Discovery of key regulators in classical monocyte phenotypes linked to COVID-19 severity using single-cell multi-omics sequencing
et al., iScience, doi:10.1016/j.isci.2026.114849, Jan 2026
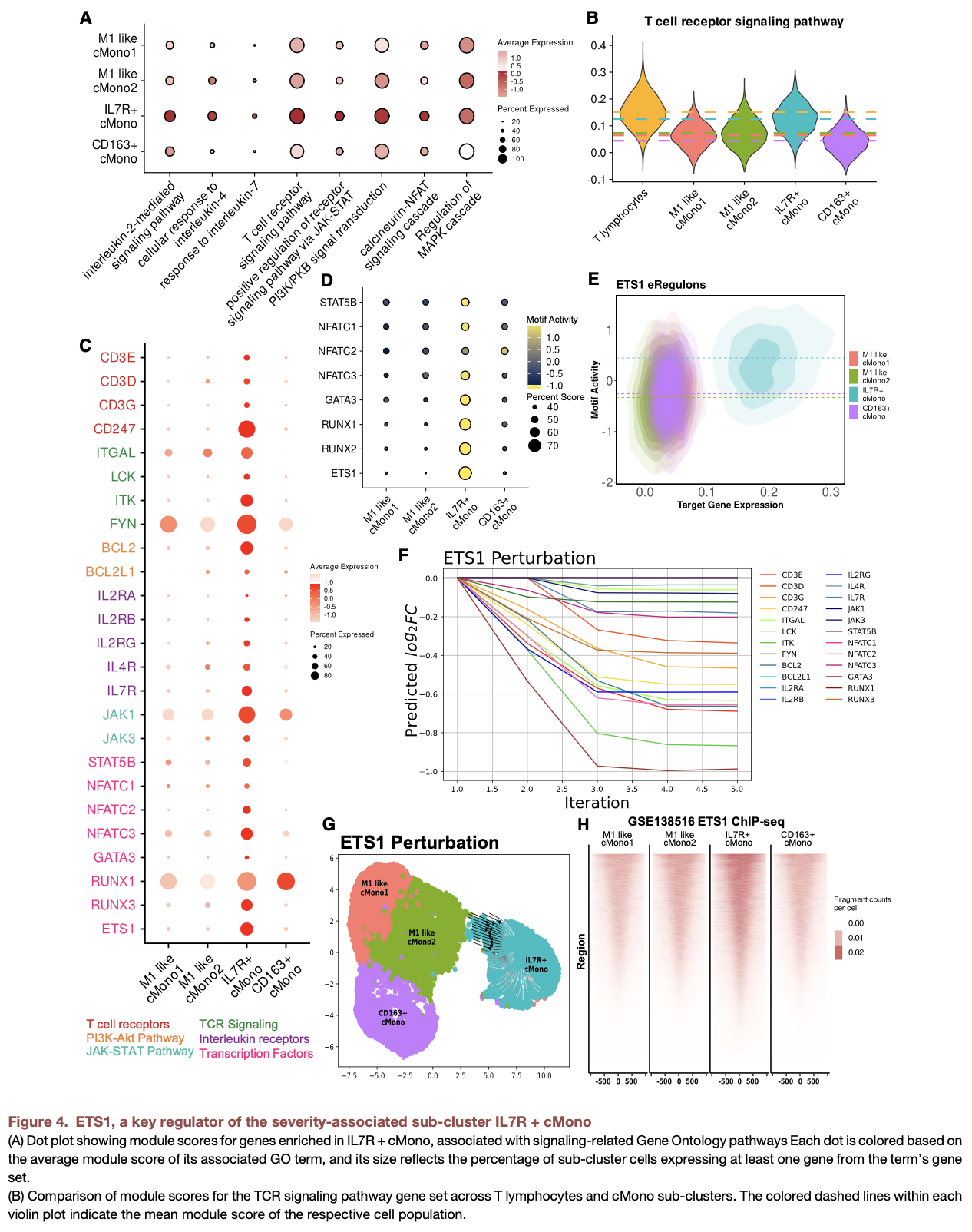
In vitro and clinical study showing that ETS1 and JDP2 are key transcriptional regulators driving COVID-19 severity-associated monocyte phenotypes.
Kim et al., 29 Jan 2026, multiple countries, peer-reviewed, 6 authors.
Contact: tae@jejunu.ac.kr (corresponding author), tae@jejunu.ac.kr (corresponding author), jihwan.park@gist.ac.kr.
Discovery of key regulators in classical monocyte phenotypes linked to COVID-19 severity using single-cell multi-omics sequencing
iScience, doi:10.1016/j.isci.2026.114849
severity-associated subtypes (IL7R+ cMono and CD163+ cMono) are identified • In IL7R+ cMono, ETS1 drives T cell-like signaling and immune suppressive programs
AUTHOR CONTRIBUTIONS This study was designed by J.P. and E.T.K.; J.R.Y. collected PBMCs from COVID-19 patients.; S.Y.C. performed the single-cell multi-omics sequencing library construction.; Data analysis was performed by D.K.; J.P. guided the process of data analysis.; C.Y.K. performed the experiments under the supervision of E.K.; J.P. and D.K. wrote the original draft of the manuscript. All authors read and approved the final manuscript.
DECLARATION OF INTERESTS S.Y.C. is an employee of Eyeoncell, and J.P. holds a leadership position at Eyeoncell.
DECLARATION OF GENERATIVE AI AND AI-ASSISTED TECHNOLOGIES IN THE WRITING PROCESS No generative AI or AI-assisted technologies were used in the writing or preparation of this manuscript.
STAR★METHODS Detailed methods are provided in the online version of this paper and include the following: • METHOD DETAILS ○ Preparation of human peripheral blood mononuclear cells (PBMCs) from COVID-19 patients ○ Single-cell multi-omics library preparation ○ Preprocessing of single-cell multi-omics data and quality check for obtaining high-quality cells ○ Demultiplexing of donors using genetic variants and doublet filtering ○ Dimension reduction and batch effect correction ○ Peak calling ○ Cell type and RNA-ATAC correlation ○ Differential abundance analysis of KNN graph in COVID-19 infection ○ Gene ontology term and module score analysis ○ Cell-cell interaction analysis using CellphoneDB ○ Gene set enrichment analysis ○ Genomic annotation..
