Metal-coding assisted serological multi-omics profiling deciphers the role of selenium in COVID-19 immunity
et al., Chemical Science, doi:10.1039/d3sc03345g, Sep 2023
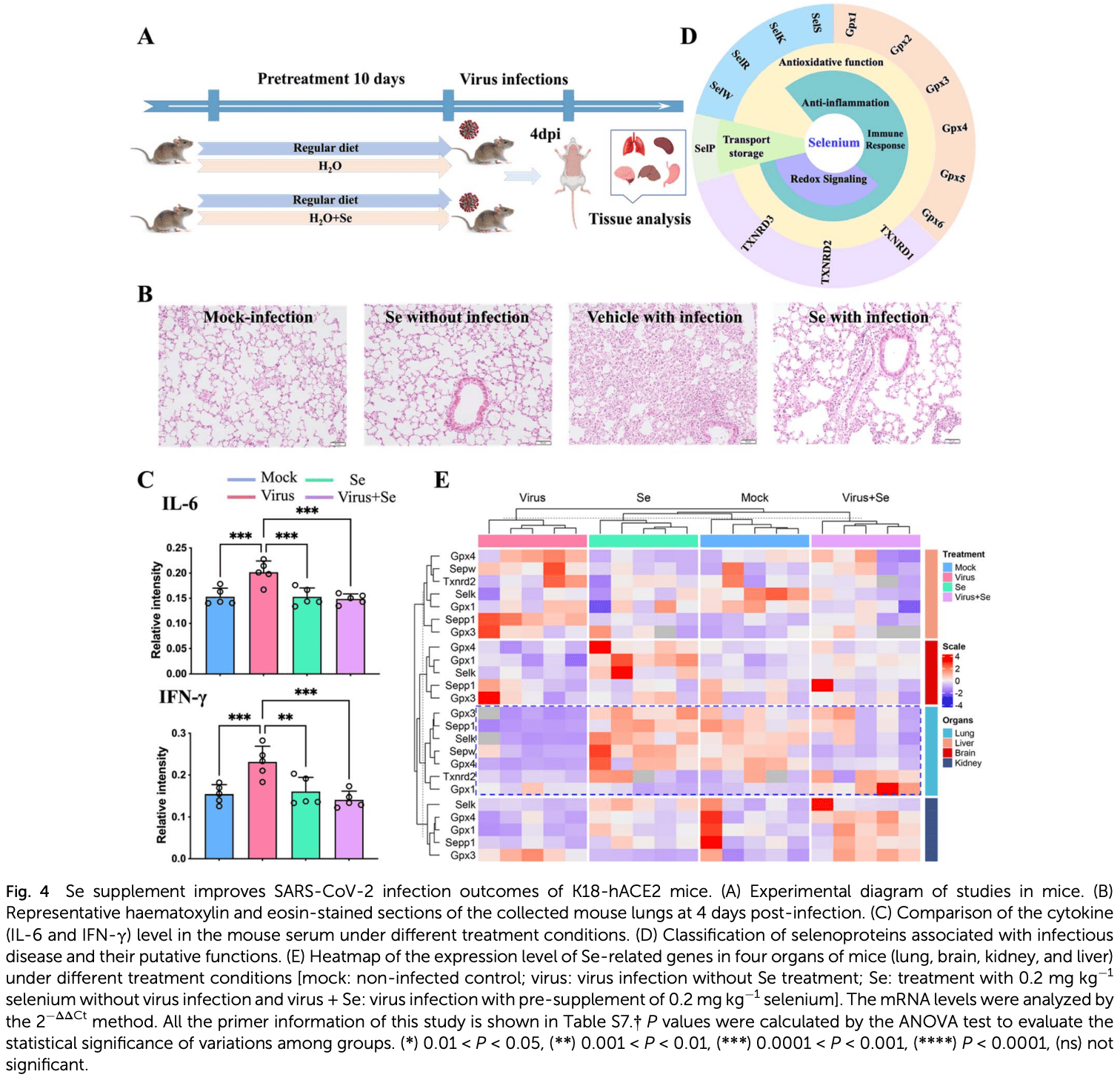
In vitro and mouse study highlighting selenium's potential for COVID-19 treatment through modulation of cytokines, selenoproteins, and immune cells. Using metal-coding mass spectrometry, authors link the COVID-19 metallome to disease severity and find selenium associates with anti-inflammatory interleukin-10. In vitro, selenium activates ERK/Akt pathways in B cells, enhancing interleukin-10 production while inhibiting proinflammatory cytokines. In SARS-CoV-2-infected mice, selenium restored virus-suppressed selenoproteins, alleviating lung damage through reduced cytokines.
