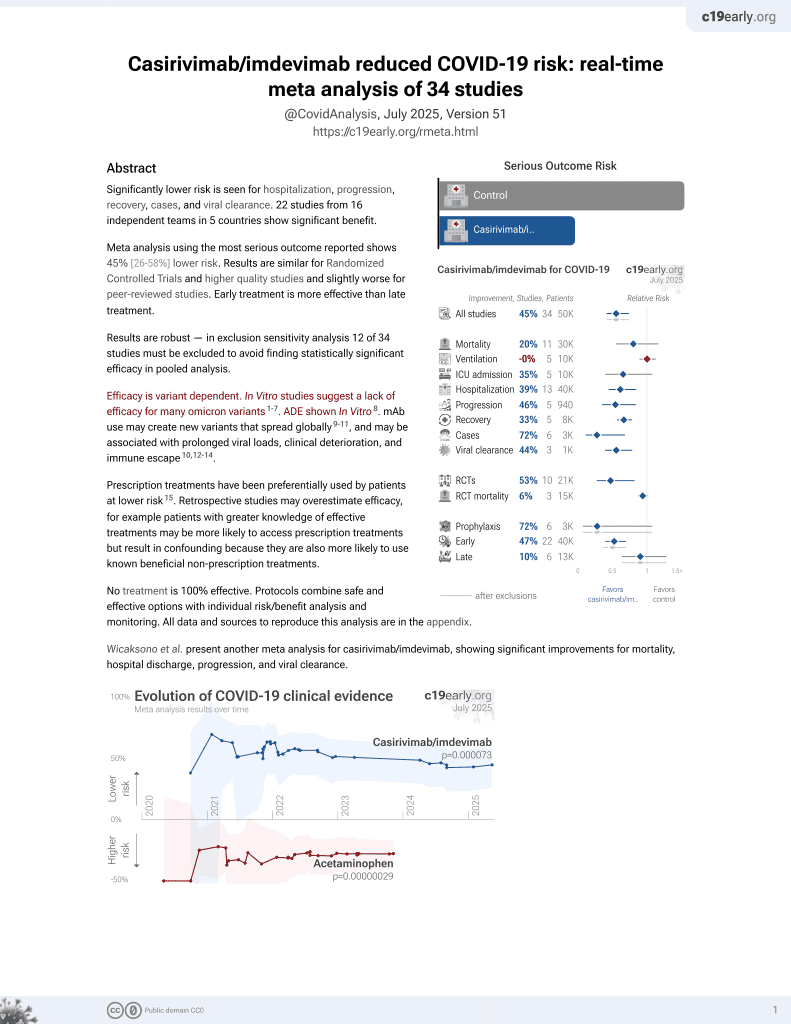
Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies
et al., Science, 21 Aug 2020, 369:6506, 1014-1018, doi:10.1126/science.abd0831, Aug 2020
18th treatment shown to reduce risk in
March 2021, now with p = 0.000095 from 34 studies, recognized in 52 countries.
