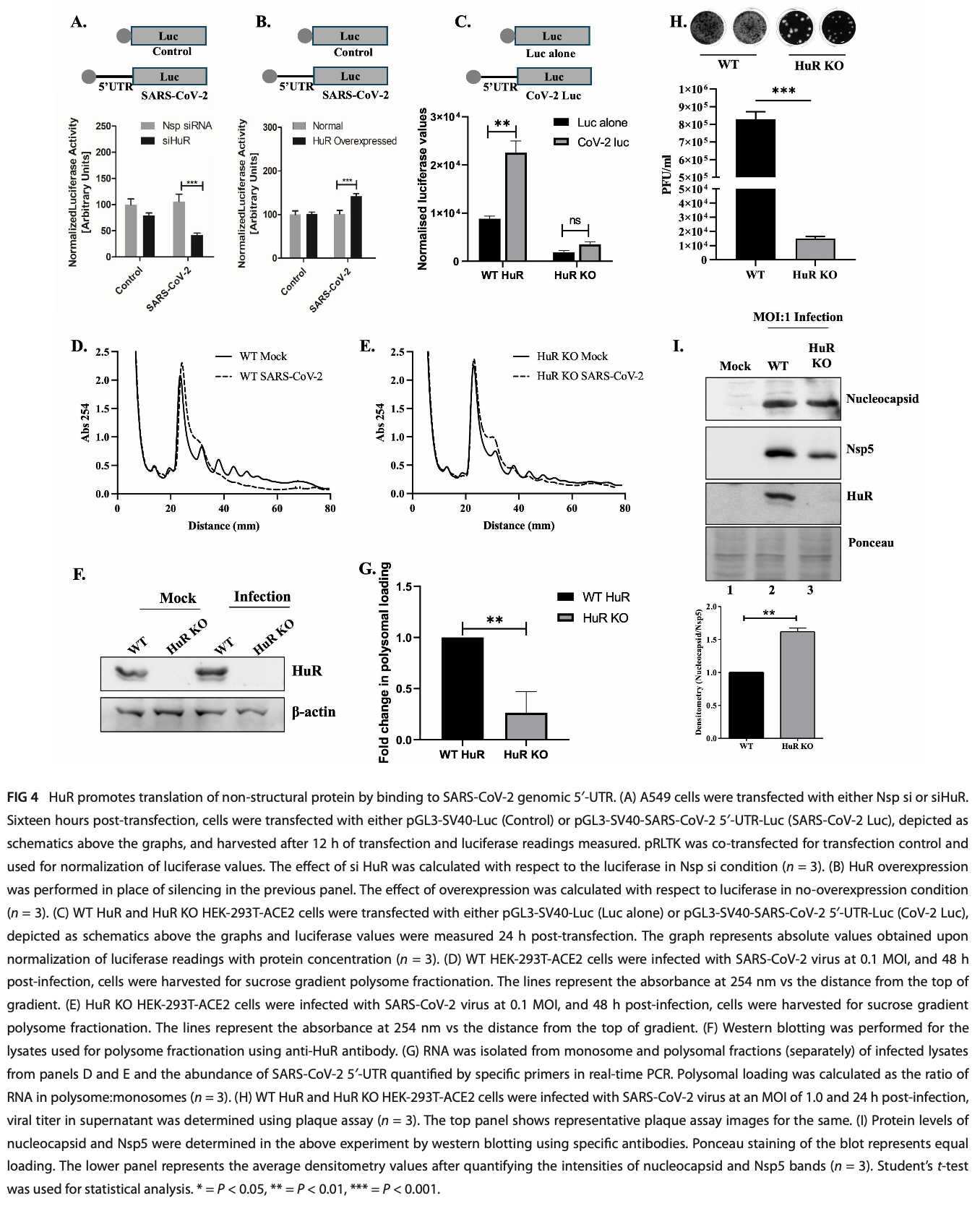
HuR enhances SARS-CoV-2 non-structural protein translation through the genomic 5′-UTR, by promoting polypyrimidine tract-binding protein binding
et al., Journal of Virology, doi:10.1128/jvi.00276-26, Apr 2026
In vitro study showing that human antigen R (HuR), an RNA-binding protein, promotes SARS-CoV-2 replication by enhancing non-structural protein translation through the viral genomic 5′-UTR in HEK-293T-ACE2 and A549 cells.
Raheja et al., 16 Apr 2026, multiple countries, peer-reviewed, 10 authors.