Zhou et al., 18 Sep 2023, China, peer-reviewed, 20 authors.
Contact: hsun@hku.hk, jfwchan@hku.hk.
Metal-coding assisted serological multi-omics profiling deciphers the role of selenium in COVID-19 immunity
Chemical Science, doi:10.1039/d3sc03345g
Uncovering how host metal(loid)s mediate the immune response against invading pathogens is critical for better understanding the pathogenesis mechanism of infectious disease. Clinical data show that imbalance of host metal(loid)s is closely associated with the severity and mortality of COVID-19. However, it remains elusive how metal(loid)s, which are essential elements for all forms of life and closely associated with multiple diseases if dysregulated, are involved in COVID-19 pathophysiology and immunopathology. Herein, we built up a metal-coding assisted multiplexed serological metallome and immunoproteome profiling system to characterize the links of metallome with COVID-19 pathogenesis and immunity. We found distinct metallome features in COVID-19 patients compared with non-infected control subjects, which may serve as a biomarker for disease diagnosis. Moreover, we generated the first correlation network between the host metallome and immunity mediators, and unbiasedly uncovered a strong association of selenium with interleukin-10 (IL-10). Supplementation of selenium to immune cells resulted in enhanced IL-10 expression in B cells and reduced induction of proinflammatory cytokines in B and CD4 + T cells. The selenium-enhanced IL-10 production in B cells was confirmed to be attributable to the activation of ERK and Akt pathways. We further validated our cellular data in SARS-CoV-2-infected K18-hACE2 mice, and found that selenium supplementation alleviated SARS-CoV-2-induced lung damage characterized by decreased alveolar inflammatory infiltrates through restoration of virusrepressed selenoproteins to alleviate oxidative stress. Our approach can be readily extended to other diseases to understand how the host defends against invading pathogens through regulation of metallome.
Author contributions
Conflicts of interest There are no conicts to declare.
References
Al-Saleh, None, BioMetals
Amporndanai, None, Nat. Commun
Andreini, Arnesano, Rosato, None, Metallomics
Baixauli, Villa, Pearce, None, Science
Becht, None, Nat. Biotechnol
Bibby, None, Nat. Commun
Bojkova, None, Nature
Budinger, None, Lancet Neurol
Chan, None, Lancet
Domingo, Marquès, None, Food Chem. Toxicol
Eil, None, Nature
Fukada, None, Nat. Immunol
Ge, None, Nat. Rev. Cancer
Glass, None, Cell Rep
Gupta, None, Nat. Med
Hiffler, Rakotoambinina, None, Front. Nutr
Huang, None, Immunity
Iyer, Cheng, None, Crit. Rev. Immunol
Jesus, None, Metallomics
Karagiannis, None, Nature
Kim, None, Sci. Signaling
Krafcikova, None, Nat. Commun
Laforge, None, Nat. Rev. Immunol
Lee, None, Nature
Li, None, Chem. Commun
Maio, None, Science
Mckee, Fontenot, None, Curr. Opin. Immunol
Merad, None, Science
Michee-Cospolite, None, Front. Immunol
Moghaddam, None, Nutrients
Murdoch, Skaar, None, Nat. Rev. Microbiol
Namkoong, None, Nature
Noori, None, Rev. Med. Virol
Rayman, None, Lancet
Rayman, Taylor, Zhang, None, Proc. Nutr. Soc
Sargsyan, None, Chem. Sci
Schomburg, None, Nat. Rev. Endocrinol
Sedgwick, None, Br. Med. J
Skalny, None, Metabolites
Stukalov, None, Nature
To, None, Emerging Microbes Infect
Vig, Kinet, None, Nat. Immunol
Wang, None, Adv. Immunol
Wang, None, Food Chem. Toxicol
Yang, None, J. Med. Virol
Yao, None, Nat. Immunol
Yuan, None, Cell Discovery
Yuen, None, Int. J. Biol. Sci
Zhang, None, Am. J. Clin. Nutr
Zhang, None, Redox Biol
Zhou, None, Annu. Rev. Biochem