References
Al-Aly, Davis, Mccorkell, Soares, Wulf-Hanson et al., Long COVID science, research and policy, Nat. Med, doi:10.1038/s41591-024-03173-6
Al-Mossawi, Yager, Taylor, Lau, Danielli et al., Context-specific regulation of surface and soluble IL7R expression by an autoimmune risk allele, Nat. Commun, doi:10.1038/s41467-019-12393-1
Alfaro, Casitas, Diaz-Garcia, Garcia-Tovar, Galera et al., TGF-beta1 overexpression in severe COVID-19 survivors and its implications for early-phase fibrotic abnormalities and long-term functional impairment, Front. Immunol, doi:10.3389/fimmu.2024.1401015
Aronheim, Zandi, Hennemann, Elledge, Isolation of an AP-1 repressor by a novel method for detecting protein-protein interactions, Mol. Cell Biol, doi:10.1128/mcb.17.6.3094
Ashburner, Ball, Blake, Botstein, Butler et al., Gene ontology: tool for the unification of biology. The Gene Ontology Consortium, Nat. Genet, doi:10.1038/75556
Austermann, Roth, Barczyk-Kahlert, The Good and the Bad: Monocytes' and Macrophages' Diverse Functions in Inflammation, Cells, doi:10.3390/cells11121979
Badia-I-Mompel, Wessels, Mu ¨ Ller-Dott, Trimbour, Ramirez Flores et al., Gene regulatory network inference in the era of single-cell multi-omics, Nat. Rev. Genet, doi:10.1038/s41576-023-00618-5
Baysoy, Bai, Satija, Fan, The technological landscape and applications of single-cell multi-omics, Nat. Rev. Mol. Cell Biol, doi:10.1038/s41580-023-00615-w
Bhat, Thompson, Lindsten, June, Fujiwara et al., Reciprocal expression of human ETS1 and ETS2 genes during T-cell activation: regulatory role for the protooncogene ETS1, Proc. Natl. Acad. Sci. USA, doi:10.1073/pnas.87.10.3723
Bhattacharya, Insights from Transcriptomics: CD163(+) Profibrotic Lung Macrophages in COVID-19, Am. J. Respir. Cell Mol. Biol, doi:10.1165/rcmb.2022-0107TR
Chae, Bothwell, Canonical and Non-Canonical Wnt Signaling in Immune Cells, Trends Immunol, doi:10.1016/j.it.2018.08.006
Cheong, Ravishankar, Sharma, Parkhurst, Grassmann et al., Epigenetic memory of coronavirus infection in innate immune cells and their progenitors, Cell, doi:10.1016/j.cell.2023.07.019
Chevrier, Zurbuchen, Cervia, Adamo, Raeber et al., A distinct innate immune signature marks progression from mild to severe COVID-19, Cell Rep. Med, doi:10.1016/j.xcrm.2020.100166
Choi, Park, Kim, Kim, Kim et al., Wnt5a and Wnt11 as acute respiratory distress syndrome biomarkers for severe acute respiratory syndrome coronavirus 2 patients, Eur. Respir. J, doi:10.1183/13993003.01531-2020
Clevers, Nusse, Wnt/beta-catenin signaling and disease, Cell, doi:10.1016/j.cell.2012.05.012
Coccia, Del Russo, Stellacci, Testa, Marziali et al., STAT1 activation during monocyte to macrophage matu-ration: role of adhesion molecules, Int. Immunol, doi:10.1093/intimm/11.7.1075
Dann, Henderson, Teichmann, Morgan, Marioni, Differential abundance testing on single-cell data using k-nearest neighbor graphs, Nat. Biotechnol, doi:10.1038/s41587-021-01033-z
Edahiro, Shirai, Takeshima, Sakakibara, Yamaguchi et al., Singlecell analyses and host genetics highlight the role of innate immune cells in COVID-19 severity, Nat. Genet, doi:10.1038/s41588-023-01375-1
Eun, Kim, Shin, Yang, Kim et al., Chromatin accessibility analysis and architectural profiling of human kidneys reveal key cell types and a regulator of diabetic kidney disease, Kidney Int, doi:10.1016/j.kint.2023.09.030
Farooq, Khan, Kim, Choi, The Role of Fibroblast Growth Factor (FGF) Signaling in Tissue Repair and Regeneration, Cells, doi:10.3390/cells10113242
Frangogiannis, Transforming growth factor-beta in tissue fibrosis, J. Exp. Med, doi:10.1084/jem.20190103
Gaud, Lesourne, Love, Regulatory mechanisms in T cell receptor signalling, Nat. Rev. Immunol, doi:10.1038/s41577-018-0020-8
Germain, Lun, Garcia Meixide, Macnair, Robinson, Doublet identification in single-cell sequencing data using scDblFinder, F1000Res, doi:10.12688/f1000research.73600.2
Giroux, Ding, Mcclain, Burke, Petzold et al., Differential chromatin accessibility in peripheral blood mononuclear cells underlies COVID-19 disease severity prior to seroconversion, Sci. Rep, doi:10.1038/s41598-022-15668-8
Gonzalez-Blas, De Winter, Hulselmans, Hecker, Matetovici et al., SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks, Nat. Methods, doi:10.1038/s41592-023-01938-4
Gu, Liu, Zhang, Xu, COVID-19 and trained immunity: the inflammatory burden of long covid, Front. Immunol, doi:10.3389/fimmu.2023.1294959
Hall, Barr, Delamare, Burkholder, Tsai et al., Profiling migration of human monocytes in response to chemotactic and barotactic guidance cues, Cell Rep. Methods, doi:10.1016/j.crmeth.2024.100846
Hao, Stuart, Kowalski, Choudhary, Hoffman et al., Dictionary learning for integrative, multimodal and scalable single-cell analysis, Nat. Biotechnol, doi:10.1038/s41587-023-01767-y
He, Gao, Xu, Lu, Wang et al., Analysis of IL-6 and IL-1beta release in cryopreserved pooled human whole blood stimulated with endotoxin, Innate Immun, doi:10.1177/1753425918777596
Ho, Tai, Pai, GATA3 and the T-cell lineage: essential functions before and after T-helper-2-cell differentiation, Nat. Rev. Immunol, doi:10.1038/nri2476
Hoffmann, Weinberg, Swaminathan, Chaudhuri, Almubarak et al., Unique molecular signatures sustained in circulating monocytes and regulatory T cells in convalescent COVID-19 patients, Clin. Immunol, doi:10.1016/j.clim.2023.109634
Hou, Shi, Chen, Lv, Chen et al., M2 macrophages promote myofibroblast differentiation of LR-MSCs and are associated with pulmonary fibrogenesis, Cell Commun. Signal, doi:10.1186/s12964-018-0300-8
Huisman, Guan, Ru ¨ Ckert, Garner, Singh et al., Highthroughput characterization of HLA-E-presented CD94/NKG2x ligands reveals peptides which modulate NK cell activation, Nat. Commun, doi:10.1038/s41467-023-40220-1
Hwang, Byeon, Kim, Park, Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development, Exp. Mol. Med, doi:10.1038/s12276-020-0435-8
Kang, Subramaniam, Targ, Nguyen, Maliskova et al., Multiplexed droplet single-cell RNA-sequencing using natural genetic variation, Nat. Biotechnol, doi:10.1038/nbt.4042
Katz, Heinrich, Aronheim, The AP-1 repressor, JDP2, is a bona fide substrate for the c-Jun N-terminal kinase, FEBS Lett, doi:10.1016/s0014-5793(01)02907-6
Kawaida, Ohtsuka, Okutsu, Takahashi, Kadono et al., dimerization protein 2 (JDP2), a member of the AP-1 family of transcription factor, mediates osteoclast differentiation induced by RANKL, J. Exp. Med, doi:10.1084/jem.20021321
Kieslinger, Woldman, Moriggl, Hofmann, Marine et al., Antiapoptotic activity of Stat5 required during terminal stages of myeloid differentiation, Genes Dev
Kim, Lim, Park, A practical handbook on single-cell RNA sequencing data quality control and downstream analysis, Mol. Cells, doi:10.1016/j.mocell.2024.100103
Kim, Park, Single-cell transcriptomics: a novel precision medicine technique in nephrology, Korean J. Intern. Med, doi:10.3904/kjim.2020.415
Kim, Shin, Jung, An, Choi et al., Cell Type-and Age-Specific Expression of lncRNAs across Kidney Cell Types, J. Am. Soc. Nephrol, doi:10.1681/ASN.0000000000000354
Kimura, Rieger, Simone, Chen, Wickre et al., The transcription factors STAT5A/B regulate GM-CSF-mediated granulopoiesis, Blood, doi:10.1182/blood-2009-04-216390
Kishimoto, Kirino, Tamura, Takeno, Kunishita et al., Dysregulated heme oxygenase-1(low) M2-like macrophages augment lupus nephritis via Bach1 induced by type I interferons, Arthritis Res. Ther, doi:10.1186/s13075-018-1568-1
Ko, Chang, Wang, Biological roles of CCAAT/ Enhancer-binding protein delta during inflammation, J. Biomed. Sci, doi:10.1186/s12929-014-0110-2
Korsunsky, Millard, Fan, Slowikowski, Zhang et al., Fast, sensitive and accurate integration of single-cell data with Harmony, Nat. Methods, doi:10.1038/s41592-019-0619-0
Laabi, Gras, Brouet, Berger, Larsen et al., The BCMA gene, preferentially expressed during B lymphoid maturation, is bidirectionally transcribed, Nucleic Acids Res, doi:10.1093/nar/22.7.1147
Lazar, Sandulescu, Barbu, Chitu-Tisu, Andreescu et al., The Role of Cytokines and Molecular Pathways in Lung Fibrosis Following SARS-CoV-2 Infection: A Physiopathologic (Re)view, Biomedicines, doi:10.3390/biomedicines12030639
Lee, Shin, An, Choi, Park, Multi-Omics Single-Cell Analysis Reveals Key Regulators of HIV-1 Persistence and Aberrant Host Immune Responses in Early Infection, eLife
Lerdrup, Holmberg, Dietrich, Shaulian, Herdegen et al., Depletion of the AP-1 repressor JDP2 induces cell death similar to apoptosis, Biochim. Biophys. Acta, doi:10.1016/j.bbamcr.2005.06.008
Li, A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data, Bioinformatics, doi:10.1093/bioinformatics/btr509
Li, Kong, He, Liu, Peng et al., Recent developments in application of single-cell RNA sequencing in the tumour immune microenvironment and cancer therapy, Mil. Med. Res, doi:10.1186/s40779-022-00414-y
