Cellular host factors for SARS-CoV-2 infection
et al., Nature Microbiology, doi:10.1038/s41564-021-00958-0, Sep 2021
Review of cellular host factors required for SARS-CoV-2 infection. Authors systematically examine proviral host factors that SARS-CoV-2 depends on for successful replication, identified through genome-wide CRISPR screens and viral interactome analyses.
Baggen et al., 1 Sep 2021, multiple countries, peer-reviewed, 4 authors.
Contact: dirk.daelemans@kuleuven.be.
Abstract: Review Article
https://doi.org/10.1038/s41564-021-00958-0
Cellular host factors for SARS-CoV-2 infection
Jim Baggen , Els Vanstreels , Sander Jansen
and Dirk Daelemans
✉
The coronavirus disease 2019 (COVID-19) pandemic has claimed millions of lives and caused a global economic crisis. No effective antiviral drugs are currently available to treat infections of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).
The medical need imposed by the pandemic has spurred unprecedented research efforts to study coronavirus biology. Every
virus depends on cellular host factors and pathways for successful replication. These proviral host factors represent attractive
targets for antiviral therapy as they are genetically more stable than viral targets and may be shared among related viruses.
The application of various ‘omics’ technologies has led to the rapid discovery of proviral host factors that are required for the
completion of the SARS-CoV-2 life cycle. In this Review, we summarize insights into the proviral host factors that are required
for SARS-CoV-2 infection that were mainly obtained using functional genetic and interactome screens. We discuss cellular processes that are important for the SARS-CoV-2 life cycle, as well as parallels with non-coronaviruses. Finally, we highlight host
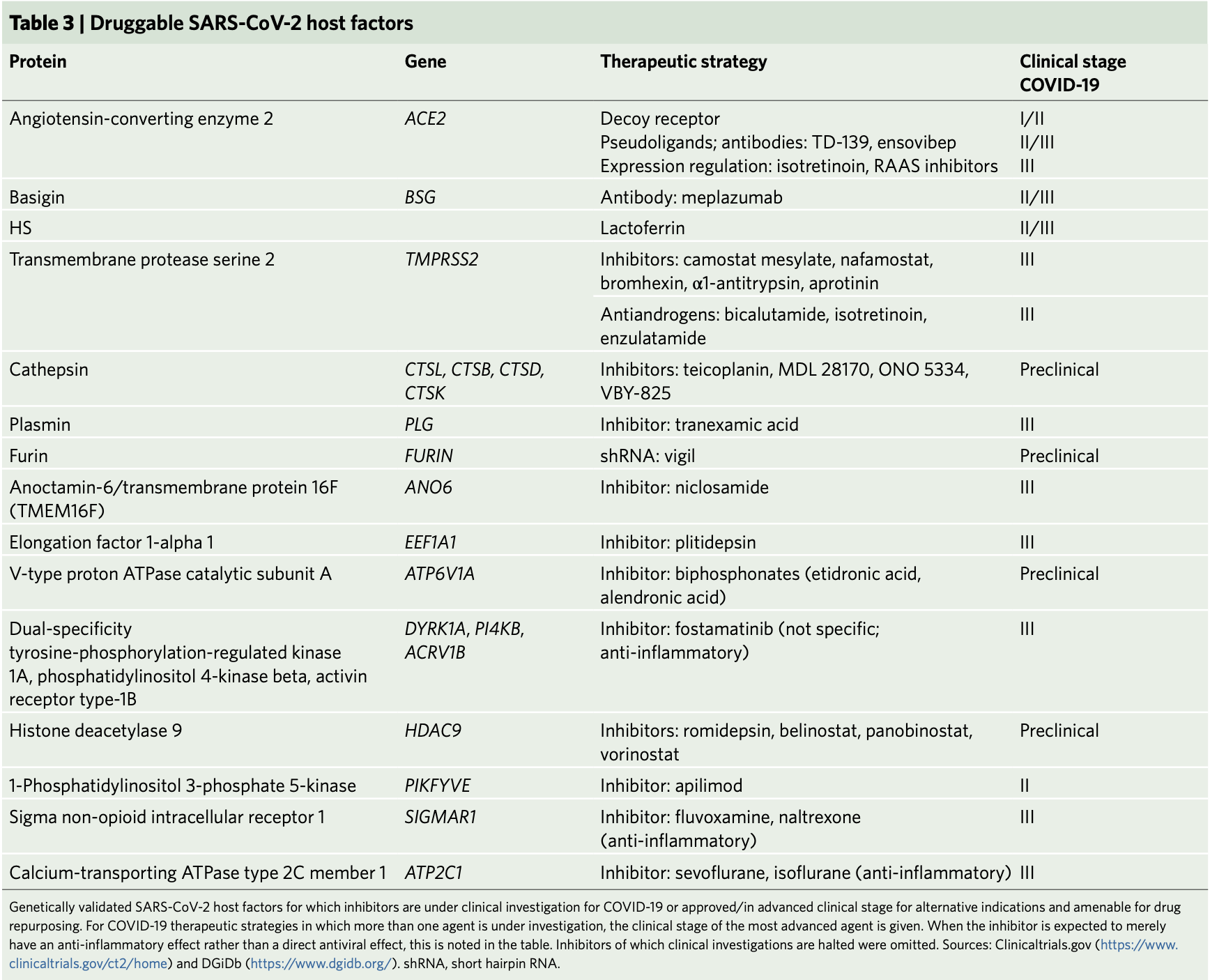
factors that could be targeted by clinically approved molecules and molecules in clinical trials as potential antiviral therapies
for COVID-19.
C
oronaviruses are positive-strand RNA viruses belonging to the subfamily Orthocoronavirinae within the family
Coronaviridae (International Committee on Taxonomy of
Viruses) and are subdivided into four genera—Alphacoronavirus, Betacoronavirus, Gammacoronavirus and Deltacoronavirus.
Coronaviruses cause intestinal and respiratory infections in a variety
of birds and mammals, including livestock and domestic animals.
Seven human coronaviruses (HCoVs) have been characterized, four of which cause mild respiratory infections (HCoV-229E,
HCoV-NL63, HCoV-OC43 and HCoV-HKU1). The emergence
of two highly pathogenic betacoronaviruses in 2002 (severe acute
respiratory syndrome coronavirus (SARS-CoV)) and 2012 (Middle
East respiratory syndrome coronavirus (MERS-CoV)) revealed that
new pathogenic coronaviruses can emerge in the human population by zoonotic transmission (reviewed in ref. 1). In late 2019, a
pathogenic coronavirus SARS-CoV-2 was first detected in Wuhan,
China, causing an outbreak of severe pneumonia. This human
pathogen is thought to have originated in horseshoe bats and was
probably transmitted to humans through an intermediate host that
remains to be identified2. Owing to its high contagiousness and the
occurrence of asymptomatic carriers, SARS-CoV-2 rapidly spread
across the globe and continues to claim human lives and obstruct
social and economic activity, while vaccination programs are ongoing. The pandemic has prompted countless efforts to develop vaccines and antiviral therapies, but also many fundamental studies
to better understand coronavirus biology. As cellular proviral host
factors are potential antiviral drug targets, numerous studies have
analysed host factor dependencies of coronaviruses, in particular
SARS-CoV-2.
In this Review, we provide an overview of proviral host factors
for SARS-CoV-2. After explaining the coronavirus life cycle, we
discuss the cellular receptors and proteases that are required for
SARS-CoV-2 entry. As only few of the identified proviral host factors have been..
DOI record:
{
"DOI": "10.1038/s41564-021-00958-0",
"ISSN": [
"2058-5276"
],
"URL": "http://dx.doi.org/10.1038/s41564-021-00958-0",
"alternative-id": [
"958"
],
"assertion": [
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Received",
"name": "received",
"order": 1,
"value": "10 May 2021"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "3 August 2021"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "First Online",
"name": "first_online",
"order": 3,
"value": "1 September 2021"
},
{
"group": {
"label": "Competing interests",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 1,
"value": "The authors declare no competing interests."
},
{
"label": "Free to read",
"name": "free",
"value": "This content has been made available to all."