Contact: sdas@iisc.ac.in.
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
Abstract:
| Virology | Full-Length Text
HuR enhances SARS-CoV-2 non-structural protein translation through the genomic 5 ′ -UTR, by promoting polypyrimidine tract-binding protein binding
Harsha Raheja, 1 Risabh Sahu, 1 Trinath Ghosh, 2 Santu Paul, 1 Priya Rani, 1 Ashish Aneja, 1 Biju George, 1 Oyahida Khatun, 1,3 Shashank Tripathi, 1,3 Saumitra Das 1,2
AUTHOR AFFILIATIONS
See affiliation list on p. 18.
ABSTRACT Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) viral RNA associates with different RNA-binding host proteins at each stage of its life cycle. We found sequence-dependent binding of one such important protein, human antigen R (HuR), to the SARS-CoV-2 5 ′ -UTR and studied its potential role in the viral life cycle. The knockdown and knockout studies revealed the importance of such binding in viral translation. We identified 5 ′ -UTR mutations in SARS-CoV-2 variants of concern that altered the HuR-binding affinity. Interestingly, HuR enhanced non-structural protein translation through the genomic 5 ′ -UTR by promoting polypyrimidine tract-binding protein binding to the 5 ′ -UTR. HuR knockout increased the sensitivity to remdesi vir treatment by decreasing its half-maximal inhibitory concentration. An antisense oligonucleotide (whose binding site overlapped the HuR-binding site) reduced viral RNA levels in wild-type cells but not HuR-knockout cells. Our results indicate that HuR regulates the balance between SARS-CoV-2 structural and non-structural proteins and guides the infection of viral variants, implying that HuR can potentially be explored as an antiviral target.
IMPORTANCE Viruses interact with various host proteins throughout their life cycle. A key protein is HuR, an RNA-binding protein regulating RNA stability and translation. HuR binds to viral RNAs at the 5 ′ -UTR or 3 ′ -UTR, influencing translation and replication. We identified conserved HuR binding sites in the SARS-CoV-2 5 ′ -UTR across beta coronavi ruses. This binding enhances translation initiation from the genomic 5 ′ -UTR, increasing non-structural protein production essential for replication. Additionally, we discovered that another host protein, PTB, is recruited by HuR to the viral 5 ′ -UTR, aiding ribosome loading. This regulation shows that the virus exploits HuR for its benefit. Targeting HuR may help control the SARS-CoV-2 life cycle. HuR knockout increased sensitivity to remdesivir, an antiviral drug. Using an antisense oligonucleotide to block HuR binding effectively reduced viral RNA levels. Our findings highlight HuR's critical role in viral protein production regulation and its potential as a therapeutic target against SARSCoV-2.
KEYWORDS 5 ′ -untranslated region, genomic RNA, human antigen R, severe acute respiratory syndrome coronavirus 2, translation
S evere acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a positive-strand RNA virus with a 30 kilobase genome. The viral life cycle begins with attachment of the virus to the angiotensin-converting enzyme 2 (ACE2) receptor. That interaction is followed by receptor-mediated endocytosis, where the virus particles are internal ized into endosomes, which release viral RNA into the host cell cytoplasm. Viral RNA
Editor Christiane E. Wobus, University of..