Liao, Liu, Yuan, Wen, Xu et al., Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19, Nat. Med, doi:10.1038/s41591-020-0901-9
Liu, Li, Empowering biologists to decode omics data: the Genekitr R package and web server, BMC Bioinf, doi:10.1186/s12859-023-05342-9
Luque-Martin, Angell, Kalxdorf, Bernard, Thompson et al., IFN-gamma Drives Human Monocyte Differentiation into Highly Proinflammatory Macrophages That Resemble a Phenotype Relevant to Psoriasis, J. Immunol, doi:10.4049/jimmunol.2001310
Mccarter, Della Gatta, Melnick, Kim, Sha et al., Combinatorial ETS1-dependent control of oncogenic NOTCH1 enhancers in T-cell leukemia, Blood Cancer Discov, doi:10.1158/2643-3230.BCD-20-0026
Mcginnis, Murrow, Gartner, DoubletFinder: Doublet Detection in Single-Cell RNA Sequencing Data Using Artificial Nearest Neighbors, Cell Syst, doi:10.1016/j.cels.2019.03.003
Murphy, Cox, Connolly, Breen, Brugman et al., Trained immunity is induced in humans after immunization with an adenoviral vector COVID-19 vaccine, J. Clin. Investig, doi:10.1172/JCI162581
Obata, Furusawa, Hase, Epigenetic modifications of the immune system in health and disease, Immunol. Cell Biol, doi:10.1038/icb.2014.114
Organization, COVID-19 Therapeutic Trial Synopsis
Park, Lee, Macrophages and Wnts in Tissue Injury and Repair, Cells, doi:10.3390/cells11223592
Ramirez, Patel, Pastar, The Role of TGFbeta Signaling in Wound Epithelialization, Adv. Wound Care, doi:10.1089/wound.2013.0466
Rigamonti, Villar, Segura, Monocyte differentiation within tissues: a renewed outlook, Trends Immunol, doi:10.1016/j.it.2023.10.005
Schaale, Brandenburg, Kispert, Leitges, Ehlers et al., Wnt6 is expressed in granulomatous lesions of Mycobacterium tuberculosis-infected mice and is involved in macrophage differentiation and proliferation, J. Immunol, doi:10.4049/jimmunol.1201819
Schep, Wu, Buenrostro, Greenleaf, chrom-VAR: inferring transcription-factor-associated accessibility from singlecell epigenomic data, Nat. Methods, doi:10.1038/nmeth.4401
Schmierer, Hill, TGFbeta-SMAD signal transduction: molecular specificity and functional flexibility, Nat. Rev. Mol. Cell Biol, doi:10.1038/nrm2297
Schulte-Schrepping, Reusch, Paclik, Baßler, Schlickeiser et al., Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment, Cell, doi:10.1016/j.cell.2020.08.001
Scott, Pearmain, Knight, Brand, Morgan et al., Monocyte migration profiles define disease severity in acute COVID-19 and unique features of long COVID, Eur. Respir. J, doi:10.1183/13993003.02226-2022
Simonis, Theobald, Koch, Mummadavarapu, Mudler et al., Persistent epigenetic memory of SARS-CoV-2 mRNA vaccination in monocyte-derived macrophages, Mol. Syst. Biol, doi:10.1038/s44320-025-00093-6
Stephenson, Reynolds, Botting, Calero-Nieto, Morgan et al., Single-cell multi-omics analysis of the immune response in COVID-19, Nat. Med, doi:10.1038/s41591-021-01329-2
Stuart, Srivastava, Madad, Lareau, Satija, Single-cell chromatin state analysis with Signac, Nat. Methods, doi:10.1038/s41592-021-01282-5
Su, Chen, Yuan, Lausted, Choi et al., Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19, Cell, doi:10.1016/j.cell.2020.10.037
Subramanian, Tamayo, Mootha, Mukherjee, Ebert et al., Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles, Proc. Natl. Acad. Sci, doi:10.1073/pnas.0506580102
Szabo, Levitin, Miron, Snyder, Senda et al., Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease, Nat. Commun, doi:10.1038/s41467-019-12464-3
Thomas, Tacke, Hedrick, Hanna, Nonclassical patrolling monocyte function in the vasculature, Arterioscler. Thromb. Vasc. Biol, doi:10.1161/ATVBAHA.114.304650
Tourki, Jia, Karampitsakos, Vera, Arsenault et al., Convergent and divergent immune aberrations in COVID-19, post-COVID-19-interstitial lung disease, and idiopathic pulmonary fibrosis, Am. J. Physiol. Cell Physiol, doi:10.1152/ajpcell.00528.2024
Troule, Petryszak, Cakir, Cranley, Harasty et al., CellPhoneDB v5: inferring cell-cell communication from singlecell multiomics data, Nat. Protoc, doi:10.1038/s41596-024-01137-1
Wan, GATA3: a master of many trades in immune regulation, Trends Immunol, doi:10.1016/j.it.2014.04.002
Wendisch, Dietrich, Mari, Von Stillfried, Ibarra et al., SARS-CoV-2 infection triggers profibrotic macrophage responses and lung fibrosis, Cell, doi:10.1016/j.cell.2021.11.033
Wilk, Lee, Wei, Parks, Pi et al., Multi-omic profiling reveals widespread dysregulation of innate immunity and hematopoiesis in COVID-19, J. Exp. Med, doi:10.1084/jem.20210582
Wu, Mcgoogan, Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72 314 Cases From the Chinese Center for Disease Control and Prevention, JAMA, doi:10.1001/jama.2020.2648
Wu, Nicoll, Ingham, AP-1 family transcription factors: a diverse family of proteins that regulate varied cellular activities in classical hodgkin lymphoma and ALK+ ALCL, Exp. Hematol. Oncol, doi:10.1186/s40164-020-00197-9
Xu, Elaish, Wong, Hassan, Lopez-Orozco et al., The Wnt/β-catenin pathway is important for replication of SARS-CoV-2 and other pathogenic RNA viruses, npj Viruses, doi:10.1038/s44298-024-00018-4
Xu, Qi, Li, Yang, Wang et al., The differential immune responses to COVID-19 in peripheral and lung revealed by single-cell RNA sequencing, Cell Discov, doi:10.1038/s41421-020-00225-2
Yang, Hegde, Rao, Zhang, Nagarkatti et al., Histone modifications are associated with Delta9-tetrahydrocannabinol-mediated alterations in antigen-specific T cell responses, J. Biol. Chem, doi:10.1074/jbc.M113.545210
Yang, Rutkovsky, Zhou, Zhong, Reese et al., Characterization of Altered Gene Expression and Histone Methylation in Peripheral Blood Mononuclear Cells Regulating Inflammation in COVID-19 Patients, J. Immunol, doi:10.4049/jimmunol.2101099
Yeamans, Wang, Paz-Priel, Torbett, Tenen et al., C/EBPalpha binds and activates the PU.1 distal enhancer to induce monocyte lineage commitment, Blood, doi:10.1182/blood-2007-03-080291
Yoon, Kim, Jung, Park, Single-cell lineage tracing approaches to track kidney cell development and maintenance, Kidney Int, doi:10.1016/j.kint.2024.01.045
You, Chen, Zhang, Zhao, Chen et al., Single-cell epigenomic landscape of peripheral immune cells reveals establishment of trained immunity in individuals convalescing from COVID-19, Nat. Cell Biol, doi:10.1038/s41556-021-00690-1
Yu, Wang, He, ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization, Bioinformatics, doi:10.1093/bioinformatics/btv145
Yue, Shan, Lasky, TGF-beta: Titan of Lung Fibrogenesis, Curr. Enzym. Inhib, doi:10.2174/10067
Zhang, Liu, Meyer, Eeckhoute, Johnson et al., Model-based analysis of ChIP-Seq (MACS), Genome Biol, doi:10.1186/gb-2008-9-9-r137
Zhang, Wang, Xing, Xu, Zhang et al., Single-cell landscape of immunological responses in patients with COVID-19, Nat. Immunol, doi:10.1038/s41590-020-0762-x
Zhang, Zhang, Xiong, Li, Zhang et al., CD127 imprints functional heterogeneity to diversify monocyte responses in inflammatory diseases, J. Exp. Med, doi:10.1084/jem.20211191
Zhao, Bartholdy, Yamamoto, Evans, Alberich-Jorda et al., PU.1-c-Jun interaction is crucial for PU.1 function in myeloid development, Commun. Biol, doi:10.1038/s42003-022-03888-7
Zhao, Li, Li, Xu, Xu, The mechanism of multiple organ dysfunction syndrome in patients with COVID-19, J. Med. Virol, doi:10.1002/jmv.27627
Zheng, Dan, Huo, Zhang, Hou, Characteristics of the early innate response induced by the aerosolized Ad5vectored COVID-19 vaccine, Mol. Biomed, doi:10.1186/s43556-024-00232-9
Zheng, Terry, Belgrader, Ryvkin, Bent et al., Massively parallel digital transcriptional profiling of single cells, Nat. Commun, doi:10.1038/ncomms14049
Zizzo, Cohen, The PPAR-gamma antagonist GW9662 elicits differentiation of M2c-like cells and upregulation of the MerTK/Gas6 axis: a key role for PPAR-gamma in human macrophage polarization, J. Inflamm, doi:10.1186/s12950-015-0081-4
DOI record:
{
"DOI": "10.1016/j.isci.2026.114849",
"ISSN": [
"2589-0042"
],
"URL": "http://dx.doi.org/10.1016/j.isci.2026.114849",
"alternative-id": [
"S2589004226002245"
],
"article-number": "114849",
"assertion": [
{
"label": "This article is maintained by",
"name": "publisher",
"value": "Elsevier"
},
{
"label": "Article Title",
"name": "articletitle",
"value": "Discovery of key regulators in classical monocyte phenotypes linked to COVID-19 severity using single-cell multi-omics sequencing"
},
{
"label": "Journal Title",
"name": "journaltitle",
"value": "iScience"
},
{
"label": "CrossRef DOI link to publisher maintained version",
"name": "articlelink",
"value": "https://doi.org/10.1016/j.isci.2026.114849"
},
{
"label": "Content Type",
"name": "content_type",
"value": "article"
},
{
"label": "Copyright",
"name": "copyright",
"value": "© 2026 The Author(s). Published by Elsevier Inc."