Zhou, None, Chem. Sci
Zhou, None, Nat. Biotechnol
DOI record:
{
"DOI": "10.1039/d3sc03345g",
"ISSN": [
"2041-6520",
"2041-6539"
],
"URL": "http://dx.doi.org/10.1039/d3sc03345g",
"abstract": "<jats:p>Uncovering how host metal(loid)s mediate the immune response against invading pathogens is critical for better understanding the pathogenesis mechanism of infectious disease.</jats:p>",
"author": [
{
"ORCID": "http://orcid.org/0009-0009-6634-3149",
"affiliation": [
{
"name": "Department of Chemistry, State Key Laboratory of Synthetic Chemistry, CAS-HKU Joint Laboratory of Metallomics on Health and Environment, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"authenticated-orcid": false,
"family": "Zhou",
"given": "Ying",
"sequence": "first"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
},
{
"name": "Department of Infectious Diseases and Microbiology, The University of Hong Kong-Shenzhen Hospital, Shenzhen, Guangdong, China"
},
{
"name": "Centre for Virology, Vaccinology and Therapeutics, Hong Kong Science and Technology Park, Hong Kong SAR, China"
}
],
"family": "Yuan",
"given": "Shuofeng",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Department of Pathology, Shenzhen Institute of Research and Innovation, The University of Hong Kong, Hong Kong SAR, China"
}
],
"family": "Xiao",
"given": "Fan",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Department of Chemistry, State Key Laboratory of Synthetic Chemistry, CAS-HKU Joint Laboratory of Metallomics on Health and Environment, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Li",
"given": "Hongyan",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "School of Biomedical Sciences, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Ye",
"given": "Ziwei",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Faculty of Dentistry, The University of Hong Kong, Pokfulam, Hong Kong SAR, Hong Kong, China"
}
],
"family": "Cheng",
"given": "Tianfan",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Luo",
"given": "Cuiting",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Tang",
"given": "Kaiming",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Cai",
"given": "Jianpiao",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Situ",
"given": "Jianwen",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
},
{
"name": "Department of Infectious Diseases and Microbiology, The University of Hong Kong-Shenzhen Hospital, Shenzhen, Guangdong, China"
},
{
"name": "Centre for Virology, Vaccinology and Therapeutics, Hong Kong Science and Technology Park, Hong Kong SAR, China"
},
{
"name": "Department of Microbiology, Queen Mary Hospital, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Sridhar",
"given": "Siddharth",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Division of Infectious Diseases, Department of Medicine, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Chu",
"given": "Wing-Ming",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Division of Infectious Diseases, Department of Medicine, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Tam",
"given": "Anthony Raymond",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Chu",
"given": "Hin",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-2554-7219",
"affiliation": [
{
"name": "Department of Chemistry, State Key Laboratory of Synthetic Chemistry, CAS-HKU Joint Laboratory of Metallomics on Health and Environment, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"authenticated-orcid": false,
"family": "Che",
"given": "Chi-Ming",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0002-5340-9710",