}
],
"author": [
{
"ORCID": "https://orcid.org/0000-0002-1628-6766",
"affiliation": [],
"authenticated-orcid": false,
"family": "Baggen",
"given": "Jim",
"sequence": "first"
},
{
"ORCID": "https://orcid.org/0000-0002-7814-8035",
"affiliation": [],
"authenticated-orcid": false,
"family": "Vanstreels",
"given": "Els",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-9344-131X",
"affiliation": [],
"authenticated-orcid": false,
"family": "Jansen",
"given": "Sander",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0001-7092-1153",
"affiliation": [],
"authenticated-orcid": false,
"family": "Daelemans",
"given": "Dirk",
"sequence": "additional"
}
],
"container-title": "Nature Microbiology",
"container-title-short": "Nat Microbiol",
"content-domain": {
"crossmark-restriction": false,
"domain": [
"link.springer.com"
]
},
"created": {
"date-parts": [
[
2021,
9,
1
]
],
"date-time": "2021-09-01T10:04:51Z",
"timestamp": 1630490691000
},
"deposited": {
"date-parts": [
[
2023,
2,
6
]
],
"date-time": "2023-02-06T03:07:36Z",
"timestamp": 1675652856000
},
"indexed": {
"date-parts": [
[
2025,
9,
13
]
],
"date-time": "2025-09-13T15:33:52Z",
"timestamp": 1757777632350,
"version": "3.37.3"
},
"is-referenced-by-count": 193,
"issue": "10",
"issued": {
"date-parts": [
[
2021,
9,
1
]
]
},
"journal-issue": {
"issue": "10",
"published-online": {
"date-parts": [
[
2021,
10
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://www.springernature.com/gp/researchers/text-and-data-mining",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2021,
9,
1
]
],
"date-time": "2021-09-01T00:00:00Z",
"timestamp": 1630454400000
}
},
{
"URL": "https://www.springernature.com/gp/researchers/text-and-data-mining",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2021,
9,
1
]
],
"date-time": "2021-09-01T00:00:00Z",
"timestamp": 1630454400000
}
}
],
"link": [
{
"URL": "https://www.nature.com/articles/s41564-021-00958-0.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.nature.com/articles/s41564-021-00958-0",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.nature.com/articles/s41564-021-00958-0.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "297",
"original-title": [],
"page": "1219-1232",
"prefix": "10.1038",
"published": {
"date-parts": [
[
2021,
9,
1
]
]
},
"published-online": {
"date-parts": [
[
2021,
9,
1
]
]
},
"publisher": "Springer Science and Business Media LLC",
"reference": [
{
"DOI": "10.1038/s41579-018-0118-9",
"author": "J Cui",
"doi-asserted-by": "publisher",
"first-page": "181",
"journal-title": "Nat. Rev. Microbiol.",
"key": "958_CR1",
"unstructured": "Cui, J., Li, F. & Shi, Z. L. Origin and evolution of pathogenic coronaviruses. Nat. Rev. Microbiol. 17, 181–192 (2019).",
"volume": "17",
"year": "2019"
},
{
"DOI": "10.1038/s41467-021-21240-1",
"author": "S Wacharapluesadee",
"doi-asserted-by": "publisher",
"first-page": "972",
"journal-title": "Nat. Commun.",
"key": "958_CR2",
"unstructured": "Wacharapluesadee, S. et al. Evidence for SARS-CoV-2 related coronaviruses circulating in bats and pangolins in Southeast Asia. Nat. Commun. 12, 972 (2021).",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2020.09.018",
"author": "H Yao",
"doi-asserted-by": "publisher",
"first-page": "730",
"journal-title": "Cell",
"key": "958_CR3",
"unstructured": "Yao, H. et al. Molecular architecture of the SARS-CoV-2 virus. Cell 183, 730–738 (2020).",
"volume": "183",
"year": "2020"
},
{
"DOI": "10.1128/JVI.00645-06",
"author": "BW Neuman",
"doi-asserted-by": "publisher",
"first-page": "7918",
"journal-title": "J. Virol.",
"key": "958_CR4",
"unstructured": "Neuman, B. W. et al. Supramolecular architecture of severe acute respiratory syndrome coronavirus revealed by electron cryomicroscopy. J. Virol. 80, 7918–7928 (2006).",
"volume": "80",
"year": "2006"
},
{
"DOI": "10.1146/annurev-virology-110615-042301",
"author": "F Li",
"doi-asserted-by": "publisher",
"first-page": "237",
"journal-title": "Annu. Rev. Virol.",
"key": "958_CR5",
"unstructured": "Li, F. Structure, function, and evolution of coronavirus spike proteins. Annu. Rev. Virol. 3, 237–261 (2016).",
"volume": "3",
"year": "2016"
},
{
"DOI": "10.1016/j.antiviral.2020.104792",
"author": "T Tang",
"doi-asserted-by": "publisher",
"first-page": "104792",
"journal-title": "Antivir. Res.",
"key": "958_CR6",
"unstructured": "Tang, T., Bidon, M., Jaimes, J. A., Whittaker, G. R. & Daniel, S. Coronavirus membrane fusion mechanism offers a potential target for antiviral development. Antivir. Res. 178, 104792 (2020).",
"volume": "178",
"year": "2020"
},
{
"DOI": "10.1016/j.coviro.2021.02.006",
"author": "G Whittaker",
"doi-asserted-by": "publisher",
"first-page": "113",
"journal-title": "Curr. Opin. Virol.",
"key": "958_CR7",
"unstructured": "Whittaker, G., Daniel, S. & Millet, J. Coronavirus entry: how we arrived at SARS-CoV-2. Curr. Opin. Virol. 47, 113–120 (2021).",
"volume": "47",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2020.04.011",
"author": "D Kim",
"doi-asserted-by": "publisher",
"first-page": "914",
"journal-title": "Cell",
"key": "958_CR8",
"unstructured": "Kim, D. et al. The architecture of SARS-CoV-2 transcriptome. Cell 181, 914–921 (2020).",
"volume": "181",
"year": "2020"
},
{
"DOI": "10.1080/22221751.2020.1719902",
"author": "JFW Chan",
"doi-asserted-by": "publisher",
"first-page": "221",
"journal-title": "Emerg. Microbes Infect.",
"key": "958_CR9",
"unstructured": "Chan, J. F. W. et al. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 9, 221–236 (2020).",
"volume": "9",
"year": "2020"
},
{
"DOI": "10.1016/j.antiviral.2014.06.013",
"author": "DX Liu",
"doi-asserted-by": "publisher",
"first-page": "97",
"journal-title": "Antivir. Res.",
"key": "958_CR10",
"unstructured": "Liu, D. X., Fung, T. S., Chong, K. K. L., Shukla, A. & Hilgenfeld, R. Accessory proteins of SARS-CoV and other coronaviruses. Antivir. Res. 109, 97–109 (2014).",
"volume": "109",
"year": "2014"
},
{
"DOI": "10.1038/s41579-020-00468-6",
"doi-asserted-by": "publisher",
"key": "958_CR11",
"unstructured": "V’kovski, P., Kratzel, A., Steiner, S., Stalder, H. & Thiel, V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-020-00468-6 (2020)."
},
{
"DOI": "10.1007/978-1-4939-2438-7_1",
"doi-asserted-by": "crossref",
"key": "958_CR12",
"unstructured": "Fehr, A. R. & Perlman, S. in Coronaviruses. Methods in Molecular Biology Vol. 1282 (eds. Maier, H. et al.) 1–23 (HumanaPress, 2015)."
},
{
"DOI": "10.1128/JVI.01358-06",
"author": "SG Sawicki",
"doi-asserted-by": "publisher",
"first-page": "20",
"journal-title": "J. Virol.",
"key": "958_CR13",
"unstructured": "Sawicki, S. G., Sawicki, D. L. & Siddell, S. G. A contemporary view of coronavirus transcription. J. Virol. 81, 20–29 (2007).",
"volume": "81",
"year": "2007"
},
{
"DOI": "10.1007/82_2017_25",
"doi-asserted-by": "crossref",
"key": "958_CR14",
"unstructured": "de Wilde, A. H., Snijder, E. J., Kikkert, M. & van Hemert, M. J. in Roles of Host Gene and Non-coding RNA Expression in Virus Infection (eds Tripp, R. & Tompkins, S.) 1–42 (Springer, Cham, 2018)."
},
{
"DOI": "10.1371/journal.pbio.3000715",
"author": "EJ Snijder",
"doi-asserted-by": "publisher",
"first-page": "e3000715",
"journal-title": "PLoS Biol.",
"key": "958_CR15",
"unstructured": "Snijder, E. J. et al. A unifying structural and functional model of the coronavirus replication organelle: tracking down RNA synthesis. PLoS Biol. 18, e3000715 (2020).",
"volume": "18",
"year": "2020"
},
{
"DOI": "10.1126/science.abd3629",
"author": "G Wolff",
"doi-asserted-by": "publisher",
"first-page": "1395",
"journal-title": "Science",
"key": "958_CR16",
"unstructured": "Wolff, G. et al. A molecular pore spans the double membrane of the coronavirus replication organelle. Science 369, 1395–1398 (2020).",
"volume": "369",
"year": "2020"
},
{
"DOI": "10.1016/j.virol.2006.11.027",
"author": "S Stertz",
"doi-asserted-by": "publisher",
"first-page": "304",
"journal-title": "Virology",
"key": "958_CR17",
"unstructured": "Stertz, S. et al. The intracellular sites of early replication and budding of SARS-coronavirus. Virology 361, 304–315 (2007).",
"volume": "361",
"year": "2007"
},
{
"DOI": "10.1016/j.cell.2020.10.039",
"author": "S Ghosh",
"doi-asserted-by": "publisher",
"first-page": "1520",
"journal-title": "Cell",
"key": "958_CR18",
"unstructured": "Ghosh, S. et al. β-Coronaviruses use lysosomes for egress instead of the biosynthetic secretory pathway. Cell 183, 1520–1535 (2020).",
"volume": "183",
"year": "2020"
},
{
"DOI": "10.1038/nm1267",
"author": "K Kuba",
"doi-asserted-by": "publisher",
"first-page": "875",
"journal-title": "Nat. Med.",
"key": "958_CR19",
"unstructured": "Kuba, K. et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 11, 875–879 (2005).",
"volume": "11",
"year": "2005"
},
{
"DOI": "10.1038/s41586-020-2180-5",
"author": "J Lan",
"doi-asserted-by": "publisher",
"first-page": "215",
"journal-title": "Nature",
"key": "958_CR20",
"unstructured": "Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).",
"volume": "581",
"year": "2020"
},
{
"DOI": "10.1126/science.abb2762",
"author": "R Yan",
"doi-asserted-by": "publisher",
"first-page": "1444",
"journal-title": "Science",
"key": "958_CR21",
"unstructured": "Yan, R. et al. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 367, 1444–1448 (2020).",
"volume": "367",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.02.052",
"doi-asserted-by": "publisher",
"key": "958_CR22",
"unstructured": "Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell https://doi.org/10.1016/j.cell.2020.02.052 (2020)."
},
{
"DOI": "10.1038/s41586-020-2012-7",
"author": "P Zhou",
"doi-asserted-by": "publisher",
"first-page": "270",
"journal-title": "Nature",
"key": "958_CR23",
"unstructured": "Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273 (2020).",
"volume": "579",
"year": "2020"
},
{
"DOI": "10.1038/s41586-020-2312-y",
"author": "L Bao",
"doi-asserted-by": "publisher",
"first-page": "830",
"journal-title": "Nature",
"key": "958_CR24",
"unstructured": "Bao, L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature 583, 830–833 (2020).",
"volume": "583",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.10.028",
"doi-asserted-by": "publisher",
"key": "958_CR25",
"unstructured": "Wei, J. et al. Genome-wide CRISPR screens reveal host factors critical for SARS-CoV-2 infection. Cell https://doi.org/10.1016/j.cell.2020.10.028 (2021)."