}
],
"author": [
{
"affiliation": [],
"family": "Kim",
"given": "Donggun",
"sequence": "first"
},
{
"affiliation": [],
"family": "Choi",
"given": "Sin Young",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Kim",
"given": "Chae Yeon",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Yoo",
"given": "Jeong Rae",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Kim",
"given": "Eui Tae",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-5728-912X",
"affiliation": [],
"authenticated-orcid": false,
"family": "Park",
"given": "Jihwan",
"sequence": "additional"
}
],
"container-title": "iScience",
"container-title-short": "iScience",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"cell.com",
"elsevier.com",
"sciencedirect.com"
]
},
"created": {
"date-parts": [
[
2026,
1,
30
]
],
"date-time": "2026-01-30T00:42:49Z",
"timestamp": 1769733769000
},
"deposited": {
"date-parts": [
[
2026,
2,
27
]
],
"date-time": "2026-02-27T07:12:28Z",
"timestamp": 1772176348000
},
"funder": [
{
"DOI": "10.13039/501100003653",
"award": [
"2025-ER1606-00"
],
"award-info": [
{
"award-number": [
"2025-ER1606-00"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003653",
"id-type": "DOI"
}
],
"name": "Korea National Institute of Health"
},
{
"DOI": "10.13039/501100003653",
"award": [
"2024-ER1605-00"
],
"award-info": [
{
"award-number": [
"2024-ER1605-00"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003653",
"id-type": "DOI"
}
],
"name": "Korea National Institute of Health"
},
{
"DOI": "10.13039/501100003653",
"award": [
"2022-ER1603-00"
],
"award-info": [
{
"award-number": [
"2022-ER1603-00"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003653",
"id-type": "DOI"
}
],
"name": "Korea National Institute of Health"
},
{
"DOI": "10.13039/501100003625",
"award": [
"RS-2024-00466906"
],
"award-info": [
{
"award-number": [
"RS-2024-00466906"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003625",
"id-type": "DOI"
}
],
"name": "Korea Ministry of Health and Welfare"
},
{
"DOI": "10.13039/501100002701",
"award": [
"RS-2023-00270936"
],
"award-info": [
{
"award-number": [
"RS-2023-00270936"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100002701",
"id-type": "DOI"
}
],
"name": "Korea Ministry of Education"
},
{
"DOI": "10.13039/501100014188",
"award": [
"RS-2024-00403622"
],
"award-info": [
{
"award-number": [
"RS-2024-00403622"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100014188",
"id-type": "DOI"
}
],
"name": "Korea Ministry of Science and ICT"
},
{
"DOI": "10.13039/501100014188",
"award": [
"RS-2024-00352590"
],
"award-info": [
{
"award-number": [
"RS-2024-00352590"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100014188",
"id-type": "DOI"
}
],
"name": "Korea Ministry of Science and ICT"
},
{
"DOI": "10.13039/501100003725",
"award": [
"RS-2024-00335026"
],
"award-info": [
{
"award-number": [
"RS-2024-00335026"
]
}
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003725",
"id-type": "DOI"
}
],
"name": "National Research Foundation of Korea"
},
{
"DOI": "10.13039/501100003716",
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/501100003716",
"id-type": "DOI"
}
],
"name": "Korea Basic Science Institute"
}
],
"indexed": {
"date-parts": [
[
2026,
2,
27
]
],
"date-time": "2026-02-27T08:26:14Z",
"timestamp": 1772180774415,
"version": "3.50.1"
},
"is-referenced-by-count": 0,
"issue": "3",
"issued": {
"date-parts": [
[
2026,
3
]
]
},
"journal-issue": {
"issue": "3",
"published-print": {
"date-parts": [
[
2026,
3
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://www.elsevier.com/tdm/userlicense/1.0/",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
3,
1
]
],
"date-time": "2026-03-01T00:00:00Z",
"timestamp": 1772323200000
}
},
{
"URL": "https://www.elsevier.com/legal/tdmrep-license",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
3,
1
]
],
"date-time": "2026-03-01T00:00:00Z",
"timestamp": 1772323200000
}
},
{
"URL": "http://creativecommons.org/licenses/by-nc-nd/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
1,
27
]
],
"date-time": "2026-01-27T00:00:00Z",
"timestamp": 1769472000000
}
}
],
"link": [
{
"URL": "https://api.elsevier.com/content/article/PII:S2589004226002245?httpAccept=text/xml",
"content-type": "text/xml",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://api.elsevier.com/content/article/PII:S2589004226002245?httpAccept=text/plain",
"content-type": "text/plain",
"content-version": "vor",
"intended-application": "text-mining"
}
],
"member": "78",
"original-title": [],
"page": "114849",
"prefix": "10.1016",
"published": {
"date-parts": [
[
2026,
3
]
]
},
"published-print": {
"date-parts": [
[
2026,
3
]
]
},
"publisher": "Elsevier BV",
"reference": [
{
"DOI": "10.1001/jama.2020.2648",
"article-title": "Characteristics of and Important Lessons From the Coronavirus Disease 2019 (COVID-19) Outbreak in China: Summary of a Report of 72 314 Cases From the Chinese Center for Disease Control and Prevention",
"author": "Wu",
"doi-asserted-by": "crossref",
"first-page": "1239",
"journal-title": "JAMA",
"key": "10.1016/j.isci.2026.114849_bib1",
"volume": "323",
"year": "2020"
},
{
"DOI": "10.1038/s41591-024-03173-6",
"article-title": "Long COVID science, research and policy",
"author": "Al-Aly",
"doi-asserted-by": "crossref",
"first-page": "2148",
"journal-title": "Nat. Med.",
"key": "10.1016/j.isci.2026.114849_bib2",
"volume": "30",
"year": "2024"
},
{
"DOI": "10.1002/jmv.27627",
"article-title": "The mechanism of multiple organ dysfunction syndrome in patients with COVID-19",
"author": "Zhao",
"doi-asserted-by": "crossref",
"first-page": "1886",
"journal-title": "J. Med. Virol.",
"key": "10.1016/j.isci.2026.114849_bib3",
"volume": "94",
"year": "2022"
},
{
"DOI": "10.1038/icb.2014.114",
"article-title": "Epigenetic modifications of the immune system in health and disease",
"author": "Obata",
"doi-asserted-by": "crossref",
"first-page": "226",
"journal-title": "Immunol. Cell Biol.",
"key": "10.1016/j.isci.2026.114849_bib4",
"volume": "93",
"year": "2015"
},
{
"DOI": "10.1074/jbc.M113.545210",
"article-title": "Histone modifications are associated with Delta9-tetrahydrocannabinol-mediated alterations in antigen-specific T cell responses",
"author": "Yang",
"doi-asserted-by": "crossref",
"first-page": "18707",
"journal-title": "J. Biol. Chem.",
"key": "10.1016/j.isci.2026.114849_bib5",
"volume": "289",
"year": "2014"
},
{
"DOI": "10.3389/fimmu.2023.1294959",
"article-title": "COVID-19 and trained immunity: the inflammatory burden of long covid",
"author": "Gu",
"doi-asserted-by": "crossref",
"journal-title": "Front. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib6",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.4049/jimmunol.2101099",
"article-title": "Characterization of Altered Gene Expression and Histone Methylation in Peripheral Blood Mononuclear Cells Regulating Inflammation in COVID-19 Patients",
"author": "Yang",
"doi-asserted-by": "crossref",
"first-page": "1968",
"journal-title": "J. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib7",
"volume": "208",
"year": "2022"
},
{
"DOI": "10.1038/s41556-021-00690-1",
"article-title": "Single-cell epigenomic landscape of peripheral immune cells reveals establishment of trained immunity in individuals convalescing from COVID-19",
"author": "You",
"doi-asserted-by": "crossref",
"first-page": "620",