DOI record:
{
"DOI": "10.1128/jvi.00276-26",
"ISSN": [
"0022-538X",
"1098-5514"
],
"URL": "http://dx.doi.org/10.1128/jvi.00276-26",
"abstract": "<jats:title>ABSTRACT</jats:title>\n <jats:sec>\n <jats:title/>\n <jats:p>Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) viral RNA associates with different RNA-binding host proteins at each stage of its life cycle. We found sequence-dependent binding of one such important protein, human antigen R (HuR), to the SARS-CoV-2 5′-UTR and studied its potential role in the viral life cycle. The knockdown and knockout studies revealed the importance of such binding in viral translation. We identified 5′-UTR mutations in SARS-CoV-2 variants of concern that altered the HuR-binding affinity. Interestingly, HuR enhanced non-structural protein translation through the genomic 5′-UTR by promoting polypyrimidine tract-binding protein binding to the 5′-UTR. HuR knockout increased the sensitivity to remdesivir treatment by decreasing its half-maximal inhibitory concentration. An antisense oligonucleotide (whose binding site overlapped the HuR-binding site) reduced viral RNA levels in wild-type cells but not HuR-knockout cells. Our results indicate that HuR regulates the balance between SARS-CoV-2 structural and non-structural proteins and guides the infection of viral variants, implying that HuR can potentially be explored as an antiviral target.</jats:p>\n <jats:sec>\n <jats:title>IMPORTANCE</jats:title>\n <jats:p>Viruses interact with various host proteins throughout their life cycle. A key protein is HuR, an RNA-binding protein regulating RNA stability and translation. HuR binds to viral RNAs at the 5′-UTR or 3′-UTR, influencing translation and replication. We identified conserved HuR binding sites in the SARS-CoV-2 5′-UTR across beta coronaviruses. This binding enhances translation initiation from the genomic 5′-UTR, increasing non-structural protein production essential for replication. Additionally, we discovered that another host protein, PTB, is recruited by HuR to the viral 5′-UTR, aiding ribosome loading. This regulation shows that the virus exploits HuR for its benefit. Targeting HuR may help control the SARS-CoV-2 life cycle. HuR knockout increased sensitivity to remdesivir, an antiviral drug. Using an antisense oligonucleotide to block HuR binding effectively reduced viral RNA levels. Our findings highlight HuR’s critical role in viral protein production regulation and its potential as a therapeutic target against SARS-CoV-2.</jats:p>\n </jats:sec>\n </jats:sec>",
"alternative-id": [
"10.1128/jvi.00276-26"
],
"article-number": "e00276-26",
"assertion": [
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Received",
"name": "received",
"order": 0,
"value": "2026-02-16"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "2026-03-05"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Published",
"name": "published",
"order": 3,
"value": "2026-04-16"
}
],
"author": [
{
"ORCID": "https://orcid.org/0000-0003-1695-7512",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"authenticated-orcid": false,
"family": "Raheja",
"given": "Harsha",
"sequence": "first"
},
{
"ORCID": "https://orcid.org/0009-0004-1027-1258",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"authenticated-orcid": false,
"family": "Sahu",
"given": "Risabh",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/057y6sk36",
"id-type": "ROR"
}
],
"name": "National Institute of Biomedical Genomics",
"place": [
"Kalyani, India"
]
}
],
"family": "Ghosh",
"given": "Trinath",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"family": "Paul",
"given": "Santu",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"family": "Rani",
"given": "Priya",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"family": "Aneja",
"given": "Ashish",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"family": "George",
"given": "Biju",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Centre for Infectious Disease Research, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"family": "Khatun",
"given": "Oyahida",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-5702-9248",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Centre for Infectious Disease Research, Indian Institute of Science",
"place": [
"Bangalore, India"
]
}
],
"authenticated-orcid": true,
"family": "Tripathi",
"given": "Shashank",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-0640-3586",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05j873a45",
"id-type": "ROR"
}
],
"name": "Department of Microbiology and Cell Biology, Indian Institute of Science",
"place": [
"Bangalore, India"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/057y6sk36",
"id-type": "ROR"
}
],
"name": "National Institute of Biomedical Genomics",
"place": [
"Kalyani, India"
]
}
],
"authenticated-orcid": true,
"family": "Das",
"given": "Saumitra",
"sequence": "additional"
}
],
"container-title": "Journal of Virology",
"container-title-short": "J Virol",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"journals.asm.org"
]
},
"created": {
"date-parts": [
[
2026,
4,
16
]
],
"date-time": "2026-04-16T14:01:31Z",
"timestamp": 1776348091000
},
"deposited": {
"date-parts": [
[
2026,
4,
16
]
],
"date-time": "2026-04-16T14:01:33Z",
"timestamp": 1776348093000
},
"editor": [
{
"affiliation": [],
"family": "Wobus",
"given": "Christiane E.",
"sequence": "additional"
}
],
"indexed": {
"date-parts": [
[
2026,
4,
16
]
],
"date-time": "2026-04-16T14:54:28Z",
"timestamp": 1776351268696,
"version": "3.51.2"