},
{
"DOI": "10.1016/j.cell.2020.12.006",
"author": "WM Schneider",
"doi-asserted-by": "publisher",
"first-page": "120",
"journal-title": "Cell",
"key": "958_CR26",
"unstructured": "Schneider, W. M. et al. Genome-scale identification of SARS-CoV-2 and pan-coronavirus host factor networks. Cell 184, 120–132 (2021).",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.1016/j.chom.2020.12.009",
"author": "H Hoffmann",
"doi-asserted-by": "publisher",
"first-page": "267",
"journal-title": "Cell Host Microbe",
"key": "958_CR27",
"unstructured": "Hoffmann, H. et al. Functional interrogation of a SARS-CoV-2 host protein interactome identifies unique and shared coronavirus host factors. Cell Host Microbe 29, 267–280 (2020).",
"volume": "29",
"year": "2020"
},
{
"DOI": "10.1038/s41586-020-2286-9",
"author": "DE Gordon",
"doi-asserted-by": "publisher",
"first-page": "459",
"journal-title": "Nature",
"key": "958_CR28",
"unstructured": "Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).",
"volume": "583",
"year": "2020"
},
{
"DOI": "10.1101/2021.04.22.440848",
"doi-asserted-by": "publisher",
"key": "958_CR29",
"unstructured": "Biering, S. B. et al. Genome-wide, bidirectional CRISPR screens identify mucins as critical host factors modulating SARS-CoV-2 infection. Preprint at bioRxiv https://doi.org/10.1101/2021.04.22.440848 (2021)."
},
{
"DOI": "10.1101/2021.05.19.444823",
"doi-asserted-by": "publisher",
"key": "958_CR30",
"unstructured": "Rebendenne, A. et al. Bidirectional genome-wide CRISPR screens reveal host factors regulating SARS-CoV-2, MERS-CoV and seasonal coronaviruses. Preprint at bioRxiv https://doi.org/10.1101/2021.05.19.444823 (2021)."
},
{
"DOI": "10.1038/s41422-020-00460-y",
"author": "S Wang",
"doi-asserted-by": "publisher",
"first-page": "126",
"journal-title": "Cell Res.",
"key": "958_CR31",
"unstructured": "Wang, S. et al. AXL is a candidate receptor for SARS-CoV-2 that promotes infection of pulmonary and bronchial epithelial cells. Cell Res. 31, 126–140 (2021).",
"volume": "31",
"year": "2021"
},
{
"DOI": "10.1016/j.celrep.2021.109364",
"doi-asserted-by": "publisher",
"key": "958_CR32",
"unstructured": "Puray-Chavez, M. et al. Systematic analysis of SARS-CoV-2 infection of an ACE2-negative human airway cell. Cell Rep. https://doi.org/10.1016/j.celrep.2021.109364 (2021)."
},
{
"DOI": "10.1016/j.cell.2021.02.053",
"doi-asserted-by": "publisher",
"key": "958_CR33",
"unstructured": "Yeung, M. L. et al. Soluble ACE2-mediated cell entry of SARS-CoV-2 via interaction with proteins related to the renin-angiotensin system. Cell https://doi.org/10.1016/j.cell.2021.02.053 (2021)."
},
{
"DOI": "10.1016/j.it.2020.10.004",
"author": "AG Harrison",
"doi-asserted-by": "publisher",
"first-page": "1100",
"journal-title": "Trends Immunol.",
"key": "958_CR34",
"unstructured": "Harrison, A. G., Lin, T. & Wang, P. Mechanisms of SARS-CoV-2 transmission and pathogenesis. Trends Immunol. 41, 1100–1115 (2020).",
"volume": "41",
"year": "2020"
},
{
"DOI": "10.1016/S2666-5247(20)30144-0",
"author": "B Schurink",
"doi-asserted-by": "publisher",
"first-page": "e290",
"journal-title": "Lancet Microbe",
"key": "958_CR35",
"unstructured": "Schurink, B. et al. Viral presence and immunopathology in patients with lethal COVID-19: a prospective autopsy cohort study. Lancet Microbe 1, e290–e299 (2020).",
"volume": "1",
"year": "2020"
},
{
"DOI": "10.1056/NEJMc2011400",
"author": "VG Puelles",
"doi-asserted-by": "publisher",
"first-page": "590",
"journal-title": "N. Engl. J. Med.",
"key": "958_CR36",
"unstructured": "Puelles, V. G. et al. Multiorgan and renal tropism of SARS-CoV-2. N. Engl. J. Med. 383, 590–592 (2020).",
"volume": "383",
"year": "2020"
},
{
"DOI": "10.1038/s41587-020-0602-4",
"author": "RL Chua",
"doi-asserted-by": "publisher",
"first-page": "970",
"journal-title": "Nat. Biotechnol.",
"key": "958_CR37",
"unstructured": "Chua, R. L. et al. COVID-19 severity correlates with airway epithelium–immune cell interactions identified by single-cell analysis. Nat. Biotechnol. 38, 970–979 (2020).",
"volume": "38",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.04.035",
"author": "CGK Ziegler",
"doi-asserted-by": "publisher",
"first-page": "1016",
"journal-title": "Cell",
"key": "958_CR38",
"unstructured": "Ziegler, C. G. K. et al. SARS-CoV-2 receptor ACE2 is an interferon-stimulated gene in human airway epithelial cells and is detected in specific cell subsets across tissues. Cell 181, 1016–1035 (2020).",
"volume": "181",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.05.006",
"author": "P Bost",
"doi-asserted-by": "publisher",
"first-page": "1475",
"journal-title": "Cell",
"key": "958_CR39",
"unstructured": "Bost, P. et al. Host-viral infection maps reveal signatures of severe COVID-19 patients. Cell 181, 1475–1488 (2020).",
"volume": "181",
"year": "2020"
},
{
"DOI": "10.1021/acscentsci.1c00010",
"author": "L Liu",
"doi-asserted-by": "publisher",
"first-page": "1009",
"journal-title": "ACS Cent. Sci.",
"key": "958_CR40",
"unstructured": "Liu, L. et al. Heparan sulfate proteoglycans as attachment factor for SARS-CoV-2. ACS Cent. Sci. 7, 1009–1018 (2021).",
"volume": "7",
"year": "2021"
},
{
"DOI": "10.1038/s41421-020-00222-5",
"author": "Q Zhang",
"doi-asserted-by": "publisher",
"first-page": "80",
"journal-title": "Cell Discov.",
"key": "958_CR41",
"unstructured": "Zhang, Q. et al. Heparan sulfate assists SARS-CoV-2 in cell entry and can be targeted by approved drugs in vitro. Cell Discov. 6, 80 (2020).",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.09.033",
"author": "TM Clausen",
"doi-asserted-by": "publisher",
"first-page": "1043",
"journal-title": "Cell",
"key": "958_CR42",
"unstructured": "Clausen, T. M. et al. SARS-CoV-2 infection depends on cellular heparan sulfate and ACE2. Cell 183, 1043–1057 (2020).",
"volume": "183",
"year": "2020"
},
{
"DOI": "10.1038/s41588-021-00805-2",
"doi-asserted-by": "publisher",
"key": "958_CR43",
"unstructured": "Baggen, J. et al. Genome-wide CRISPR screening identifies TMEM106B as a proviral host factor for SARS-CoV-2. Nat. Genet. https://doi.org/10.1038/s41588-021-00805-2 (2021)."
},
{
"DOI": "10.3390/v11070596",
"author": "V Cagno",
"doi-asserted-by": "publisher",
"first-page": "596",
"journal-title": "Viruses",
"key": "958_CR44",
"unstructured": "Cagno, V., Tseligka, E. D., Jones, S. T. & Tapparel, C. Heparan sulfate proteoglycans and viral attachment: true receptors or adaptation bias? Viruses 11, 596 (2019).",
"volume": "11",
"year": "2019"
},
{
"DOI": "10.1038/s42255-020-00324-0",
"author": "C Wei",
"doi-asserted-by": "publisher",
"first-page": "1391",
"journal-title": "Nat. Metab.",
"key": "958_CR45",
"unstructured": "Wei, C. et al. HDL-scavenger receptor B type 1 facilitates SARS-CoV-2 entry. Nat. Metab. 2, 1391–1400 (2020).",
"volume": "2",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.02.058",
"author": "AC Walls",
"doi-asserted-by": "publisher",
"first-page": "281",
"journal-title": "Cell",
"key": "958_CR46",
"unstructured": "Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 Spike glycoprotein. Cell 181, 281–292 (2020).",
"volume": "181",
"year": "2020"
},
{
"DOI": "10.1126/science.abd3072",
"author": "JL Daly",
"doi-asserted-by": "publisher",
"first-page": "861",
"journal-title": "Science",
"key": "958_CR47",
"unstructured": "Daly, J. L. et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 370, 861–865 (2020).",
"volume": "370",
"year": "2020"
},
{
"DOI": "10.1126/science.abd2985",
"author": "L Cantuti-Castelvetri",
"doi-asserted-by": "publisher",
"first-page": "856",
"journal-title": "Science",
"key": "958_CR48",
"unstructured": "Cantuti-Castelvetri, L. et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 370, 856–860 (2020).",
"volume": "370",
"year": "2020"
},
{
"DOI": "10.1101/2021.04.08.438924",
"doi-asserted-by": "publisher",
"key": "958_CR49",
"unstructured": "Zhu, S. et al. Genome-wide CRISPR activation screen identifies novel receptors for SARS-CoV-2 entry. Preprint at bioRxiv https://doi.org/10.1101/2021.04.08.438924 (2021)."
},
{
"DOI": "10.1038/s41392-020-00426-x",
"author": "K Wang",
"doi-asserted-by": "publisher",
"first-page": "283",
"journal-title": "Signal Transduct. Target. Ther.",
"key": "958_CR50",
"unstructured": "Wang, K. et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target. Ther. 5, 283 (2020).",
"volume": "5",
"year": "2020"
},
{
"DOI": "10.1038/s41598-020-80464-1",
"author": "J Shilts",
"doi-asserted-by": "publisher",
"first-page": "413",
"journal-title": "Sci. Rep.",
"key": "958_CR51",
"unstructured": "Shilts, J., Crozier, T. W. M., Greenwood, E. J. D., Lehner, P. J. & Wright, G. J. No evidence for basigin/CD147 as a direct SARS-CoV-2 spike binding receptor. Sci. Rep. 11, 413 (2021).",
"volume": "11",
"year": "2021"
},
{
"DOI": "10.1101/2020.09.09.287508",
"doi-asserted-by": "publisher",
"key": "958_CR52",
"unstructured": "Gu, Y. et al. Interaction network of SARS-CoV-2 with host receptome through spike protein. Preprint at bioRxiv https://doi.org/10.1101/2020.09.09.287508 (2020)."