Efficacy is variant dependent.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
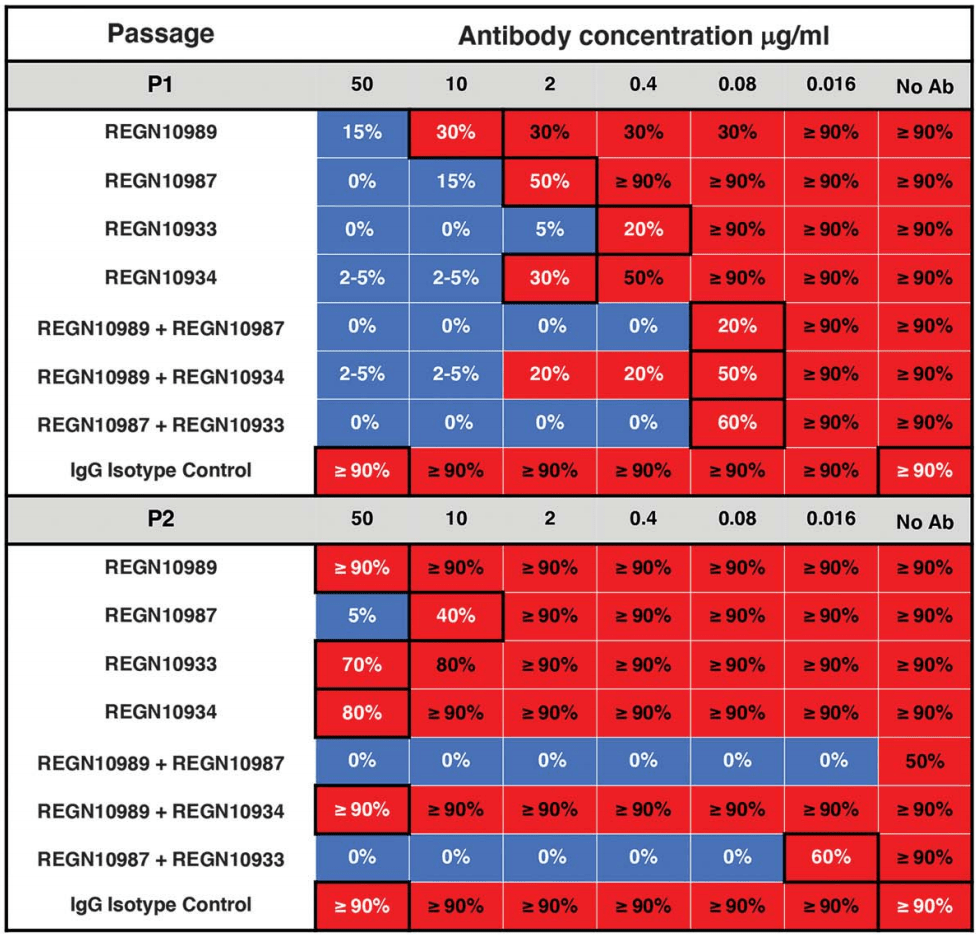
In vitro study showing that, under pressure from individual antibodies, mutant viruses were rapidly selected that evaded the blocking function of all individual antibodies tested, including antibodies that potently bind to highly-conserved regions on the spike protein. However, escape mutants could not be efficiently generated following exposure to the REGN-COV2 cocktail since it utilizes two antibodies that can simultaneously bind to distinct regions of the RBD.
Efficacy is variant dependent. In Vitro research suggests a lack of efficacy for many omicron variants1-7.
1.
Liu et al., Striking Antibody Evasion Manifested by the Omicron Variant of SARS-CoV-2, bioRxiv, doi:10.1101/2021.12.14.472719.
2.
Sheward et al., Variable loss of antibody potency against SARS-CoV-2 B.1.1.529 (Omicron), bioRxiv, doi:10.1101/2021.12.19.473354.
3.
VanBlargan et al., An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by several therapeutic monoclonal antibodies, bioRxiv, doi:10.1101/2021.12.15.472828.
4.
Tatham et al., Lack of Ronapreve (REGN-CoV; casirivimab and imdevimab) virological efficacy against the SARS-CoV 2 Omicron variant (B.1.1.529) in K18-hACE2 mice, bioRxiv, doi:10.1101/2022.01.23.477397.
5.
Pochtovyi et al., In Vitro Efficacy of Antivirals and Monoclonal Antibodies against SARS-CoV-2 Omicron Lineages XBB.1.9.1, XBB.1.9.3, XBB.1.5, XBB.1.16, XBB.2.4, BQ.1.1.45, CH.1.1, and CL.1, Vaccines, doi:10.3390/vaccines11101533.
Baum et al., 21 Aug 2020, peer-reviewed, 17 authors.
In vitro studies are an important part of preclinical research, however results may be very different in vivo.