"affiliation": [
{
"name": "Faculty of Dentistry, The University of Hong Kong, Pokfulam, Hong Kong SAR, Hong Kong, China"
}
],
"authenticated-orcid": false,
"family": "Jin",
"given": "Lijian",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Department of Infectious Diseases and Microbiology, The University of Hong Kong-Shenzhen Hospital, Shenzhen, Guangdong, China"
},
{
"name": "Division of Infectious Diseases, Department of Medicine, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"family": "Hung",
"given": "Ivan Fan-Ngai",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Department of Pathology, Shenzhen Institute of Research and Innovation, The University of Hong Kong, Hong Kong SAR, China"
}
],
"family": "Lu",
"given": "Liwei",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0001-6336-6657",
"affiliation": [
{
"name": "State Key Laboratory of Emerging Infectious Diseases, Carol Yu Centre for Infection, Department of Microbiology, School of Clinical Medicine, Li Ka Shing Faculty of Medicine, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
},
{
"name": "Department of Infectious Diseases and Microbiology, The University of Hong Kong-Shenzhen Hospital, Shenzhen, Guangdong, China"
},
{
"name": "Centre for Virology, Vaccinology and Therapeutics, Hong Kong Science and Technology Park, Hong Kong SAR, China"
},
{
"name": "Department of Microbiology, Queen Mary Hospital, Pokfulam, Hong Kong SAR, China"
},
{
"name": "Academician Workstation of Hainan Province, Hainan Medical University-The University of Hong Kong Joint Laboratory of Tropical Infectious Diseases, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"authenticated-orcid": false,
"family": "Chan",
"given": "Jasper Fuk-Woo",
"sequence": "additional"
},
{
"ORCID": "http://orcid.org/0000-0001-6697-6899",
"affiliation": [
{
"name": "Department of Chemistry, State Key Laboratory of Synthetic Chemistry, CAS-HKU Joint Laboratory of Metallomics on Health and Environment, The University of Hong Kong, Pokfulam, Hong Kong SAR, China"
}
],
"authenticated-orcid": false,
"family": "Sun",
"given": "Hongzhe",
"sequence": "additional"
}
],
"container-title": "Chemical Science",
"container-title-short": "Chem. Sci.",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"rsc.org"
]
},
"created": {
"date-parts": [
[
2023,
9,
18
]
],
"date-time": "2023-09-18T11:58:01Z",
"timestamp": 1695038281000
},
"deposited": {
"date-parts": [
[
2023,
10,
4
]
],
"date-time": "2023-10-04T10:56:24Z",
"timestamp": 1696416984000
},
"funder": [
{
"DOI": "10.13039/501100002920",
"award": [
"C7060-21G",
"C7034-20EF",
"T11-709/21-N",
"17318322",
"2122-7S04",
"17306323"
],
"doi-asserted-by": "publisher",
"name": "Research Grants Council, University Grants Committee"
},
{
"DOI": "10.13039/501100005847",
"award": [
"20190572",
"COVID1903010"
],
"doi-asserted-by": "publisher",
"name": "Health and Medical Research Fund"
},
{
"DOI": "10.13039/501100005407",
"doi-asserted-by": "publisher",
"name": "Food and Health Bureau"
},
{
"DOI": "10.13039/501100003452",
"doi-asserted-by": "publisher",
"name": "Innovation and Technology Commission"
},
{
"DOI": "10.13039/501100012166",
"award": [
"2020YFA0707500",
"2020YFA0707504"
],
"doi-asserted-by": "publisher",
"name": "National Key Research and Development Program of China"
},
{
"DOI": "10.13039/501100012151",
"award": [
"SZSM201911014"
],
"doi-asserted-by": "publisher",
"name": "Sanming Project of Medicine in Shenzhen"
},
{
"DOI": "10.13039/501100018609",
"doi-asserted-by": "publisher",
"name": "High-level Hospital Construction Project of Guangdong Provincial People's Hospital"