},
{
"DOI": "10.1101/2020.10.23.350348",
"doi-asserted-by": "publisher",
"key": "958_CR53",
"unstructured": "Tang, X. et al. Transferrin receptor is another receptor for SARS-CoV-2 entry. Preprint at bioRxiv https://doi.org/10.1101/2020.10.23.350348 (2020)."
},
{
"DOI": "10.1101/2020.11.05.369264",
"doi-asserted-by": "publisher",
"key": "958_CR54",
"unstructured": "Soh, W. T. et al. The N-terminal domain of spike glycoprotein mediates SARS-CoV-2 infection by associating with L-SIGN and DC-SIGN. Preprint at bioRxiv https://doi.org/10.1101/2020.11.05.369264 (2020)."
},
{
"DOI": "10.1101/2020.07.29.227462",
"doi-asserted-by": "publisher",
"key": "958_CR55",
"unstructured": "Gao, C. et al. SARS-CoV-2 spike protein interacts with multiple innate immune receptors. Preprint at bioRxiv https://doi.org/10.1101/2020.07.29.227462 (2020)."
},
{
"DOI": "10.1021/acscentsci.0c01537",
"author": "R Amraei",
"doi-asserted-by": "publisher",
"first-page": "1156",
"journal-title": "ACS Cent. Sci.",
"key": "958_CR56",
"unstructured": "Amraei, R. et al. CD209L/L-SIGN and CD209/DC-SIGN act as receptors for SARS-CoV-2. ACS Cent. Sci. 7, 1156–1165 (2021).",
"volume": "7",
"year": "2021"
},
{
"DOI": "10.1371/journal.ppat.1009576",
"author": "M Thépaut",
"doi-asserted-by": "publisher",
"first-page": "e1009576",
"journal-title": "PLoS Pathog.",
"key": "958_CR57",
"unstructured": "Thépaut, M. et al. DC/L-SIGN recognition of spike glycoprotein promotes SARS-CoV-2 trans-infection and can be inhibited by a glycomimetic antagonist. PLoS Pathog. 17, e1009576 (2021).",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.3390/biology10010001",
"author": "N Rahimi",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "Biology",
"key": "958_CR58",
"unstructured": "Rahimi, N. C-type lectin CD209L/L-SIGN and CD209/DC-SIGN: cell adhesion molecules turned to pathogen recognition receptors. Biology 10, 1 (2021).",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1371/journal.ppat.1009500",
"author": "M Laporte",
"doi-asserted-by": "publisher",
"first-page": "e1009500",
"journal-title": "PLOS Pathogens",
"key": "958_CR59",
"unstructured": "Laporte, M. et al. The SARS-CoV-2 and other human coronavirus spike proteins are fine-tuned towards temperature and proteases of the human airways. PLOS Pathogens 17, e1009500 (2021).",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.1016/j.molcel.2020.04.022",
"author": "M Hoffmann",
"doi-asserted-by": "publisher",
"first-page": "779",
"journal-title": "Mol. Cell",
"key": "958_CR60",
"unstructured": "Hoffmann, M., Kleine-Weber, H. & Pöhlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 78, 779–784 (2020).",
"volume": "78",
"year": "2020"
},
{
"DOI": "10.1371/journal.ppat.1009246",
"author": "G Papa",
"doi-asserted-by": "publisher",
"first-page": "e1009246",
"journal-title": "PLoS Pathog.",
"key": "958_CR61",
"unstructured": "Papa, G. et al. Furin cleavage of SARS-CoV-2 Spike promotes but is not essential for infection and cell-cell fusion. PLoS Pathog. 17, e1009246 (2021).",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.1038/s41586-021-03237-4",
"author": "BA Johnson",
"doi-asserted-by": "publisher",
"first-page": "293",
"journal-title": "Nature",
"key": "958_CR62",
"unstructured": "Johnson, B. A. et al. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature 591, 293–299 (2021).",
"volume": "591",
"year": "2021"
},
{
"DOI": "10.1080/22221751.2020.1756700",
"author": "SY Lau",
"doi-asserted-by": "publisher",
"first-page": "837",
"journal-title": "Emerg. Microbes Infect.",
"key": "958_CR63",
"unstructured": "Lau, S. Y. et al. Attenuated SARS-CoV-2 variants with deletions at the S1/S2 junction. Emerg. Microbes Infect. 9, 837–842 (2020).",
"volume": "9",
"year": "2020"
},
{
"DOI": "10.1038/s41467-021-21213-4",
"doi-asserted-by": "publisher",
"key": "958_CR64",
"unstructured": "Zhu, Y. et al. A genome-wide CRISPR screen identifies host factors that regulate SARS-CoV-2 entry. Nat. Commun. https://doi.org/10.1038/s41467-021-21213-4 (2021)."
},
{
"DOI": "10.1038/s41467-020-15562-9",
"author": "X Ou",
"doi-asserted-by": "publisher",
"first-page": "1620",
"journal-title": "Nat. Commun.",
"key": "958_CR65",
"unstructured": "Ou, X. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 11, 1620 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1021/acsinfecdis.0c00701",
"doi-asserted-by": "publisher",
"key": "958_CR66",
"unstructured": "Tang, T. et al. Proteolytic activation of SARS-CoV-2 Spike at the S1/S2 boundary: potential role of proteases beyond furin. ACS Infect. Dis. https://doi.org/10.1021/acsinfecdis.0c00701 (2021)."
},
{
"DOI": "10.1126/sciimmunol.abc3582",
"author": "R Zang",
"doi-asserted-by": "publisher",
"first-page": "eabc3582",
"journal-title": "Sci. Immunol.",
"key": "958_CR67",
"unstructured": "Zang, R. et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 5, eabc3582 (2020).",
"volume": "5",
"year": "2020"
},
{
"DOI": "10.1038/s41591-020-0868-6",
"author": "W Sungnak",
"doi-asserted-by": "publisher",
"first-page": "681",
"journal-title": "Nat. Med.",
"key": "958_CR68",
"unstructured": "Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 26, 681–687 (2020).",
"volume": "26",
"year": "2020"
},
{
"DOI": "10.1016/j.isci.2020.101212",
"author": "JA Jaimes",
"doi-asserted-by": "publisher",
"first-page": "101212",
"journal-title": "Iscience",
"key": "958_CR69",
"unstructured": "Jaimes, J. A., Millet, J. K. & Whittaker, G. R. Proteolytic cleavage of the SARS-CoV-2 Spike protein and the role of the novel S1/S2 site. Iscience 23, 101212 (2020).",
"volume": "23",
"year": "2020"
},
{
"DOI": "10.1152/physrev.00013.2020",
"author": "HL Ji",
"doi-asserted-by": "publisher",
"first-page": "1065",
"journal-title": "Physiol. Rev.",
"key": "958_CR70",
"unstructured": "Ji, H. L., Zhao, R., Matalon, S. & Matthay, M. A. Elevated plasmin(Ogen) as a common risk factor for COVID-19 susceptibility. Physiol. Rev. 100, 1065–1075 (2020).",
"volume": "100",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.10.030",
"author": "Z Daniloski",
"doi-asserted-by": "publisher",
"first-page": "92",
"journal-title": "Cell",
"key": "958_CR71",
"unstructured": "Daniloski, Z. et al. Identification of required host factors for SARS-CoV-2 infection in human cells. Cell 184, 92–105 (2021).",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2020.12.004",
"doi-asserted-by": "publisher",
"key": "958_CR72",
"unstructured": "Wang, R. et al. Genetic screens identify host factors for SARS-CoV-2 and common cold coronaviruses. Cell https://doi.org/10.1016/j.cell.2020.12.004 (2020)."
},
{
"DOI": "10.1016/j.cell.2021.03.012",
"doi-asserted-by": "publisher",
"key": "958_CR73",
"unstructured": "Flynn, R. A. et al. Discovery and functional interrogation of SARS-CoV-2 RNA-host protein interactions. Cell https://doi.org/10.1016/j.cell.2021.03.012 (2021)."