Abstract: RES EARCH
CORONAVIRUS
Antibody cocktail to SARS-CoV-2 spike protein
prevents rapid mutational escape seen with
individual antibodies
Alina Baum, Benjamin O. Fulton, Elzbieta Wloga, Richard Copin, Kristen E. Pascal, Vincenzo Russo,
Stephanie Giordano, Kathryn Lanza, Nicole Negron, Min Ni, Yi Wei, Gurinder S. Atwal,
Andrew J. Murphy, Neil Stahl, George D. Yancopoulos, Christos A. Kyratsous*
Antibodies targeting the spike protein of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)
present a promising approach to combat the coronavirus disease 2019 (COVID-19) pandemic; however,
concerns remain that mutations can yield antibody resistance. We investigated the development of
resistance against four antibodies to the spike protein that potently neutralize SARS-CoV-2, individually as
well as when combined into cocktails. These antibodies remain effective against spike variants that have
arisen in the human population. However, novel spike mutants rapidly appeared after in vitro passaging
in the presence of individual antibodies, resulting in loss of neutralization; such escape also occurred
with combinations of antibodies binding diverse but overlapping regions of the spike protein. Escape
mutants were not generated after treatment with a noncompeting antibody cocktail.
O
Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA.
*Corresponding author. Email: christos.kyratsous@regeneron.com
Baum et al., Science 369, 1014–1018 (2020)
in response to selective pressure from singleantibody treatments.
To assess the efficacy of our recently described antiviral antibodies against the breadth
of spike RBD variants represented in publicly
available SARS-CoV-2 sequences identified
through the end of March 2020 (representing
more than 7000 unique genomes), we used
the VSV pseudoparticle system expressing the
SARS-CoV-2 spike variants. Our top eight
neutralizing antibodies maintained their potency against all tested variants (Table 1),
demonstrating broad coverage against circulating SARS-CoV-2.
Next, escape mutants were selected under
pressure of single antibodies, as well as of
antibody combinations, by using a replicating
VSV-SARS-CoV-2-S virus (Fig. 1A). We rapidly
identified multiple independent escape mutants for each of the four individual antibodies
within the first passage (Fig. 1, B and C, and
Fig. 2). Some of these mutants became readily
fixed in the population by the second passage,
representing 100% of sequencing reads, and
are resistant to antibody concentrations of up to
50 mg/ml [a factor of ~10,000 to 100,000 greater
concentration than half-maximal inhibitory
concentration (IC50) against parental virus].
Sequencing of escape mutants (Fig. 2) revealed
that single amino acid changes can ablate
binding even to antibodies that were selected
for breadth against all known RBD variants
(Table 1) and that neutralize parental virus
at IC50 values in the low picomolar range (3).
Analysis of 22,872 publicly available unique
genome sequences (through the end of May
2020) demonstrated the presence of polymorphisms analogous to two of the escape
amino acid residues identified in our study,
albeit at an extremely low frequency of one
each. Thus, although natural variants resist-
21 August 2020
1 of 4
ne promising approach to combat the
coronavirus disease 2019 (COVID-19)
pandemic involves development of antiviral antibodies targeting the spike protein of severe acute..