},
{
"DOI": "10.13039/501100013072",
"award": [
"ZDKJ202003"
],
"doi-asserted-by": "publisher",
"name": "Major Science and Technology Project of Hainan Province"
},
{
"DOI": "10.13039/501100014818",
"doi-asserted-by": "publisher",
"name": "Hong Kong Hainan Commercial Association"
},
{
"DOI": "10.13039/100018831",
"doi-asserted-by": "publisher",
"name": "Lo Ying Shek Chi Wai Foundation"
},
{
"DOI": "10.13039/501100003803",
"doi-asserted-by": "publisher",
"name": "University of Hong Kong"
},
{
"DOI": "10.13039/501100001809",
"award": [
"General Program (82272337)"
],
"doi-asserted-by": "publisher",
"name": "National Natural Science Foundation of China"
}
],
"indexed": {
"date-parts": [
[
2023,
10,
5
]
],
"date-time": "2023-10-05T10:50:33Z",
"timestamp": 1696503033157
},
"is-referenced-by-count": 0,
"issue": "38",
"issued": {
"date-parts": [
[
2023
]
]
},
"journal-issue": {
"issue": "38",
"published-print": {
"date-parts": [
[
2023,
10,
4
]
]
}
},
"language": "en",
"license": [
{
"URL": "http://creativecommons.org/licenses/by-nc/3.0/",
"content-version": "vor",
"delay-in-days": 260,
"start": {
"date-parts": [
[
2023,
9,
18
]
],
"date-time": "2023-09-18T00:00:00Z",
"timestamp": 1694995200000
}
}
],
"link": [
{
"URL": "http://pubs.rsc.org/en/content/articlepdf/2023/SC/D3SC03345G",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "292",
"original-title": [],
"page": "10570-10579",
"prefix": "10.1039",
"published": {
"date-parts": [
[
2023
]
]
},
"published-online": {
"date-parts": [
[
2023
]
]
},
"publisher": "Royal Society of Chemistry (RSC)",
"reference": [
{
"DOI": "10.1016/S0140-6736(20)30154-9",
"author": "Chan",
"doi-asserted-by": "crossref",
"first-page": "514",
"journal-title": "Lancet",
"key": "D3SC03345G/cit1/1",
"volume": "395",
"year": "2020"
},
{
"DOI": "10.1080/22221751.2021.1898291",
"author": "To",
"doi-asserted-by": "crossref",
"first-page": "507",
"journal-title": "Emerging Microbes Infect.",
"key": "D3SC03345G/cit2/1",
"volume": "10",
"year": "2021"
},
{
"DOI": "10.1126/science.abm8108",
"author": "Merad",
"doi-asserted-by": "crossref",
"first-page": "1122",
"journal-title": "Science",
"key": "D3SC03345G/cit3/1",
"volume": "375",
"year": "2022"
},
{
"DOI": "10.1038/s41586-022-05163-5",
"author": "Namkoong",
"doi-asserted-by": "crossref",
"first-page": "754",
"journal-title": "Nature",
"key": "D3SC03345G/cit4/1",
"volume": "609",
"year": "2022"
},
{
"DOI": "10.1038/s41586-022-05128-8",
"author": "Karagiannis",
"doi-asserted-by": "crossref",
"first-page": "801",
"journal-title": "Nature",
"key": "D3SC03345G/cit5/1",
"volume": "609",
"year": "2022"
},
{
"author": "Yuan",
"first-page": "1",
"journal-title": "Cell Discovery",
"key": "D3SC03345G/cit6/1",
"volume": "7",
"year": "2021"
},
{
"DOI": "10.1038/s41586-021-03493-4",
"author": "Stukalov",
"doi-asserted-by": "crossref",
"first-page": "246",
"journal-title": "Nature",
"key": "D3SC03345G/cit7/1",
"volume": "594",
"year": "2021"
},
{
"DOI": "10.1038/s41586-020-2332-7",
"author": "Bojkova",
"doi-asserted-by": "crossref",
"first-page": "469",
"journal-title": "Nature",
"key": "D3SC03345G/cit8/1",
"volume": "583",
"year": "2020"
},
{
"author": "Zhou",
"first-page": "1",
"journal-title": "Nat. Biotechnol.",
"key": "D3SC03345G/cit9/1",
"volume": "374",
"year": "2022"
},
{
"DOI": "10.7150/ijbs.72709",
"author": "Yuen",
"doi-asserted-by": "crossref",
"first-page": "4714",
"journal-title": "Int. J. Biol. Sci.",
"key": "D3SC03345G/cit10/1",
"volume": "18",
"year": "2022"
},
{
"DOI": "10.1038/s41579-022-00745-6",
"author": "Murdoch",
"doi-asserted-by": "crossref",