},
{
"DOI": "10.1038/s41467-020-19032-0",
"author": "DJH van den Boomen",
"doi-asserted-by": "publisher",
"first-page": "5559",
"journal-title": "Nat. Commun.",
"key": "958_CR74",
"unstructured": "van den Boomen, D. J. H. et al. A trimeric Rab7 GEF controls NPC1-dependent lysosomal cholesterol export. Nat. Commun. 11, 5559 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1371/journal.ppat.1004502",
"author": "C Burkard",
"doi-asserted-by": "publisher",
"first-page": "e1004502",
"journal-title": "PLoS Pathog.",
"key": "958_CR75",
"unstructured": "Burkard, C. et al. Coronavirus cell entry occurs through the endo-/lysosomal pathway in a proteolysis-dependent manner. PLoS Pathog. 10, e1004502 (2014).",
"volume": "10",
"year": "2014"
},
{
"DOI": "10.1038/nature10348",
"author": "JE Carette",
"doi-asserted-by": "publisher",
"first-page": "340",
"journal-title": "Nature",
"key": "958_CR76",
"unstructured": "Carette, J. E. et al. Ebola virus entry requires the cholesterol transporter Niemann-Pick C1. Nature 477, 340–343 (2011).",
"volume": "477",
"year": "2011"
},
{
"DOI": "10.1016/j.devcel.2020.12.010",
"author": "G Miao",
"doi-asserted-by": "publisher",
"first-page": "427",
"journal-title": "Dev. Cell",
"key": "958_CR77",
"unstructured": "Miao, G. et al. ORF3a of the COVID-19 virus SARS-CoV-2 blocks HOPS complex-mediated assembly of the SNARE complex required for autolysosome formation. Dev. Cell 56, 427–442 (2021).",
"volume": "56",
"year": "2021"
},
{
"DOI": "10.1126/science.abe9403",
"author": "DE Gordon",
"doi-asserted-by": "publisher",
"first-page": "eabe9403",
"journal-title": "Science",
"key": "958_CR78",
"unstructured": "Gordon, D. E. et al. Comparative host-coronavirus protein interaction networks reveal pan-viral disease mechanisms. Science 370, eabe9403 (2020).",
"volume": "370",
"year": "2020"
},
{
"DOI": "10.1016/j.omtn.2017.12.016",
"author": "S Peng",
"doi-asserted-by": "publisher",
"first-page": "361",
"journal-title": "Mol. Ther. Nucleic Acids",
"key": "958_CR79",
"unstructured": "Peng, S. et al. Endogenous cellular microRNAs mediate antiviral defense against influenza A virus. Mol. Ther. Nucleic Acids 10, 361–375 (2018).",
"volume": "10",
"year": "2018"
},
{
"DOI": "10.1371/journal.pone.0197246",
"author": "MA Estrin",
"doi-asserted-by": "publisher",
"first-page": "e0197246",
"journal-title": "PLoS ONE",
"key": "958_CR80",
"unstructured": "Estrin, M. A. et al. Host-directed combinatorial RNAi improves inhibition of diverse strains of influenza A virus in human respiratory epithelial cells. PLoS ONE 13, e0197246 (2018).",
"volume": "13",
"year": "2018"
},
{
"DOI": "10.1016/j.neuron.2017.06.026",
"author": "ZA Klein",
"doi-asserted-by": "publisher",
"first-page": "281",
"journal-title": "Neuron",
"key": "958_CR81",
"unstructured": "Klein, Z. A. et al. Loss of TMEM106B ameliorates lysosomal and frontotemporal dementia-related phenotypes in article loss of TMEM106B ameliorates lysosomal and frontotemporal dementia-related phenotypes in progranulin-deficient mice. Neuron 95, 281–296 (2017).",
"volume": "95",
"year": "2017"
},
{
"DOI": "10.1016/j.tcb.2018.06.005",
"author": "KE McNally",
"doi-asserted-by": "publisher",
"first-page": "807",
"journal-title": "Trends Cell Biol.",
"key": "958_CR82",
"unstructured": "McNally, K. E. & Cullen, P. J. Endosomal retrieval of cargo: retromer is not alone. Trends Cell Biol. 28, 807–822 (2018).",
"volume": "28",
"year": "2018"
},
{
"DOI": "10.1016/j.devcel.2009.09.010",
"author": "E Derivery",
"doi-asserted-by": "publisher",
"first-page": "712",
"journal-title": "Dev. Cell",
"key": "958_CR83",
"unstructured": "Derivery, E. et al. The Arp2/3 activator WASH controls the fission of endosomes through a large multiprotein complex. Dev. Cell 17, 712–723 (2009).",
"volume": "17",
"year": "2009"
},
{
"DOI": "10.1038/ncb1505",
"author": "X Zuo",
"doi-asserted-by": "publisher",
"first-page": "1383",
"journal-title": "Nat. Cell Biol.",
"key": "958_CR84",
"unstructured": "Zuo, X. et al. Exo70 interacts with the Arp2/3 complex and regulates cell migration. Nat. Cell Biol. 8, 1383–1388 (2006).",
"volume": "8",
"year": "2006"
},
{
"DOI": "10.1371/journal.pone.0214059",
"author": "J Al-Saleem",
"doi-asserted-by": "publisher",
"first-page": "e0214059",
"journal-title": "PLoS ONE",
"key": "958_CR85",
"unstructured": "Al-Saleem, J. et al. HTLV-1 Tax-1 interacts with SNX27 to regulate cellular localization of the HTLV-1 receptor molecule, GLUT1. PLoS ONE 14, e0214059 (2019).",
"volume": "14",
"year": "2019"
},
{
"DOI": "10.1073/pnas.1302164110",
"author": "A Lipovsky",
"doi-asserted-by": "publisher",
"first-page": "7452",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "958_CR86",
"unstructured": "Lipovsky, A. et al. Genome-wide siRNA screen identifies the retromer as a cellular entry factor for human papillomavirus. Proc. Natl Acad. Sci. USA 110, 7452–7457 (2013).",
"volume": "110",
"year": "2013"
},
{
"DOI": "10.1007/s00018-015-2027-7",
"author": "P Yin",
"doi-asserted-by": "publisher",
"first-page": "869",
"journal-title": "Cell. Mol. Life Sci.",
"key": "958_CR87",
"unstructured": "Yin, P., Hong, Z., Yang, X., Chung, R. T. & Zhang, L. A role for retromer in hepatitis C virus replication. Cell. Mol. Life Sci. 73, 869–881 (2016).",
"volume": "73",
"year": "2016"
},
{
"DOI": "10.1042/BCJ20160170",
"author": "JM Backer",
"doi-asserted-by": "publisher",
"first-page": "2251",
"journal-title": "Biochem. J.",
"key": "958_CR88",
"unstructured": "Backer, J. M. The intricate regulation and complex functions of the class III phosphoinositide 3-kinase Vps34. Biochem. J. 473, 2251–2271 (2016).",
"volume": "473",
"year": "2016"
},
{
"DOI": "10.1101/2021.02.24.432634",
"doi-asserted-by": "publisher",
"key": "958_CR89",
"unstructured": "Kratzel, A. et al. A genome-wide CRISPR screen identifies interactors of the autophagy pathway as conserved coronavirus targets. Preprint at bioRxiv https://doi.org/10.1101/2021.02.24.432634 (2021)."
},
{
"DOI": "10.15252/embr.201845889",
"author": "F Moretti",
"doi-asserted-by": "publisher",
"first-page": "e45889",
"journal-title": "EMBO Rep.",
"key": "958_CR90",
"unstructured": "Moretti, F. et al. TMEM41B is a novel regulator of autophagy and lipid mobilization. EMBO Rep. 19, e45889 (2018).",
"volume": "19",
"year": "2018"
},
{
"DOI": "10.1016/j.cell.2020.12.005",
"doi-asserted-by": "publisher",
"key": "958_CR91",
"unstructured": "Hoffmann, H. et al. TMEM41B is a pan-flavivirus host factor. Cell https://doi.org/10.1016/j.cell.2020.12.005 (2020)."
},
{
"DOI": "10.1073/pnas.96.20.11041",
"author": "MS Brown",
"doi-asserted-by": "publisher",
"first-page": "11041",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "958_CR92",
"unstructured": "Brown, M. S. & Goldstein, J. L. A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood. Proc. Natl Acad. Sci. USA 96, 11041–11048 (1999).",
"volume": "96",
"year": "1999"
},
{
"DOI": "10.1016/j.tcb.2020.02.007",
"author": "Y Meng",
"doi-asserted-by": "publisher",
"first-page": "452",
"journal-title": "Trends Cell Biol.",
"key": "958_CR93",
"unstructured": "Meng, Y., Heybrock, S., Neculai, D. & Saftig, P. Cholesterol handling in lysosomes and beyond. Trends Cell Biol. 30, 452–466 (2020).",
"volume": "30",
"year": "2020"
},
{
"DOI": "10.15252/embj.2020106057",
"author": "S Wang",
"doi-asserted-by": "publisher",
"first-page": "e106057",
"journal-title": "EMBO J.",
"key": "958_CR94",
"unstructured": "Wang, S. et al. Cholesterol 25-hydroxylase inhibits SARS-CoV-2 and other coronaviruses by depleting membrane cholesterol. EMBO J. 39, e106057 (2020).",
"volume": "39",
"year": "2020"
},
{
"DOI": "10.3389/fcimb.2018.00388",
"author": "JF Osuna-Ramos",
"doi-asserted-by": "publisher",
"first-page": "388",
"journal-title": "Front. Cell. Infect. Microbiol.",
"key": "958_CR95",
"unstructured": "Osuna-Ramos, J. F., Reyes-Ruiz, J. M. & Del Ángel, R. M. The role of host cholesterol during flavivirus infection. Front. Cell. Infect. Microbiol. 8, 388 (2018).",
"volume": "8",
"year": "2018"
},
{
"DOI": "10.1101/2021.06.10.447768",
"doi-asserted-by": "publisher",
"key": "958_CR96",
"unstructured": "Sherman, E. J. et al. Identification of ACE2 modifiers by CRISPR screening. Preprint at bioRxiv https://doi.org/10.1101/2021.06.10.447768 (2021)."
},
{
"DOI": "10.3389/fmicb.2020.00024",
"author": "AR Murphy Schafer",
"doi-asserted-by": "publisher",
"first-page": "24",
"journal-title": "Front. Microbiol.",
"key": "958_CR97",
"unstructured": "Murphy Schafer, A. R., Smith, J. L., Pryke, K. M., DeFilippis, V. R. & Hirsch, A. J. The E3 ubiquitin ligase SIAH1 targets MyD88 for proteasomal degradation during dengue virus infection. Front. Microbiol. 11, 24 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1371/journal.ppat.1009366",
"author": "W Zhang",
"doi-asserted-by": "publisher",
"first-page": "e1009366",
"journal-title": "PLOS Pathog.",
"key": "958_CR98",
"unstructured": "Zhang, W. et al. JMJD6 negatively regulates cytosolic RNA induced antiviral signaling by recruiting RNF5 to promote activated IRF3 K48 ubiquitination. PLOS Pathog. 17, e1009366 (2021).",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.1016/j.molcel.2021.04.022",
"author": "S Lee",
"doi-asserted-by": "publisher",
"first-page": "2838",
"journal-title": "Mol. Cell",
"key": "958_CR99",
"unstructured": "Lee, S. et al. The SARS-CoV-2 RNA interactome. Mol. Cell 81, 2838–2850 (2021).",
"volume": "81",
"year": "2021"
},
{
"DOI": "10.1038/s41564-020-00846-z",
"author": "N Schmidt",
"doi-asserted-by": "publisher",
"first-page": "339",
"journal-title": "Nat. Microbiol.",
"key": "958_CR100",
"unstructured": "Schmidt, N. et al. The SARS-CoV-2 RNA–protein interactome in infected human cells. Nat. Microbiol. 6, 339–353 (2020).",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1016/j.molcel.2021.05.023",
"author": "W Kamel",
"doi-asserted-by": "publisher",
"first-page": "2851",
"journal-title": "Mol. Cell",
"key": "958_CR101",
"unstructured": "Kamel, W. et al. Global analysis of protein-RNA interactions in SARS-CoV-2-infected cells reveals key regulators of infection. Mol. Cell 81, 2851–2867 (2021).",
"volume": "81",
"year": "2021"
},
{
"DOI": "10.1101/2021.03.23.436611",
"doi-asserted-by": "publisher",
"key": "958_CR102",
"unstructured": "Labeau, A. et al. Characterization and functional interrogation of SARS-CoV-2 RNA interactome. Preprint at bioRxiv https://doi.org/10.1101/2021.03.23.436611 (2021)."