"journal-title": "Nat. Cell Biol.",
"key": "10.1016/j.isci.2026.114849_bib8",
"volume": "23",
"year": "2021"
},
{
"DOI": "10.1038/s44320-025-00093-6",
"article-title": "Persistent epigenetic memory of SARS-CoV-2 mRNA vaccination in monocyte-derived macrophages",
"author": "Simonis",
"doi-asserted-by": "crossref",
"first-page": "341",
"journal-title": "Mol. Syst. Biol.",
"key": "10.1016/j.isci.2026.114849_bib9",
"volume": "21",
"year": "2025"
},
{
"article-title": "Profiling migration of human monocytes in response to chemotactic and barotactic guidance cues",
"author": "Hall",
"journal-title": "Cell Rep. Methods",
"key": "10.1016/j.isci.2026.114849_bib10",
"volume": "4",
"year": "2024"
},
{
"DOI": "10.1183/13993003.02226-2022",
"article-title": "Monocyte migration profiles define disease severity in acute COVID-19 and unique features of long COVID",
"author": "Scott",
"doi-asserted-by": "crossref",
"journal-title": "Eur. Respir. J.",
"key": "10.1016/j.isci.2026.114849_bib11",
"volume": "61",
"year": "2023"
},
{
"DOI": "10.3390/cells11121979",
"article-title": "The Good and the Bad: Monocytes' and Macrophages' Diverse Functions in Inflammation",
"author": "Austermann",
"doi-asserted-by": "crossref",
"journal-title": "Cells",
"key": "10.1016/j.isci.2026.114849_bib12",
"volume": "11",
"year": "2022"
},
{
"DOI": "10.1016/j.it.2023.10.005",
"article-title": "Monocyte differentiation within tissues: a renewed outlook",
"author": "Rigamonti",
"doi-asserted-by": "crossref",
"first-page": "999",
"journal-title": "Trends Immunol.",
"key": "10.1016/j.isci.2026.114849_bib13",
"volume": "44",
"year": "2023"
},
{
"DOI": "10.1016/j.cell.2020.10.037",
"article-title": "Multi-Omics Resolves a Sharp Disease-State Shift between Mild and Moderate COVID-19",
"author": "Su",
"doi-asserted-by": "crossref",
"first-page": "1479",
"journal-title": "Cell",
"key": "10.1016/j.isci.2026.114849_bib14",
"volume": "183",
"year": "2020"
},
{
"DOI": "10.1038/s41591-021-01329-2",
"article-title": "Single-cell multi-omics analysis of the immune response in COVID-19",
"author": "Stephenson",
"doi-asserted-by": "crossref",
"first-page": "904",
"journal-title": "Nat. Med.",
"key": "10.1016/j.isci.2026.114849_bib15",
"volume": "27",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2023.07.019",
"article-title": "Epigenetic memory of coronavirus infection in innate immune cells and their progenitors",
"author": "Cheong",
"doi-asserted-by": "crossref",
"first-page": "3882",
"journal-title": "Cell",
"key": "10.1016/j.isci.2026.114849_bib16",
"volume": "186",
"year": "2023"
},
{
"DOI": "10.1084/jem.20211191",
"article-title": "CD127 imprints functional heterogeneity to diversify monocyte responses in inflammatory diseases",
"author": "Zhang",
"doi-asserted-by": "crossref",
"journal-title": "J. Exp. Med.",
"key": "10.1016/j.isci.2026.114849_bib17",
"volume": "219",
"year": "2022"
},
{
"DOI": "10.1016/j.cell.2020.08.001",
"article-title": "Severe COVID-19 Is Marked by a Dysregulated Myeloid Cell Compartment",
"author": "Schulte-Schrepping",
"doi-asserted-by": "crossref",
"first-page": "1419",
"journal-title": "Cell",
"key": "10.1016/j.isci.2026.114849_bib18",
"volume": "182",
"year": "2020"
},
{
"article-title": "Recent developments in application of single-cell RNA sequencing in the tumour immune microenvironment and cancer therapy",
"author": "Li",
"first-page": "52",
"journal-title": "Mil. Med. Res.",
"key": "10.1016/j.isci.2026.114849_bib19",
"volume": "9",
"year": "2022"
},
{
"DOI": "10.1016/j.kint.2024.01.045",
"article-title": "Single-cell lineage tracing approaches to track kidney cell development and maintenance",
"author": "Yoon",
"doi-asserted-by": "crossref",
"first-page": "1186",
"journal-title": "Kidney Int.",
"key": "10.1016/j.isci.2026.114849_bib20",
"volume": "105",
"year": "2024"
},
{
"DOI": "10.3904/kjim.2020.415",
"article-title": "Single-cell transcriptomics: a novel precision medicine technique in nephrology",
"author": "Kim",
"doi-asserted-by": "crossref",
"first-page": "479",
"journal-title": "Korean J. Intern. Med.",
"key": "10.1016/j.isci.2026.114849_bib21",
"volume": "36",
"year": "2021"
},
{
"DOI": "10.1016/j.kint.2023.09.030",
"article-title": "Chromatin accessibility analysis and architectural profiling of human kidneys reveal key cell types and a regulator of diabetic kidney disease",
"author": "Eun",
"doi-asserted-by": "crossref",
"first-page": "150",
"journal-title": "Kidney Int.",
"key": "10.1016/j.isci.2026.114849_bib22",
"volume": "105",
"year": "2024"
},
{
"DOI": "10.1038/s41580-023-00615-w",
"article-title": "The technological landscape and applications of single-cell multi-omics",
"author": "Baysoy",
"doi-asserted-by": "crossref",
"first-page": "695",
"journal-title": "Nat. Rev. Mol. Cell Biol.",
"key": "10.1016/j.isci.2026.114849_bib23",
"volume": "24",
"year": "2023"
},
{
"article-title": "Multi-Omics Single-Cell Analysis Reveals Key Regulators of HIV-1 Persistence and Aberrant Host Immune Responses in Early Infection",
"author": "Dayeon Lee",
"journal-title": "eLife",
"key": "10.1016/j.isci.2026.114849_bib24",
"volume": "14",
"year": "2025"
},
{
"DOI": "10.1038/s41576-023-00618-5",
"article-title": "Gene regulatory network inference in the era of single-cell multi-omics",
"author": "Badia-I-Mompel",
"doi-asserted-by": "crossref",
"first-page": "739",
"journal-title": "Nat. Rev. Genet.",
"key": "10.1016/j.isci.2026.114849_bib25",
"volume": "24",
"year": "2023"
},
{
"DOI": "10.1038/s41587-021-01033-z",
"article-title": "Differential abundance testing on single-cell data using k-nearest neighbor graphs",
"author": "Dann",
"doi-asserted-by": "crossref",
"first-page": "245",
"journal-title": "Nat. Biotechnol.",
"key": "10.1016/j.isci.2026.114849_bib27",
"volume": "40",
"year": "2022"
},
{
"DOI": "10.1038/s41590-020-0762-x",
"article-title": "Single-cell landscape of immunological responses in patients with COVID-19",
"author": "Zhang",
"doi-asserted-by": "crossref",
"first-page": "1107",
"journal-title": "Nat. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib28",
"volume": "21",
"year": "2020"
},
{
"DOI": "10.1152/ajpcell.00528.2024",
"article-title": "Convergent and divergent immune aberrations in COVID-19, post-COVID-19-interstitial lung disease, and idiopathic pulmonary fibrosis",
"author": "Tourki",
"doi-asserted-by": "crossref",
"first-page": "C199",
"journal-title": "Am. J. Physiol. Cell Physiol.",
"key": "10.1016/j.isci.2026.114849_bib29",
"volume": "328",
"year": "2025"
},
{
"DOI": "10.1084/jem.20210582",
"article-title": "Multi-omic profiling reveals widespread dysregulation of innate immunity and hematopoiesis in COVID-19",
"author": "Wilk",
"doi-asserted-by": "crossref",
"journal-title": "J. Exp. Med.",
"key": "10.1016/j.isci.2026.114849_bib30",
"volume": "218",
"year": "2021"
},
{
"DOI": "10.1038/s41467-023-40220-1",
"article-title": "High-throughput characterization of HLA-E-presented CD94/NKG2x ligands reveals peptides which modulate NK cell activation",
"author": "Huisman",
"doi-asserted-by": "crossref",
"first-page": "4809",
"journal-title": "Nat. Commun.",
"key": "10.1016/j.isci.2026.114849_bib31",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.1161/ATVBAHA.114.304650",
"article-title": "Nonclassical patrolling monocyte function in the vasculature",
"author": "Thomas",