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2026,
4,
16
]
]
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
4,
16
]
],
"date-time": "2026-04-16T00:00:00Z",
"timestamp": 1776297600000
}
},
{
"URL": "https://journals.asm.org/non-commercial-tdm-license",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
4,
16
]
],
"date-time": "2026-04-16T00:00:00Z",
"timestamp": 1776297600000
}
}
],
"link": [
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/jvi.00276-26",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/jvi.00276-26",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "235",
"original-title": [],
"prefix": "10.1128",
"published": {
"date-parts": [
[
2026,
4,
16
]
]
},
"published-online": {
"date-parts": [
[
2026,
4,
16
]
]
},
"publisher": "American Society for Microbiology",
"reference": [
{
"DOI": "10.1038/s41579-020-00468-6",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_2_2"
},
{
"DOI": "10.3390/v13060952",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_3_2"
},
{
"DOI": "10.3390/v13112172",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_4_2"
},
{
"DOI": "10.1016/j.molcel.2021.05.023",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_5_2"
},
{
"DOI": "10.1038/s41586-020-2286-9",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_6_2"
},
{
"DOI": "10.1016/j.molcel.2011.06.007",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_7_2"
},
{
"DOI": "10.1016/j.molcel.2011.06.008",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_8_2"
},
{
"DOI": "10.1002/wrna.1372",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_9_2"
},
{
"DOI": "10.1128/JVI.01714-15",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_10_2"
},
{
"DOI": "10.1371/journal.ppat.1011552",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_11_2"
},
{
"DOI": "10.1128/JVI.00915-21",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_12_2"
},
{
"DOI": "10.1016/j.bbrc.2017.12.036",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_13_2"
},
{
"DOI": "10.1016/j.chom.2010.07.003",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_14_2"
},
{
"DOI": "10.1186/1742-4690-5-47",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_15_2"
},
{
"DOI": "10.1016/j.compbiomed.2022.105575",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_16_2"
},
{
"DOI": "10.1080/15476286.2020.1814556",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_17_2"
},
{
"DOI": "10.1093/nar/gkq1069",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_18_2"
},
{
"DOI": "10.1111/cei.13110",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_19_2"
},
{
"DOI": "10.1038/s41467-020-19883-7",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_20_2"
},
{
"DOI": "10.1038/s41467-023-39091-3",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_21_2"
},
{
"DOI": "10.3390/v13101923",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_22_2"
},
{
"DOI": "10.1038/s41467-022-32216-0",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_23_2"
},
{
"DOI": "10.1016/S1473-3099(22)00001-9",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_24_2"
},
{
"DOI": "10.1101/2020.10.14.339515",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_25_2",
"unstructured": "Tidu A Janvier A Schaeffer L Sosnowski P Kuhn L Hammann P Westhof E Eriani G Martin F. 2020. The viral protein NSP1 acts as a ribosome gatekeeper for shutting down host translation and fostering SARS-CoV-2 translation. Molecular Biology. doi:10.1101/2020.10.14.339515"
},
{
"DOI": "10.1038/s41594-020-0511-8",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_26_2"
},
{
"DOI": "10.1101/2022.10.18.512708",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_27_2",
"unstructured": "Kehrer T Cupic A Ye C Yildiz S Bouhhadou M Crossland NA Barrall E Cohen P Tseng A Çağatay T et al.. 2022. Impact of SARS-CoV-2 ORF6 and its variant polymorphisms on host responses and viral pathogenesis. bioRxiv:2022.10.18.512708. doi:10.1101/2022.10.18.512708"
},
{
"DOI": "10.1016/j.cell.2023.09.002",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_28_2"
},
{
"DOI": "10.2741/3921",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_29_2"
},
{
"DOI": "10.1016/j.isci.2021.102477",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_30_2"
},
{
"DOI": "10.1016/j.virusres.2018.09.006",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_31_2"
},
{
"DOI": "10.1016/j.addr.2021.114068",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_32_2"
},
{
"DOI": "10.1016/j.addr.2022.114442",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_33_2"
},
{
"DOI": "10.1128/AAC.01168-19",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_34_2"
},
{
"DOI": "10.1128/AAC.00900-20",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_35_2"
},
{
"DOI": "10.1038/s41594-021-00570-0",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_36_2"
},
{
"DOI": "10.1038/s42003-023-04789-z",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_37_2"
},
{
"DOI": "10.1016/j.virusres.2023.199221",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_38_2"
},
{
"DOI": "10.1016/j.virol.2020.05.015",
"doi-asserted-by": "publisher",
"key": "e_1_3_3_39_2"
}
],
"reference-count": 38,
"references-count": 38,
"relation": {},
"resource": {
"primary": {
"URL": "https://journals.asm.org/doi/10.1128/jvi.00276-26"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "HuR enhances SARS-CoV-2 non-structural protein translation through the genomic 5′-UTR, by promoting polypyrimidine tract-binding protein binding",
"type": "journal-article",
"update-policy": "https://doi.org/10.1128/asmj-crossmark-policy-page"
}