"first-page": "657",
"journal-title": "Nat. Rev. Microbiol.",
"key": "D3SC03345G/cit11/1",
"volume": "20",
"year": "2022"
},
{
"DOI": "10.1146/annurev-biochem-040320-104628",
"author": "Zhou",
"doi-asserted-by": "crossref",
"first-page": "449",
"journal-title": "Annu. Rev. Biochem.",
"key": "D3SC03345G/cit12/1",
"volume": "91",
"year": "2022"
},
{
"DOI": "10.1016/S1474-4422(21)00238-6",
"author": "Budinger",
"doi-asserted-by": "crossref",
"first-page": "956",
"journal-title": "Lancet Neurol.",
"key": "D3SC03345G/cit13/1",
"volume": "20",
"year": "2021"
},
{
"DOI": "10.1038/s41568-021-00417-2",
"author": "Ge",
"doi-asserted-by": "crossref",
"first-page": "102",
"journal-title": "Nat. Rev. Cancer",
"key": "D3SC03345G/cit14/1",
"volume": "22",
"year": "2022"
},
{
"DOI": "10.1016/S0140-6736(11)61452-9",
"author": "Rayman",
"doi-asserted-by": "crossref",
"first-page": "1256",
"journal-title": "Lancet",
"key": "D3SC03345G/cit15/1",
"volume": "379",
"year": "2012"
},
{
"DOI": "10.1002/jmv.26515",
"author": "Yang",
"doi-asserted-by": "crossref",
"first-page": "1639",
"journal-title": "J. Med. Virol.",
"key": "D3SC03345G/cit16/1",
"volume": "93",
"year": "2021"
},
{
"DOI": "10.1038/nature19364",
"author": "Eil",
"doi-asserted-by": "crossref",
"first-page": "539",
"journal-title": "Nature",
"key": "D3SC03345G/cit17/1",
"volume": "537",
"year": "2016"
},
{
"DOI": "10.1038/nature11588",
"author": "Lee",
"doi-asserted-by": "crossref",
"first-page": "123",
"journal-title": "Nature",
"key": "D3SC03345G/cit18/1",
"volume": "492",
"year": "2012"
},
{
"DOI": "10.1126/scisignal.abi7400",
"author": "Kim",
"doi-asserted-by": "crossref",
"first-page": "eabi7400",
"journal-title": "Sci. Signaling",
"key": "D3SC03345G/cit19/1",
"volume": "15",
"year": "2022"
},
{
"DOI": "10.1016/bs.ai.2019.11.007",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "187",
"journal-title": "Adv. Immunol.",
"key": "D3SC03345G/cit20/1",
"volume": "145",
"year": "2020"
},
{
"DOI": "10.1016/j.coi.2016.05.001",
"author": "McKee",
"doi-asserted-by": "crossref",
"first-page": "25",
"journal-title": "Curr. Opin. Immunol.",
"key": "D3SC03345G/cit21/1",
"volume": "42",
"year": "2016"
},
{
"DOI": "10.1126/science.aaw8800",
"author": "Baixauli",
"doi-asserted-by": "crossref",
"first-page": "1395",
"journal-title": "Science",
"key": "D3SC03345G/cit22/1",
"volume": "363",
"year": "2019"
},
{
"DOI": "10.1038/s41590-019-0319-z",
"author": "Fukada",
"doi-asserted-by": "crossref",
"first-page": "248",
"journal-title": "Nat. Immunol.",
"key": "D3SC03345G/cit23/1",
"volume": "20",
"year": "2019"
},
{
"DOI": "10.1038/ni.f.220",
"author": "Vig",
"doi-asserted-by": "crossref",
"first-page": "21",
"journal-title": "Nat. Immunol.",
"key": "D3SC03345G/cit24/1",
"volume": "10",
"year": "2009"
},
{
"DOI": "10.1039/D2CC01772E",
"author": "Li",
"doi-asserted-by": "crossref",
"first-page": "7466",
"journal-title": "Chem. Commun.",
"key": "D3SC03345G/cit25/1",
"volume": "58",
"year": "2022"
},
{
"DOI": "10.1126/science.abi5224",
"author": "Maio",
"doi-asserted-by": "crossref",
"first-page": "236",
"journal-title": "Science",
"key": "D3SC03345G/cit26/1",
"volume": "373",
"year": "2021"
},
{
"DOI": "10.1093/mtomcs/mfac047",
"author": "Andreini",
"doi-asserted-by": "crossref",
"first-page": "mfac047",
"journal-title": "Metallomics",
"key": "D3SC03345G/cit27/1",
"volume": "14",
"year": "2022"
},
{
"DOI": "10.1038/s41467-020-17495-9",
"author": "Krafcikova",
"doi-asserted-by": "crossref",
"first-page": "3717",
"journal-title": "Nat. Commun.",
"key": "D3SC03345G/cit28/1",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.3390/metabo11040244",