"doi-asserted-by": "crossref",
"first-page": "1306",
"journal-title": "Arterioscler. Thromb. Vasc. Biol.",
"key": "10.1016/j.isci.2026.114849_bib32",
"volume": "35",
"year": "2015"
},
{
"DOI": "10.4049/jimmunol.2001310",
"article-title": "IFN-gamma Drives Human Monocyte Differentiation into Highly Proinflammatory Macrophages That Resemble a Phenotype Relevant to Psoriasis",
"author": "Luque-Martin",
"doi-asserted-by": "crossref",
"first-page": "555",
"journal-title": "J. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib33",
"volume": "207",
"year": "2021"
},
{
"DOI": "10.1093/nar/22.7.1147",
"article-title": "The BCMA gene, preferentially expressed during B lymphoid maturation, is bidirectionally transcribed",
"author": "Laabi",
"doi-asserted-by": "crossref",
"first-page": "1147",
"journal-title": "Nucleic Acids Res.",
"key": "10.1016/j.isci.2026.114849_bib34",
"volume": "22",
"year": "1994"
},
{
"DOI": "10.1177/1753425918777596",
"article-title": "Analysis of IL-6 and IL-1beta release in cryopreserved pooled human whole blood stimulated with endotoxin",
"author": "He",
"doi-asserted-by": "crossref",
"first-page": "316",
"journal-title": "Innate Immun.",
"key": "10.1016/j.isci.2026.114849_bib35",
"volume": "24",
"year": "2018"
},
{
"DOI": "10.1038/s41598-022-15668-8",
"article-title": "Differential chromatin accessibility in peripheral blood mononuclear cells underlies COVID-19 disease severity prior to seroconversion",
"author": "Giroux",
"doi-asserted-by": "crossref",
"journal-title": "Sci. Rep.",
"key": "10.1016/j.isci.2026.114849_bib36",
"volume": "12",
"year": "2022"
},
{
"DOI": "10.1038/s41588-023-01375-1",
"article-title": "Single-cell analyses and host genetics highlight the role of innate immune cells in COVID-19 severity",
"author": "Edahiro",
"doi-asserted-by": "crossref",
"first-page": "753",
"journal-title": "Nat. Genet.",
"key": "10.1016/j.isci.2026.114849_bib39",
"volume": "55",
"year": "2023"
},
{
"DOI": "10.1038/s41421-020-00225-2",
"article-title": "The differential immune responses to COVID-19 in peripheral and lung revealed by single-cell RNA sequencing",
"author": "Xu",
"doi-asserted-by": "crossref",
"first-page": "73",
"journal-title": "Cell Discov.",
"key": "10.1016/j.isci.2026.114849_bib40",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1038/s41467-019-12464-3",
"article-title": "Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease",
"author": "Szabo",
"doi-asserted-by": "crossref",
"first-page": "4706",
"journal-title": "Nat. Commun.",
"key": "10.1016/j.isci.2026.114849_bib41",
"volume": "10",
"year": "2019"
},
{
"DOI": "10.1038/s41467-019-12393-1",
"article-title": "Context-specific regulation of surface and soluble IL7R expression by an autoimmune risk allele",
"author": "Al-Mossawi",
"doi-asserted-by": "crossref",
"first-page": "4575",
"journal-title": "Nat. Commun.",
"key": "10.1016/j.isci.2026.114849_bib38",
"volume": "10",
"year": "2019"
},
{
"DOI": "10.1186/s12950-015-0081-4",
"article-title": "The PPAR-gamma antagonist GW9662 elicits differentiation of M2c-like cells and upregulation of the MerTK/Gas6 axis: a key role for PPAR-gamma in human macrophage polarization",
"author": "Zizzo",
"doi-asserted-by": "crossref",
"first-page": "36",
"journal-title": "J. Inflamm.",
"key": "10.1016/j.isci.2026.114849_bib42",
"volume": "12",
"year": "2015"
},
{
"DOI": "10.1016/j.cell.2021.11.033",
"article-title": "SARS-CoV-2 infection triggers profibrotic macrophage responses and lung fibrosis",
"author": "Wendisch",
"doi-asserted-by": "crossref",
"first-page": "6243",
"journal-title": "Cell",
"key": "10.1016/j.isci.2026.114849_bib43",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.1186/s43556-024-00232-9",
"article-title": "Characteristics of the early innate response induced by the aerosolized Ad5-vectored COVID-19 vaccine",
"author": "Zheng",
"doi-asserted-by": "crossref",
"first-page": "64",
"journal-title": "Mol. Biomed.",
"key": "10.1016/j.isci.2026.114849_bib44",
"volume": "5",
"year": "2024"
},
{
"DOI": "10.1172/JCI162581",
"article-title": "Trained immunity is induced in humans after immunization with an adenoviral vector COVID-19 vaccine",
"author": "Murphy",
"doi-asserted-by": "crossref",
"journal-title": "J. Clin. Investig.",
"key": "10.1016/j.isci.2026.114849_bib45",
"volume": "133",
"year": "2023"
},
{
"DOI": "10.1038/s41591-020-0901-9",
"article-title": "Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19",
"author": "Liao",
"doi-asserted-by": "crossref",
"first-page": "842",
"journal-title": "Nat. Med.",
"key": "10.1016/j.isci.2026.114849_bib37",
"volume": "26",
"year": "2020"
},
{
"DOI": "10.1101/gad.14.2.232",
"article-title": "Antiapoptotic activity of Stat5 required during terminal stages of myeloid differentiation",
"author": "Kieslinger",
"doi-asserted-by": "crossref",
"first-page": "232",
"journal-title": "Genes Dev.",
"key": "10.1016/j.isci.2026.114849_bib46",
"volume": "14",
"year": "2000"
},
{
"DOI": "10.1182/blood-2009-04-216390",
"article-title": "The transcription factors STAT5A/B regulate GM-CSF-mediated granulopoiesis",
"author": "Kimura",
"doi-asserted-by": "crossref",
"first-page": "4721",
"journal-title": "Blood",
"key": "10.1016/j.isci.2026.114849_bib47",
"volume": "114",
"year": "2009"
},
{
"DOI": "10.1089/wound.2013.0466",
"article-title": "The Role of TGFbeta Signaling in Wound Epithelialization",
"author": "Ramirez",
"doi-asserted-by": "crossref",
"first-page": "482",
"journal-title": "Adv. Wound Care",
"key": "10.1016/j.isci.2026.114849_bib48",
"volume": "3",
"year": "2014"
},
{
"DOI": "10.3390/cells10113242",
"article-title": "The Role of Fibroblast Growth Factor (FGF) Signaling in Tissue Repair and Regeneration",
"author": "Farooq",
"doi-asserted-by": "crossref",
"journal-title": "Cells",
"key": "10.1016/j.isci.2026.114849_bib49",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1084/jem.20190103",
"article-title": "Transforming growth factor-beta in tissue fibrosis",
"author": "Frangogiannis",
"doi-asserted-by": "crossref",
"journal-title": "J. Exp. Med.",
"key": "10.1016/j.isci.2026.114849_bib50",
"volume": "217",
"year": "2020"
},
{
"DOI": "10.2174/157340810791233033",
"article-title": "TGF-beta: Titan of Lung Fibrogenesis",
"author": "Yue",
"doi-asserted-by": "crossref",
"first-page": "67",
"journal-title": "Curr. Enzym. Inhib.",
"key": "10.1016/j.isci.2026.114849_bib51",
"volume": "6",
"year": "2010"
},
{
"DOI": "10.1093/intimm/11.7.1075",
"article-title": "STAT1 activation during monocyte to macrophage maturation: role of adhesion molecules",
"author": "Coccia",
"doi-asserted-by": "crossref",
"first-page": "1075",
"journal-title": "Int. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib52",
"volume": "11",
"year": "1999"
},
{
"DOI": "10.1182/blood-2007-03-080291",
"article-title": "C/EBPalpha binds and activates the PU.1 distal enhancer to induce monocyte lineage commitment",
"author": "Yeamans",
"doi-asserted-by": "crossref",
"first-page": "3136",
"journal-title": "Blood",
"key": "10.1016/j.isci.2026.114849_bib53",
"volume": "110",
"year": "2007"
},
{
"DOI": "10.1038/s42003-022-03888-7",
"article-title": "PU.1-c-Jun interaction is crucial for PU.1 function in myeloid development",
"author": "Zhao",
"doi-asserted-by": "crossref",
"first-page": "961",
"journal-title": "Commun. Biol.",
"key": "10.1016/j.isci.2026.114849_bib54",