},
{
"DOI": "10.1021/acsinfecdis.0c00500",
"author": "JP Davies",
"doi-asserted-by": "publisher",
"first-page": "3174",
"journal-title": "ACS Infect. Dis.",
"key": "958_CR103",
"unstructured": "Davies, J. P., Almasy, K. M., McDonald, E. F. & Plate, L. Comparative multiplexed interactomics of SARS-CoV-2 and homologous coronavirus nonstructural proteins identifies unique and shared host-cell dependencies. ACS Infect. Dis. 6, 3174–3189 (2020).",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1016/j.medj.2020.07.002",
"author": "J Li",
"doi-asserted-by": "publisher",
"first-page": "99",
"journal-title": "Med",
"key": "958_CR104",
"unstructured": "Li, J. et al. Virus-host interactome and proteomic survey reveal potential virulence factors influencing SARS-CoV-2 pathogenesis. Med 2, 99–112 (2021).",
"volume": "2",
"year": "2021"
},
{
"DOI": "10.1038/s41586-021-03493-4",
"doi-asserted-by": "publisher",
"key": "958_CR105",
"unstructured": "Stukalov, A. et al. Multilevel proteomics reveals host perturbations by SARS-CoV-2 and SARS-CoV. Nature https://doi.org/10.1038/s41586-021-03493-4 (2021)."
},
{
"DOI": "10.1101/2020.09.03.282103",
"doi-asserted-by": "publisher",
"key": "958_CR106",
"unstructured": "Samavarchi-Tehrani, P. et al. A SARS-CoV-2–host proximity interactome. Preprint at bioRxiv https://doi.org/10.1101/2020.09.03.282103 (2020)."
},
{
"DOI": "10.1101/2020.08.28.269175",
"doi-asserted-by": "publisher",
"key": "958_CR107",
"unstructured": "St-Germain, J. R. et al. A SARS-CoV-2 BioID-based virus-host membrane protein interactome and virus peptide compendium: new proteomics resources for COVID-19 research. Preprint at bioRxiv https://doi.org/10.1101/2020.08.28.269175 (2020)."
},
{
"DOI": "10.1101/2020.08.28.272955",
"doi-asserted-by": "publisher",
"key": "958_CR108",
"unstructured": "Laurent, E. M. N. et al. Global BioID-based SARS-CoV-2 proteins proximal interactome unveils novel ties between viral polypeptides and host factors involved in multiple COVID19-associated mechanisms. Preprint at bioRxiv https://doi.org/10.1101/2020.08.28.272955 (2020)."
},
{
"DOI": "10.1016/j.virol.2015.03.001",
"author": "RE Lloyd",
"doi-asserted-by": "publisher",
"first-page": "457",
"journal-title": "Virology",
"key": "958_CR109",
"unstructured": "Lloyd, R. E. Nuclear proteins hijacked by mammalian cytoplasmic plus strand RNA viruses. Virology 479, 457–474 (2015).",
"volume": "479",
"year": "2015"
},
{
"DOI": "10.3390/v10020070",
"author": "L Carrasco",
"doi-asserted-by": "publisher",
"first-page": "70",
"journal-title": "Viruses",
"key": "958_CR110",
"unstructured": "Carrasco, L., Sanz, M. A. & González-Almela, E. The regulation of translation in alphavirus-infected cells. Viruses 10, 70 (2018).",
"volume": "10",
"year": "2018"
},
{
"DOI": "10.1038/s41467-020-19996-z",
"author": "M Pietzner",
"doi-asserted-by": "publisher",
"first-page": "6397",
"journal-title": "Nat. Commun.",
"key": "958_CR111",
"unstructured": "Pietzner, M. et al. Genetic architecture of host proteins involved in SARS-CoV-2 infection. Nat. Commun. 11, 6397 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.3390/jcm9040982",
"author": "PH Guzzi",
"doi-asserted-by": "publisher",
"first-page": "982",
"journal-title": "J. Clin. Med.",
"key": "958_CR112",
"unstructured": "Guzzi, P. H., Mercatelli, D., Ceraolo, C. & Giorgi, F. M. Master regulator analysis of the SARS-CoV-2/human interactome. J. Clin. Med. 9, 982 (2020).",
"volume": "9",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.06.034",
"author": "M Bouhaddou",
"doi-asserted-by": "publisher",
"first-page": "685",
"journal-title": "Cell",
"key": "958_CR113",
"unstructured": "Bouhaddou, M. et al. The global phosphorylation landscape of SARS-CoV-2 infection. Cell 182, 685–712 (2020).",
"volume": "182",
"year": "2020"
},
{
"DOI": "10.1016/j.celrep.2017.08.076",
"author": "V Malikov",
"doi-asserted-by": "publisher",
"first-page": "2792",
"journal-title": "Cell Rep.",
"key": "958_CR114",
"unstructured": "Malikov, V. & Naghavi, M. H. Localized phosphorylation of a kinesin-1 adaptor by a capsid-associated kinase regulates HIV-1 motility and uncoating. Cell Rep. 20, 2792–2799 (2017).",
"volume": "20",
"year": "2017"
},
{
"DOI": "10.1128/JVI.01772-09",
"author": "MJ Gadlage",
"doi-asserted-by": "publisher",
"first-page": "280",
"journal-title": "J. Virol.",
"key": "958_CR115",
"unstructured": "Gadlage, M. J. et al. Murine hepatitis virus nonstructural protein 4 regulates virus-induced membrane modifications and replication complex function. J. Virol. 84, 280–290 (2010).",
"volume": "84",
"year": "2010"
},
{
"DOI": "10.1016/j.virol.2006.10.034",
"author": "CM Petit",
"doi-asserted-by": "publisher",
"first-page": "264",
"journal-title": "Virology",
"key": "958_CR116",
"unstructured": "Petit, C. M. et al. Palmitoylation of the cysteine-rich endodomain of the SARS–coronavirus spike glycoprotein is important for spike-mediated cell fusion. Virology 360, 264–274 (2007).",
"volume": "360",
"year": "2007"
},
{
"DOI": "10.1101/2020.12.20.423603",
"doi-asserted-by": "publisher",
"key": "958_CR117",
"unstructured": "Lee, M. et al. Fatty acid synthase inhibition prevents palmitoylation of SARS-CoV2 Spike protein and improves survival of mice infected with murine hepatitis virus. Preprint at bioRxiv https://doi.org/10.1101/2020.12.20.423603 (2020)."
},
{
"DOI": "10.3389/fphar.2020.592737",
"author": "CZ Chen",
"doi-asserted-by": "publisher",
"first-page": "592737",
"journal-title": "Front. Pharmacol.",
"key": "958_CR118",
"unstructured": "Chen, C. Z. et al. Drug repurposing screen for compounds inhibiting the cytopathic effect of SARS-CoV-2. Front. Pharmacol. 11, 592737 (2021).",
"volume": "11",
"year": "2021"
},
{
"DOI": "10.1007/s00406-020-01231-x",
"author": "K Hashimoto",
"doi-asserted-by": "publisher",
"first-page": "249",
"journal-title": "Eur. Arch. Psychiatry Clin. Neurosci.",
"key": "958_CR119",
"unstructured": "Hashimoto, K. Repurposing of CNS drugs to treat COVID-19 infection: targeting the sigma-1 receptor. Eur. Arch. Psychiatry Clin. Neurosci. 271, 249–258 (2021).",
"volume": "271",
"year": "2021"
},
{
"DOI": "10.1038/s41586-021-03491-6",
"doi-asserted-by": "publisher",
"key": "958_CR120",
"unstructured": "Braga, L. et al. Drugs inhibiting TMEM16 proteins block SARS-CoV-2 Spike-induced syncytia. Nature https://doi.org/10.1038/s41586-021-03491-6 (2021)."
},
{
"DOI": "10.1038/s41586-020-2577-1",
"author": "L Riva",
"doi-asserted-by": "publisher",
"first-page": "113",
"journal-title": "Nature",
"key": "958_CR121",
"unstructured": "Riva, L. et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 586, 113–119 (2020).",
"volume": "586",
"year": "2020"
},
{
"DOI": "10.15252/msb.20209628",
"author": "S Sinha",
"doi-asserted-by": "publisher",
"first-page": "e9628",
"journal-title": "Mol. Syst. Biol.",
"key": "958_CR122",
"unstructured": "Sinha, S. et al. In vitro and in vivo identification of clinically approved drugs that modify ACE 2 expression. Mol. Syst. Biol. 16, e9628 (2020).",
"volume": "16",
"year": "2020"
},
{
"DOI": "10.1165/rcmb.2020-0322ps",
"doi-asserted-by": "publisher",
"key": "958_CR123",
"unstructured": "Jia, H., Neptune, E. & Cui, H. Targeting ACE2 for COVID-19 therapy: opportunities and challenges. Am. J. Respir. Cell Mol. Biol. https://doi.org/10.1165/rcmb.2020-0322ps (2020)."