DOI record:
{
"DOI": "10.1126/science.abd0831",
"ISSN": [
"0036-8075",
"1095-9203"
],
"URL": "http://dx.doi.org/10.1126/science.abd0831",
"abstract": "<jats:title>An antibody cocktail against SARS-CoV-2</jats:title>\n <jats:p>\n There is an urgent focus on antibodies that target the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) viral spike and prevent the virus from entering host cells. Hansen\n <jats:italic>et al.</jats:italic>\n generated a large panel of antibodies against the spike protein from humanized mice and recovered patients. From this panel, they identified several neutralizing antibodies, including pairs that do not compete for binding to the receptor binding domain. Baum\n <jats:italic>et al.</jats:italic>\n focused in on four of these antibodies. All four are effective against known spike variants. However, by growing a pseudovirus that expresses the spike in the presence of individual antibodies, the authors were able to select for spike mutants resistant to that antibody. In contrast, escape mutants are not selected when pseudovirus is grown in the presence of pairs of antibodies that either do not compete or only partially compete for binding to the RBD. Such a pair might be used in a therapeutic antibody cocktail.\n </jats:p>\n <jats:p>\n <jats:italic>Science</jats:italic>\n , this issue p.\n <jats:related-article xmlns:xlink=\"http://www.w3.org/1999/xlink\" ext-link-type=\"doi\" issue=\"6506\" page=\"1010\" related-article-type=\"in-this-issue\" vol=\"369\" xlink:href=\"10.1126/science.abd0827\">1010</jats:related-article>\n , p.\n <jats:related-article xmlns:xlink=\"http://www.w3.org/1999/xlink\" ext-link-type=\"doi\" issue=\"6506\" page=\"1014\" related-article-type=\"in-this-issue\" vol=\"369\" xlink:href=\"10.1126/science.abd0831\">1014</jats:related-article>\n </jats:p>",
"alternative-id": [
"10.1126/science.abd0831"
],
"author": [
{
"ORCID": "http://orcid.org/0000-0001-7179-8679",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Baum",
"given": "Alina",
"sequence": "first"
},
{
"ORCID": "http://orcid.org/0000-0003-1224-4988",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Fulton",
"given": "Benjamin O.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-1494-9765",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Wloga",
"given": "Elzbieta",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Copin",
"given": "Richard",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0003-4826-4606",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Pascal",
"given": "Kristen E.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-3295-7654",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Russo",
"given": "Vincenzo",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0001-5879-3275",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Giordano",
"given": "Stephanie",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Lanza",
"given": "Kathryn",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Negron",
"given": "Nicole",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Ni",
"given": "Min",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Wei",
"given": "Yi",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Atwal",
"given": "Gurinder S.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0003-4152-4081",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Murphy",
"given": "Andrew J.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0001-7265-2575",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Stahl",
"given": "Neil",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"family": "Yancopoulos",
"given": "George D.",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-2596-2906",
"affiliation": [
{
"name": "Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA."
}
],
"authenticated-orcid": true,
"family": "Kyratsous",
"given": "Christos A.",
"sequence": "additional"
}
],
"container-title": "Science",
"container-title-short": "Science",
"content-domain": {
"crossmark-restriction": false,
"domain": []
},
"created": {
"date-parts": [
[
2020,
6,
15
]
],
"date-time": "2020-06-15T19:12:08Z",
"timestamp": 1592248328000
},
"deposited": {
"date-parts": [
[
2024,
1,
15
]
],
"date-time": "2024-01-15T18:35:26Z",
"timestamp": 1705343726000
},
"funder": [
{
"DOI": "10.13039/100012399",
"award": [
"HHSO100201700020C"
],
"doi-asserted-by": "publisher",
"name": "Biomedical Advanced Research and Development Authority"
}
],
"indexed": {
"date-parts": [
[
2024,
5,
14
]
],
"date-time": "2024-05-14T17:54:23Z",
"timestamp": 1715709263876
},
"is-referenced-by-count": 1159,
"issue": "6506",