"volume": "5",
"year": "2022"
},
{
"DOI": "10.1186/s40164-020-00197-9",
"article-title": "AP-1 family transcription factors: a diverse family of proteins that regulate varied cellular activities in classical hodgkin lymphoma and ALK+ ALCL",
"author": "Wu",
"doi-asserted-by": "crossref",
"first-page": "4",
"journal-title": "Exp. Hematol. Oncol.",
"key": "10.1016/j.isci.2026.114849_bib55",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1038/nrm2297",
"article-title": "TGFbeta-SMAD signal transduction: molecular specificity and functional flexibility",
"author": "Schmierer",
"doi-asserted-by": "crossref",
"first-page": "970",
"journal-title": "Nat. Rev. Mol. Cell Biol.",
"key": "10.1016/j.isci.2026.114849_bib56",
"volume": "8",
"year": "2007"
},
{
"DOI": "10.1038/s41592-023-01938-4",
"article-title": "SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks",
"author": "Bravo Gonzalez-Blas",
"doi-asserted-by": "crossref",
"first-page": "1355",
"journal-title": "Nat. Methods",
"key": "10.1016/j.isci.2026.114849_bib57",
"volume": "20",
"year": "2023"
},
{
"DOI": "10.1186/s13075-018-1568-1",
"article-title": "Dysregulated heme oxygenase-1(low) M2-like macrophages augment lupus nephritis via Bach1 induced by type I interferons",
"author": "Kishimoto",
"doi-asserted-by": "crossref",
"first-page": "64",
"journal-title": "Arthritis Res. Ther.",
"key": "10.1016/j.isci.2026.114849_bib58",
"volume": "20",
"year": "2018"
},
{
"DOI": "10.1186/s12929-014-0110-2",
"article-title": "Biological roles of CCAAT/Enhancer-binding protein delta during inflammation",
"author": "Ko",
"doi-asserted-by": "crossref",
"first-page": "6",
"journal-title": "J. Biomed. Sci.",
"key": "10.1016/j.isci.2026.114849_bib59",
"volume": "22",
"year": "2015"
},
{
"DOI": "10.1073/pnas.0506580102",
"article-title": "Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles",
"author": "Subramanian",
"doi-asserted-by": "crossref",
"first-page": "15545",
"journal-title": "Proc. Natl. Acad. Sci. USA",
"key": "10.1016/j.isci.2026.114849_bib60",
"volume": "102",
"year": "2005"
},
{
"DOI": "10.1038/s12276-020-0435-8",
"article-title": "Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development",
"author": "Hwang",
"doi-asserted-by": "crossref",
"first-page": "750",
"journal-title": "Exp. Mol. Med.",
"key": "10.1016/j.isci.2026.114849_bib61",
"volume": "52",
"year": "2020"
},
{
"DOI": "10.1073/pnas.87.10.3723",
"article-title": "Reciprocal expression of human ETS1 and ETS2 genes during T-cell activation: regulatory role for the protooncogene ETS1",
"author": "Bhat",
"doi-asserted-by": "crossref",
"first-page": "3723",
"journal-title": "Proc. Natl. Acad. Sci. USA",
"key": "10.1016/j.isci.2026.114849_bib62",
"volume": "87",
"year": "1990"
},
{
"DOI": "10.1038/s41577-018-0020-8",
"article-title": "Regulatory mechanisms in T cell receptor signalling",
"author": "Gaud",
"doi-asserted-by": "crossref",
"first-page": "485",
"journal-title": "Nat. Rev. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib63",
"volume": "18",
"year": "2018"
},
{
"DOI": "10.1038/nri2476",
"article-title": "GATA3 and the T-cell lineage: essential functions before and after T-helper-2-cell differentiation",
"author": "Ho",
"doi-asserted-by": "crossref",
"first-page": "125",
"journal-title": "Nat. Rev. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib64",
"volume": "9",
"year": "2009"
},
{
"DOI": "10.1016/j.it.2014.04.002",
"article-title": "GATA3: a master of many trades in immune regulation",
"author": "Wan",
"doi-asserted-by": "crossref",
"first-page": "233",
"journal-title": "Trends Immunol.",
"key": "10.1016/j.isci.2026.114849_bib65",
"volume": "35",
"year": "2014"
},
{
"DOI": "10.1158/2643-3230.BCD-20-0026",
"article-title": "Combinatorial ETS1-dependent control of oncogenic NOTCH1 enhancers in T-cell leukemia",
"author": "McCarter",
"doi-asserted-by": "crossref",
"first-page": "178",
"journal-title": "Blood Cancer Discov.",
"key": "10.1016/j.isci.2026.114849_bib66",
"volume": "1",
"year": "2020"
},
{
"DOI": "10.1016/S0014-5793(01)02907-6",
"article-title": "The AP-1 repressor, JDP2, is a bona fide substrate for the c-Jun N-terminal kinase",
"author": "Katz",
"doi-asserted-by": "crossref",
"first-page": "196",
"journal-title": "FEBS Lett.",
"key": "10.1016/j.isci.2026.114849_bib67",
"volume": "506",
"year": "2001"
},
{
"DOI": "10.1084/jem.20021321",
"article-title": "dimerization protein 2 (JDP2), a member of the AP-1 family of transcription factor, mediates osteoclast differentiation induced by RANKL",
"author": "Kawaida",
"doi-asserted-by": "crossref",
"first-page": "1029",
"journal-title": "J. Exp. Med.",
"key": "10.1016/j.isci.2026.114849_bib68",
"volume": "197",
"year": "2003"
},
{
"DOI": "10.1016/j.it.2018.08.006",
"article-title": "Canonical and Non-Canonical Wnt Signaling in Immune Cells",
"author": "Chae",
"doi-asserted-by": "crossref",
"first-page": "830",
"journal-title": "Trends Immunol.",
"key": "10.1016/j.isci.2026.114849_bib69",
"volume": "39",
"year": "2018"
},
{
"DOI": "10.1016/j.cell.2012.05.012",
"article-title": "Wnt/beta-catenin signaling and disease",
"author": "Clevers",
"doi-asserted-by": "crossref",
"first-page": "1192",
"journal-title": "Cell",
"key": "10.1016/j.isci.2026.114849_bib70",
"volume": "149",
"year": "2012"
},
{
"DOI": "10.1038/s44298-024-00018-4",
"article-title": "The Wnt/β-catenin pathway is important for replication of SARS-CoV-2 and other pathogenic RNA viruses",
"author": "Xu",
"doi-asserted-by": "crossref",
"first-page": "6",
"journal-title": "npj Viruses",
"key": "10.1016/j.isci.2026.114849_bib71",
"volume": "2",
"year": "2024"
},
{
"DOI": "10.1183/13993003.01531-2020",
"article-title": "Wnt5a and Wnt11 as acute respiratory distress syndrome biomarkers for severe acute respiratory syndrome coronavirus 2 patients",
"author": "Choi",
"doi-asserted-by": "crossref",
"journal-title": "Eur. Respir. J.",
"key": "10.1016/j.isci.2026.114849_bib72",
"volume": "56",
"year": "2020"
},
{
"DOI": "10.3390/cells11223592",
"article-title": "Macrophages and Wnts in Tissue Injury and Repair",
"author": "Park",
"doi-asserted-by": "crossref",
"journal-title": "Cells",
"key": "10.1016/j.isci.2026.114849_bib73",
"volume": "11",
"year": "2022"
},
{
"DOI": "10.4049/jimmunol.1201819",
"article-title": "Wnt6 is expressed in granulomatous lesions of Mycobacterium tuberculosis-infected mice and is involved in macrophage differentiation and proliferation",
"author": "Schaale",
"doi-asserted-by": "crossref",
"first-page": "5182",
"journal-title": "J. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib74",
"volume": "191",
"year": "2013"
},
{
"DOI": "10.1186/s12964-018-0300-8",
"article-title": "M2 macrophages promote myofibroblast differentiation of LR-MSCs and are associated with pulmonary fibrogenesis",
"author": "Hou",
"doi-asserted-by": "crossref",
"first-page": "89",
"journal-title": "Cell Commun. Signal.",
"key": "10.1016/j.isci.2026.114849_bib75",
"volume": "16",
"year": "2018"
},
{
"DOI": "10.3389/fimmu.2024.1401015",
"article-title": "TGF-beta1 overexpression in severe COVID-19 survivors and its implications for early-phase fibrotic abnormalities and long-term functional impairment",
"author": "Alfaro",
"doi-asserted-by": "crossref",
"journal-title": "Front. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib76",
"volume": "15",