},
{
"DOI": "10.1161/CIRCULATIONAHA.104.510461",
"author": "CM Ferrario",
"doi-asserted-by": "publisher",
"first-page": "2605",
"journal-title": "Circulation",
"key": "958_CR124",
"unstructured": "Ferrario, C. M. et al. Effect of angiotensin-converting enzyme inhibition and angiotensin II receptor blockers on cardiac angiotensin-converting enzyme 2. Circulation 111, 2605–2610 (2005).",
"volume": "111",
"year": "2005"
},
{
"DOI": "10.1016/S2213-2600(20)30558-0",
"author": "JB Cohen",
"doi-asserted-by": "publisher",
"first-page": "275",
"journal-title": "Lancet Respir. Med.",
"key": "958_CR125",
"unstructured": "Cohen, J. B. et al. Continuation versus discontinuation of renin–angiotensin system inhibitors in patients admitted to hospital with COVID-19: a prospective, randomised, open-label trial. Lancet Respir. Med. 9, 275–284 (2021).",
"volume": "9",
"year": "2021"
},
{
"DOI": "10.1016/S2213-2600(21)00003-5",
"author": "B Williams",
"doi-asserted-by": "publisher",
"first-page": "221",
"journal-title": "Lancet Respir. Med.",
"key": "958_CR126",
"unstructured": "Williams, B. Renin-angiotensin system inhibitors in hospitalised patients with COVID-19. Lancet Respir. Med. 9, 221–222 (2021).",
"volume": "9",
"year": "2021"
},
{
"DOI": "10.1038/s41467-021-21972-0",
"author": "L Wettstein",
"doi-asserted-by": "publisher",
"first-page": "1726",
"journal-title": "Nat. Commun.",
"key": "958_CR127",
"unstructured": "Wettstein, L. et al. Alpha-1 antitrypsin inhibits TMPRSS2 protease activity and SARS-CoV-2 infection. Nat. Commun. 12, 1726 (2021).",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1210/me.2013-1098",
"author": "L Clinckemalie",
"doi-asserted-by": "publisher",
"first-page": "2028",
"journal-title": "Mol. Endocrinol.",
"key": "958_CR128",
"unstructured": "Clinckemalie, L. et al. Androgen regulation of the TMPRSS2 gene and the effect of a SNP in an androgen response element. Mol. Endocrinol. 27, 2028–2040 (2013).",
"volume": "27",
"year": "2013"
},
{
"DOI": "10.7150/ijbs.45498",
"author": "N Yang",
"doi-asserted-by": "publisher",
"first-page": "1724",
"journal-title": "Int. J. Biol. Sci.",
"key": "958_CR129",
"unstructured": "Yang, N. & Shen, H. M. Targeting the endocytic pathway and autophagy process as a novel therapeutic strategy in COVID-19. Int. J. Biol. Sci. 16, 1724–1731 (2020).",
"volume": "16",
"year": "2020"
},
{
"DOI": "10.1038/s41586-020-2575-3",
"author": "M Hoffmann",
"doi-asserted-by": "publisher",
"first-page": "588",
"journal-title": "Nature",
"key": "958_CR130",
"unstructured": "Hoffmann, M. et al. Chloroquine does not inhibit infection of human lung cells with SARS-CoV-2. Nature 585, 588–590 (2020).",
"volume": "585",
"year": "2020"
},
{
"DOI": "10.1074/jbc.M116.716100",
"author": "N Zhou",
"doi-asserted-by": "publisher",
"first-page": "9218",
"journal-title": "J. Biol. Chem.",
"key": "958_CR131",
"unstructured": "Zhou, N. et al. Glycopeptide antibiotics potently inhibit cathepsin l in the late endosome/lysosome and block the entry of ebola virus, middle east respiratory syndrome coronavirus (MERS-CoV), and severe acute respiratory syndrome coronavirus (SARS-CoV). J. Biol. Chem. 291, 9218–9232 (2016).",
"volume": "291",
"year": "2016"
},
{
"DOI": "10.1016/j.ijantimicag.2020.105944",
"author": "SA Baron",
"doi-asserted-by": "publisher",
"first-page": "105944",
"journal-title": "Int. J. Antimicrob. Agents",
"key": "958_CR132",
"unstructured": "Baron, S. A., Devaux, C., Colson, P., Raoult, D. & Rolain, J. Teicoplanin: an alternative drug for the treatment of COVID-19? Int. J. Antimicrob. Agents 55, 105944 (2020).",
"volume": "55",
"year": "2020"
},
{
"DOI": "10.2217/fvl-2020-0068",
"author": "J Nemunaitis",
"doi-asserted-by": "publisher",
"first-page": "381",
"journal-title": "Future Virol.",
"key": "958_CR133",
"unstructured": "Nemunaitis, J., Stanbery, L. & Senzer, N. Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2) infection: let the virus be its own demise. Future Virol. 15, 381–395 (2020).",
"volume": "15",
"year": "2020"
},
{
"DOI": "10.1038/s41598-021-82970-2",
"author": "Y Takahashi",
"doi-asserted-by": "publisher",
"first-page": "3379",
"journal-title": "Sci. Rep.",
"key": "958_CR134",
"unstructured": "Takahashi, Y. et al. Histone deacetylase inhibitors suppress ACE2 and ABO simultaneously, suggesting a preventive potential against COVID-19. Sci. Rep. 11, 3379 (2021).",
"volume": "11",
"year": "2021"
},
{
"DOI": "10.1126/science.abf4058",
"author": "KM White",
"doi-asserted-by": "publisher",
"first-page": "926",
"journal-title": "Science",
"key": "958_CR135",
"unstructured": "White, K. M. et al. Plitidepsin has potent preclinical efficacy against SARS-CoV-2 by targeting the host protein eEF1A. Science 371, 926–931 (2021).",
"volume": "371",
"year": "2021"
},
{
"DOI": "10.1016/j.chom.2020.05.020",
"author": "SH Sun",
"doi-asserted-by": "publisher",
"first-page": "124",
"journal-title": "Cell Host Microbe",
"key": "958_CR136",
"unstructured": "Sun, S. H. et al. A mouse model of SARS-CoV-2 infection and pathogenesis. Cell Host Microbe 28, 124–133 (2020).",
"volume": "28",
"year": "2020"
},
{
"DOI": "10.1101/2020.09.16.20190694",
"doi-asserted-by": "publisher",
"key": "958_CR137",
"unstructured": "Ichimura, T. et al. KIM-1/TIM-1 is a receptor for SARS-CoV-2 in lung and kidney. Preprint at medRxiv https://doi.org/10.1101/2020.09.16.20190694 (2020)."
},
{
"DOI": "10.1093/jmcb/mjab003",
"doi-asserted-by": "publisher",
"key": "958_CR138",
"unstructured": "Yang, C. et al. Kidney injury molecule-1 is a potential receptor for SARS-CoV-2. J. Mol. Cell Biol. https://doi.org/10.1093/jmcb/mjab003 (2021)."
},
{
"DOI": "10.1016/j.jbc.2021.100759",
"author": "AJ Carlos",
"doi-asserted-by": "publisher",
"first-page": "100759",
"journal-title": "J. Biol. Chem.",
"key": "958_CR139",
"unstructured": "Carlos, A. J. et al. The chaperone GRP78 is a host auxiliary factor for SARS-CoV- 2 and GRP78 depleting antibody blocks viral entry and infection. J. Biol. Chem. 296, 100759 (2021).",
"volume": "296",
"year": "2021"
},
{
"DOI": "10.1021/acscentsci.0c00855",
"author": "AN Baker",
"doi-asserted-by": "publisher",
"first-page": "2046",
"journal-title": "ACS Cent. Sci.",
"key": "958_CR140",
"unstructured": "Baker, A. N. et al. The SARS-COV-2 Spike protein binds sialic acids and enables rapid detection in a lateral flow point of care diagnostic device. ACS Cent. Sci. 6, 2046–2052 (2020).",
"volume": "6",
"year": "2020"
},
{
"DOI": "10.1128/mBio.02233-17",
"author": "F Xiao",
"doi-asserted-by": "publisher",
"first-page": "e02233",
"journal-title": "Mbio",
"key": "958_CR141",
"unstructured": "Xiao, F. et al. Interactions between the hepatitis C virus nonstructural 2 protein and host adaptor proteins 1 and 4 orchestrate virus release. Mbio 9, e02233–17 (2018).",
"volume": "9",
"year": "2018"
},
{
"DOI": "10.1091/mbc.e06-12-1147",
"author": "G Camus",
"doi-asserted-by": "publisher",
"first-page": "986",
"journal-title": "Mol. Biol. Cell",
"key": "958_CR142",
"unstructured": "Camus, G. et al. The clathrin adaptor complex AP-1 binds HIV-1 and MLV gag and facilitates their budding. Mol. Biol. Cell 18, 986–994 (2007).",
"volume": "18",
"year": "2007"
},
{
"DOI": "10.1371/journal.ppat.1002347",
"author": "M Caillet",
"doi-asserted-by": "publisher",
"first-page": "e1002347",
"journal-title": "PLoS Pathog.",
"key": "958_CR143",
"unstructured": "Caillet, M. et al. Rab7A is required for efficient production of infectious HIV-1. PLoS Pathog. 7, e1002347 (2011).",
"volume": "7",
"year": "2011"
},
{
"DOI": "10.1371/journal.ppat.1003278",
"author": "M Qi",
"doi-asserted-by": "publisher",
"first-page": "e1003278",
"journal-title": "PLoS Pathog.",
"key": "958_CR144",
"unstructured": "Qi, M. et al. Rab11-FIP1C and Rab14 direct plasma membrane sorting and particle incorporation of the HIV-1 envelope glycoprotein complex. PLoS Pathog. 9, e1003278 (2013).",
"volume": "9",
"year": "2013"
},
{
"DOI": "10.1371/journal.ppat.1006062",
"author": "M Mehedi",
"doi-asserted-by": "publisher",
"first-page": "e1006062",
"journal-title": "PLoS Pathog.",
"key": "958_CR145",
"unstructured": "Mehedi, M. et al. Actin-related protein 2 (ARP2) and virus-induced filopodia facilitate human respiratory syncytial virus spread. PLoS Pathog. 12, e1006062 (2016).",
"volume": "12",
"year": "2016"
},
{
"DOI": "10.1371/journal.ppat.1003911",
"author": "J Petersen",