"issued": {
"date-parts": [
[
2020,
8,
21
]
]
},
"journal-issue": {
"issue": "6506",
"published-print": {
"date-parts": [
[
2020,
8,
21
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2020,
8,
21
]
],
"date-time": "2020-08-21T00:00:00Z",
"timestamp": 1597968000000
}
}
],
"link": [
{
"URL": "https://syndication.highwire.org/content/doi/10.1126/science.abd0831",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "syndication"
},
{
"URL": "https://www.science.org/doi/pdf/10.1126/science.abd0831",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "221",
"original-title": [],
"page": "1014-1018",
"prefix": "10.1126",
"published": {
"date-parts": [
[
2020,
8,
21
]
]
},
"published-print": {
"date-parts": [
[
2020,
8,
21
]
]
},
"publisher": "American Association for the Advancement of Science (AAAS)",
"reference": [
{
"DOI": "10.1126/science.abb2762",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_2_2"
},
{
"DOI": "10.1016/j.cell.2020.03.045",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_3_2"
},
{
"DOI": "10.1126/science.abd0827",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_4_2"
},
{
"DOI": "10.1080/22221751.2020.1756700",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_5_2"
},
{
"DOI": "10.1101/2020.05.20.105247",
"doi-asserted-by": "crossref",
"key": "e_1_3_2_6_2",
"unstructured": "M. E. Dieterle D. Haslwanter R. H. Bortz 3rd A. S. Wirchnianski G. Lasso O. Vergnolle S. A. Abbasi J. M. Fels E. Laudermilch C. Florez A. Mengotto D. Kimmel R. J. Malonis G. Georgiev J. Quiroz J. Barnhill L. A. Pirofski J. P. Daily J. M. Dye J. R. Lai A. S. Herbert K. Chandran R. K. Jangra A replication-competent vesicular stomatitis virus for studies of SARS-CoV-2 spike-mediated cell entry and its inhibition. bioRxiv (2020).32511365"
},
{
"DOI": "10.1038/s41586-020-2349-y",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_7_2"
},
{
"DOI": "10.1016/j.chom.2014.04.009",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_8_2"
},
{
"DOI": "10.1128/JVI.01639-18",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_9_2"
},
{
"DOI": "10.1101/2020.05.22.111005",
"doi-asserted-by": "crossref",
"key": "e_1_3_2_10_2",
"unstructured": "S. J. Zost P. Gilchuk J. B. Case E. Binshtein R. E. Chen J. X. Reidy A. Trivette R. S. Nargi R. E. Sutton N. Suryadevara L. E. Williamson E. C. Chen T. Jones S. Day L. Myers A. O. Hassan N. M. Kafai E. S. Winkler J. M. Fox J. J. Steinhardt K. Ren Y. M. Loo N. L. Kallewaard D. R. Martinez A. Schäfer L. E. Gralinski R. S. Baric L. B. Thackray M. S. Diamond R. H. Carnahan J. E. Crowe Potently neutralizing human antibodies that block SARS-CoV-2 receptor binding and protect animals. bioRxiv (2020).32511409"
},
{
"DOI": "10.1038/s41586-020-2456-9",
"doi-asserted-by": "crossref",
"key": "e_1_3_2_11_2",
"unstructured": "D. F. Robbiani C. Gaebler F. Muecksch J. C. C. Lorenzi Z. Wang A. Cho M. Agudelo C. O. Barnes A. Gazumyan S. Finkin T. Hagglof T. Y. Oliveira C. Viant A. Hurley H. H. Hoffmann K. G. Millard R. G. Kost M. Cipolla K. Gordon F. Bianchini S. T. Chen V. Ramos R. Patel J. Dizon I. Shimeliovich P. Mendoza H. Hartweger L. Nogueira M. Pack J. Horowitz F. Schmidt Y. Weisblum E. Michailidis A. W. Ashbrook E. Waltari J. E. Pak K. E. Huey-Tubman N. Koranda P. R. Hoffman A. P. West Jr.. C. M. Rice T. Hatziioannou P. J. Bjorkman P. D. Bieniasz M. Caskey M. C. Nussenzweig Convergent Antibody Responses to SARS-CoV-2 Infection in Convalescent Individuals. bioRxiv (2020).32511384"
},
{
"DOI": "10.1016/j.cell.2020.05.025",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_12_2"
},
{
"DOI": "10.1073/pnas.1510830112",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_13_2"
},
{
"DOI": "10.1038/nmeth.2413",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_14_2"
},
{
"DOI": "10.1073/pnas.92.10.4477",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_15_2"
},
{
"DOI": "10.1128/JVI.69.5.2946-2953.1995",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_16_2"
},
{
"DOI": "10.1016/j.jviromet.2010.08.006",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_17_2"
},
{
"DOI": "10.1073/pnas.94.26.14764",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_18_2"
},
{
"DOI": "10.1099/vir.0.80955-0",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_19_2"
},
{
"DOI": "10.1080/22221751.2020.1743767",
"doi-asserted-by": "publisher",
"key": "e_1_3_2_20_2"
}
],
"reference-count": 19,
"references-count": 19,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.science.org/doi/10.1126/science.abd0831"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies",
"type": "journal-article",
"volume": "369"
}