"year": "2024"
},
{
"DOI": "10.3390/biomedicines12030639",
"article-title": "The Role of Cytokines and Molecular Pathways in Lung Fibrosis Following SARS-CoV-2 Infection: A Physiopathologic (Re)view",
"author": "Lazar",
"doi-asserted-by": "crossref",
"journal-title": "Biomedicines",
"key": "10.1016/j.isci.2026.114849_bib77",
"volume": "12",
"year": "2024"
},
{
"article-title": "A distinct innate immune signature marks progression from mild to severe COVID-19",
"author": "Chevrier",
"journal-title": "Cell Rep. Med.",
"key": "10.1016/j.isci.2026.114849_bib78",
"volume": "2",
"year": "2021"
},
{
"DOI": "10.1165/rcmb.2022-0107TR",
"article-title": "Insights from Transcriptomics: CD163(+) Profibrotic Lung Macrophages in COVID-19",
"author": "Bhattacharya",
"doi-asserted-by": "crossref",
"first-page": "520",
"journal-title": "Am. J. Respir. Cell Mol. Biol.",
"key": "10.1016/j.isci.2026.114849_bib79",
"volume": "67",
"year": "2022"
},
{
"DOI": "10.1016/j.bbamcr.2005.06.008",
"article-title": "Depletion of the AP-1 repressor JDP2 induces cell death similar to apoptosis",
"author": "Lerdrup",
"doi-asserted-by": "crossref",
"first-page": "29",
"journal-title": "Biochim. Biophys. Acta",
"key": "10.1016/j.isci.2026.114849_bib80",
"volume": "1745",
"year": "2005"
},
{
"DOI": "10.1128/MCB.17.6.3094",
"article-title": "Isolation of an AP-1 repressor by a novel method for detecting protein-protein interactions",
"author": "Aronheim",
"doi-asserted-by": "crossref",
"first-page": "3094",
"journal-title": "Mol. Cell Biol.",
"key": "10.1016/j.isci.2026.114849_bib81",
"volume": "17",
"year": "1997"
},
{
"DOI": "10.1016/j.clim.2023.109634",
"article-title": "Unique molecular signatures sustained in circulating monocytes and regulatory T cells in convalescent COVID-19 patients",
"author": "Hoffmann",
"doi-asserted-by": "crossref",
"journal-title": "Clin. Immunol.",
"key": "10.1016/j.isci.2026.114849_bib82",
"volume": "252",
"year": "2023"
},
{
"DOI": "10.1038/ncomms14049",
"article-title": "Massively parallel digital transcriptional profiling of single cells",
"author": "Zheng",
"doi-asserted-by": "crossref",
"journal-title": "Nat. Commun.",
"key": "10.1016/j.isci.2026.114849_bib83",
"volume": "8",
"year": "2017"
},
{
"DOI": "10.1038/s41587-023-01767-y",
"article-title": "Dictionary learning for integrative, multimodal and scalable single-cell analysis",
"author": "Hao",
"doi-asserted-by": "crossref",
"first-page": "293",
"journal-title": "Nat. Biotechnol.",
"key": "10.1016/j.isci.2026.114849_bib84",
"volume": "42",
"year": "2024"
},
{
"DOI": "10.1038/s41592-021-01282-5",
"article-title": "Single-cell chromatin state analysis with Signac",
"author": "Stuart",
"doi-asserted-by": "crossref",
"first-page": "1333",
"journal-title": "Nat. Methods",
"key": "10.1016/j.isci.2026.114849_bib85",
"volume": "18",
"year": "2021"
},
{
"DOI": "10.1681/ASN.0000000000000354",
"article-title": "Cell Type- and Age-Specific Expression of lncRNAs across Kidney Cell Types",
"author": "Kim",
"doi-asserted-by": "crossref",
"first-page": "870",
"journal-title": "J. Am. Soc. Nephrol.",
"key": "10.1016/j.isci.2026.114849_bib86",
"volume": "35",
"year": "2024"
},
{
"DOI": "10.1016/j.mocell.2024.100103",
"article-title": "A practical handbook on single-cell RNA sequencing data quality control and downstream analysis",
"author": "Kim",
"doi-asserted-by": "crossref",
"journal-title": "Mol. Cells",
"key": "10.1016/j.isci.2026.114849_bib87",
"volume": "47",
"year": "2024"
},
{
"DOI": "10.1038/nbt.4042",
"article-title": "Multiplexed droplet single-cell RNA-sequencing using natural genetic variation",
"author": "Kang",
"doi-asserted-by": "crossref",
"first-page": "89",
"journal-title": "Nat. Biotechnol.",
"key": "10.1016/j.isci.2026.114849_bib88",
"volume": "36",
"year": "2018"
},
{
"DOI": "10.1093/bioinformatics/btr509",
"article-title": "A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data",
"author": "Li",
"doi-asserted-by": "crossref",
"first-page": "2987",
"journal-title": "Bioinformatics",
"key": "10.1016/j.isci.2026.114849_bib89",
"volume": "27",
"year": "2011"
},
{
"DOI": "10.1016/j.cels.2019.03.003",
"article-title": "DoubletFinder: Doublet Detection in Single-Cell RNA Sequencing Data Using Artificial Nearest Neighbors",
"author": "McGinnis",
"doi-asserted-by": "crossref",
"first-page": "329",
"journal-title": "Cell Syst.",
"key": "10.1016/j.isci.2026.114849_bib90",
"volume": "8",
"year": "2019"
},
{
"DOI": "10.12688/f1000research.73600.1",
"article-title": "Doublet identification in single-cell sequencing data using scDblFinder",
"author": "Germain",
"doi-asserted-by": "crossref",
"first-page": "979",
"journal-title": "F1000Res.",
"key": "10.1016/j.isci.2026.114849_bib91",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1038/s41592-019-0619-0",
"article-title": "Fast, sensitive and accurate integration of single-cell data with Harmony",
"author": "Korsunsky",
"doi-asserted-by": "crossref",
"first-page": "1289",
"journal-title": "Nat. Methods",
"key": "10.1016/j.isci.2026.114849_bib92",
"volume": "16",
"year": "2019"
},
{
"DOI": "10.1186/gb-2008-9-9-r137",
"article-title": "Model-based analysis of ChIP-Seq (MACS)",
"author": "Zhang",
"doi-asserted-by": "crossref",
"journal-title": "Genome Biol.",
"key": "10.1016/j.isci.2026.114849_bib93",
"volume": "9",
"year": "2008"
},
{
"DOI": "10.1038/75556",
"article-title": "Gene ontology: tool for the unification of biology. The Gene Ontology Consortium",
"author": "Ashburner",
"doi-asserted-by": "crossref",
"first-page": "25",
"journal-title": "Nat. Genet.",
"key": "10.1016/j.isci.2026.114849_bib94",
"volume": "25",
"year": "2000"
},
{
"DOI": "10.1038/s41596-024-01137-1",
"article-title": "CellPhoneDB v5: inferring cell-cell communication from single-cell multiomics data",
"author": "Troule",
"doi-asserted-by": "crossref",
"first-page": "3412",
"journal-title": "Nat. Protoc.",
"key": "10.1016/j.isci.2026.114849_bib95",
"volume": "20",
"year": "2025"
},
{
"DOI": "10.1186/s12859-023-05342-9",
"article-title": "Empowering biologists to decode omics data: the Genekitr R package and web server",
"author": "Liu",
"doi-asserted-by": "crossref",
"first-page": "214",
"journal-title": "BMC Bioinf.",
"key": "10.1016/j.isci.2026.114849_bib96",
"volume": "24",
"year": "2023"
},
{
"DOI": "10.1093/bioinformatics/btv145",
"article-title": "ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization",
"author": "Yu",
"doi-asserted-by": "crossref",
"first-page": "2382",
"journal-title": "Bioinformatics",
"key": "10.1016/j.isci.2026.114849_bib97",
"volume": "31",
"year": "2015"
},
{
"DOI": "10.1038/nmeth.4401",
"article-title": "chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data",
"author": "Schep",
"doi-asserted-by": "crossref",
"first-page": "975",
"journal-title": "Nat. Methods",
"key": "10.1016/j.isci.2026.114849_bib98",
"volume": "14",
"year": "2017"
}
],
"reference-count": 97,
"references-count": 97,
"relation": {},
"resource": {
"primary": {
"URL": "https://linkinghub.elsevier.com/retrieve/pii/S2589004226002245"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Discovery of key regulators in classical monocyte phenotypes linked to COVID-19 severity using single-cell multi-omics sequencing",
"type": "journal-article",
"update-policy": "https://doi.org/10.1016/elsevier_cm_policy",
"volume": "29"
}