"doi-asserted-by": "publisher",
"first-page": "e1003911",
"journal-title": "PLoS Pathog.",
"key": "958_CR146",
"unstructured": "Petersen, J. et al. The major cellular sterol regulatory pathway is required for Andes virus infection. PLoS Pathog. 10, e1003911 (2014).",
"volume": "10",
"year": "2014"
},
{
"DOI": "10.1038/s41467-018-08015-x",
"author": "S Yuan",
"doi-asserted-by": "publisher",
"first-page": "120",
"journal-title": "Nat. Commun.",
"key": "958_CR147",
"unstructured": "Yuan, S. et al. SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat. Commun. 10, 120 (2019).",
"volume": "10",
"year": "2019"
},
{
"DOI": "10.1038/s41598-017-03316-5",
"author": "S Wichit",
"doi-asserted-by": "publisher",
"first-page": "3145",
"journal-title": "Sci. Rep.",
"key": "958_CR148",
"unstructured": "Wichit, S. et al. Imipramine inhibits chikungunya virus replication in human skin fibroblasts through interference with intracellular cholesterol trafficking. Sci. Rep. 7, 3145 (2017).",
"volume": "7",
"year": "2017"
},
{
"DOI": "10.1128/JVI.03557-12",
"author": "M Friesland",
"doi-asserted-by": "publisher",
"first-page": "6377",
"journal-title": "J. Virol.",
"key": "958_CR149",
"unstructured": "Friesland, M., Mingorance, L., Chung, J., Chisari, F. V. & Gastaminza, P. Sigma-1 receptor regulates early steps of viral RNA replication at the onset of hepatitis C virus infection. J. Virol. 87, 6377–6390 (2013).",
"volume": "87",
"year": "2013"
},
{
"DOI": "10.1073/pnas.0911373107",
"author": "D Sir",
"doi-asserted-by": "publisher",
"first-page": "4383",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "958_CR150",
"unstructured": "Sir, D. et al. The early autophagic pathway is activated by hepatitis B virus and required for viral DNA replication. Proc. Natl Acad. Sci. USA 107, 4383–4388 (2010).",
"volume": "107",
"year": "2010"
},
{
"DOI": "10.1128/mBio.00597-17",
"author": "YC Lin",
"doi-asserted-by": "publisher",
"first-page": "e00597",
"journal-title": "Mbio",
"key": "958_CR151",
"unstructured": "Lin, Y. C., Jeng, K. S. & Lai, M. M. C. CNOT4-mediated ubiquitination of influenza A virus nucleoprotein promotes viral RNA replication. Mbio 8, e00597–17 (2017).",
"volume": "8",
"year": "2017"
},
{
"DOI": "10.1128/JVI.01006-12",
"author": "TE Oakland",
"doi-asserted-by": "publisher",
"first-page": "6625",
"journal-title": "J. Virol.",
"key": "958_CR152",
"unstructured": "Oakland, T. E., Haselton, K. J. & Randall, G. EWSR1 binds the hepatitis C virus cis-acting replication element and is required for efficient viral replication. J. Virol. 87, 6625–6634 (2013).",
"volume": "87",
"year": "2013"
},
{
"DOI": "10.1101/gr.230250.117",
"author": "HS Kim",
"doi-asserted-by": "publisher",
"first-page": "859",
"journal-title": "Genome Res.",
"key": "958_CR153",
"unstructured": "Kim, H. S. et al. Arrayed CRISPR screen with image-based assay reliably uncovers host genes required for coxsackievirus infection. Genome Res. 28, 859–868 (2018).",
"volume": "28",
"year": "2018"
},
{
"DOI": "10.7554/eLife.21470",
"author": "P Bagchi",
"doi-asserted-by": "publisher",
"first-page": "e21470",
"journal-title": "Elife",
"key": "958_CR154",
"unstructured": "Bagchi, P., Inoue, T. & Tsai, B. EMC1-dependent stabilization drives membrane penetration of a partially destabilized non-enveloped virus. Elife 5, e21470 (2016).",
"volume": "5",
"year": "2016"
},
{
"DOI": "10.1128/JVI.05314-11",
"author": "M Maric",
"doi-asserted-by": "publisher",
"first-page": "9667",
"journal-title": "J. Virol.",
"key": "958_CR155",
"unstructured": "Maric, M. et al. A functional role for torsina in herpes simplex virus 1 nuclear egress. J. Virol. 85, 9667–9679 (2011).",
"volume": "85",
"year": "2011"
},
{
"DOI": "10.3390/cells9030738",
"author": "JE Hölper",
"doi-asserted-by": "publisher",
"first-page": "738",
"journal-title": "Cells",
"key": "958_CR156",
"unstructured": "Hölper, J. E., Klupp, B. G., Luxton, G. W. G., Franzke, K. & Mettenleiter, T. C. Function of torsin AAA+ ATPases in pseudorabies virus nuclear egress. Cells 9, 738 (2020).",
"volume": "9",
"year": "2020"
},
{
"DOI": "10.1038/nrmicro.2017.29",
"author": "AS Puschnik",
"doi-asserted-by": "publisher",
"first-page": "351",
"journal-title": "Nat. Rev. Microbiol.",
"key": "958_CR157",
"unstructured": "Puschnik, A. S., Majzoub, K., Ooi, Y. S. & Carette, J. E. A CRISPR toolbox to study virus-host interactions. Nat. Rev. Microbiol. 15, 351–364 (2017).",
"volume": "15",
"year": "2017"
},
{
"DOI": "10.1038/s41467-019-13965-x",
"author": "B Li",
"doi-asserted-by": "publisher",
"first-page": "164",
"journal-title": "Nat. Commun.",
"key": "958_CR158",
"unstructured": "Li, B. et al. Genome-wide CRISPR screen identifies host dependency factors for influenza A virus infection. Nat. Commun. 11, 164 (2020).",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1016/j.celrep.2017.07.060",
"author": "BE Heaton",
"doi-asserted-by": "publisher",
"first-page": "1503",
"journal-title": "Cell Rep.",
"key": "958_CR159",
"unstructured": "Heaton, B. E. et al. A CRISPR activation screen identifies a pan-avian influenza virus inhibitory host factor. Cell Rep. 20, 1503–1512 (2017).",
"volume": "20",
"year": "2017"
},
{
"DOI": "10.1016/j.molcel.2006.05.012",
"author": "E Meylan",
"doi-asserted-by": "publisher",
"first-page": "561",
"journal-title": "Mol. Cell",
"key": "958_CR160",
"unstructured": "Meylan, E. & Tschopp, J. Toll-like receptors and RNA helicases: two parallel ways to trigger antiviral responses. Mol. Cell 22, 561–569 (2006).",
"volume": "22",
"year": "2006"
},
{
"DOI": "10.1016/j.immuni.2011.05.003",
"author": "Y-M Loo",
"doi-asserted-by": "publisher",
"first-page": "680",
"journal-title": "Immunity",
"key": "958_CR161",
"unstructured": "Loo, Y.-M. & Gale, M. Immune signaling by RIG-I-like receptors. Immunity 34, 680–692 (2011).",
"volume": "34",
"year": "2011"
},
{
"DOI": "10.1128/JVI.01918-14",
"author": "C Vazquez",
"doi-asserted-by": "publisher",
"first-page": "6974",
"journal-title": "J. Virol.",
"key": "958_CR162",
"unstructured": "Vazquez, C. & Horner, S. M. MAVS coordination of antiviral innate immunity. J. Virol. 89, 6974–7 (2015).",
"volume": "89",
"year": "2015"
},
{
"DOI": "10.1016/j.chom.2007.08.006",
"author": "JP White",
"doi-asserted-by": "publisher",
"first-page": "295",
"journal-title": "Cell Host Microbe",
"key": "958_CR163",
"unstructured": "White, J. P., Cardenas, A. M., Marissen, W. E. & Lloyd, R. E. Inhibition of cytoplasmic mRNA stress granule formation by a viral proteinase. Cell Host Microbe 2, 295–305 (2007).",
"volume": "2",
"year": "2007"
},
{
"DOI": "10.1038/s41467-020-20768-y",
"author": "S Lu",
"doi-asserted-by": "publisher",
"first-page": "502",
"journal-title": "Nat. Commun.",
"key": "958_CR164",
"unstructured": "Lu, S. et al. The SARS-CoV-2 nucleocapsid phosphoprotein forms mutually exclusive condensates with RNA and the membrane-associated M protein. Nat. Commun. 12, 502 (2021).",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1038/cr.2010.103",
"author": "X-Y Liu",
"doi-asserted-by": "publisher",
"first-page": "994",
"journal-title": "Cell Res.",
"key": "958_CR165",
"unstructured": "Liu, X.-Y., Wei, B., Shi, H.-X., Shan, Y.-F. & Wang, C. Tom70 mediates activation of interferon regulatory factor 3 on mitochondria. Cell Res. 20, 994–1011 (2010).",
"volume": "20",
"year": "2010"
},
{
"DOI": "10.1038/s41423-020-0514-8",
"author": "H Jiang",
"doi-asserted-by": "publisher",
"first-page": "998",
"journal-title": "Cell. Mol. Immunol.",
"key": "958_CR166",
"unstructured": "Jiang, H. et al. SARS-CoV-2 Orf9b suppresses type I interferon responses by targeting TOM70. Cell. Mol. Immunol. 17, 998–1000 (2020).",
"volume": "17",
"year": "2020"
},
{
"DOI": "10.1073/pnas.2016650117",
"author": "L Miorin",
"doi-asserted-by": "publisher",
"first-page": "28344",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "958_CR167",
"unstructured": "Miorin, L. et al. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Natl Acad. Sci. USA 117, 28344–28354 (2020).",
"volume": "117",
"year": "2020"
}
],
"reference-count": 167,
"references-count": 167,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.nature.com/articles/s41564-021-00958-0"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Cellular host factors for SARS-CoV-2 infection",
"type": "journal-article",
"update-policy": "https://doi.org/10.1007/springer_crossmark_policy",
"volume": "6"
}