"author": "Skalny",
"doi-asserted-by": "crossref",
"first-page": "244",
"journal-title": "Metabolites",
"key": "D3SC03345G/cit29/1",
"volume": "11",
"year": "2021"
},
{
"DOI": "10.3390/nu12072098",
"author": "Moghaddam",
"doi-asserted-by": "crossref",
"first-page": "2098",
"journal-title": "Nutrients",
"key": "D3SC03345G/cit30/1",
"volume": "12",
"year": "2020"
},
{
"DOI": "10.1016/j.fct.2021.112161",
"author": "Domingo",
"doi-asserted-by": "crossref",
"first-page": "112161",
"journal-title": "Food Chem. Toxicol.",
"key": "D3SC03345G/cit31/1",
"volume": "152",
"year": "2021"
},
{
"DOI": "10.1039/D1SC05852E",
"author": "Zhou",
"doi-asserted-by": "crossref",
"first-page": "3216",
"journal-title": "Chem. Sci.",
"key": "D3SC03345G/cit32/1",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.1038/nbt.4314",
"author": "Becht",
"doi-asserted-by": "crossref",
"first-page": "38",
"journal-title": "Nat. Biotechnol.",
"key": "D3SC03345G/cit33/1",
"volume": "37",
"year": "2019"
},
{
"DOI": "10.1136/bmj.g7327",
"author": "Sedgwick",
"doi-asserted-by": "crossref",
"first-page": "g7327",
"journal-title": "Br. Med. J.",
"key": "D3SC03345G/cit34/1",
"volume": "349",
"year": "2014"
},
{
"DOI": "10.1038/s41577-020-0407-1",
"author": "Laforge",
"doi-asserted-by": "crossref",
"first-page": "515",
"journal-title": "Nat. Rev. Immunol.",
"key": "D3SC03345G/cit35/1",
"volume": "20",
"year": "2020"
},
{
"DOI": "10.1016/j.celrep.2022.110728",
"author": "Glass",
"doi-asserted-by": "crossref",
"first-page": "110728",
"journal-title": "Cell Rep.",
"key": "D3SC03345G/cit36/1",
"volume": "39",
"year": "2022"
},
{
"DOI": "10.3389/fimmu.2022.818814",
"author": "Michee-Cospolite",
"doi-asserted-by": "crossref",
"first-page": "818814",
"journal-title": "Front. Immunol.",
"key": "D3SC03345G/cit37/1",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.1038/s41467-020-17179-4",
"author": "Bibby",
"doi-asserted-by": "crossref",
"first-page": "3412",
"journal-title": "Nat. Commun.",
"key": "D3SC03345G/cit38/1",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1038/s41590-021-00996-0",
"author": "Yao",
"doi-asserted-by": "crossref",
"first-page": "1127",
"journal-title": "Nat. Immunol.",
"key": "D3SC03345G/cit39/1",
"volume": "22",
"year": "2021"
},
{
"DOI": "10.1016/j.redox.2020.101715",
"author": "Zhang",
"doi-asserted-by": "crossref",
"first-page": "101715",
"journal-title": "Redox Biol.",
"key": "D3SC03345G/cit40/1",
"volume": "37",
"year": "2020"
},
{
"DOI": "10.1038/nrendo.2011.174",
"author": "Schomburg",
"doi-asserted-by": "crossref",
"first-page": "160",
"journal-title": "Nat. Rev. Endocrinol.",
"key": "D3SC03345G/cit41/1",
"volume": "8",
"year": "2012"
},
{
"DOI": "10.1039/d0mt00220h",
"author": "Jesus",
"doi-asserted-by": "crossref",
"first-page": "1912",
"journal-title": "Metallomics",
"key": "D3SC03345G/cit42/1",
"volume": "12",
"year": "2020"
},
{
"DOI": "10.1007/s10534-021-00355-4",
"author": "Al-Saleh",
"doi-asserted-by": "crossref",
"first-page": "125",
"journal-title": "BioMetals",
"key": "D3SC03345G/cit43/1",
"volume": "35",
"year": "2022"
},
{
"DOI": "10.1039/D0SC02646H",
"author": "Sargsyan",
"doi-asserted-by": "crossref",
"first-page": "9904",
"journal-title": "Chem. Sci.",
"key": "D3SC03345G/cit44/1",
"volume": "11",
"year": "2020"
},
{
"DOI": "10.1093/ajcn/nqaa095",
"author": "Zhang",
"doi-asserted-by": "crossref",
"first-page": "1297",
"journal-title": "Am. J. Clin. Nutr.",
"key": "D3SC03345G/cit45/1",
"volume": "111",
"year": "2020"
},
{
"DOI": "10.1038/s41467-021-23313-7",
"author": "Amporndanai",
"doi-asserted-by": "crossref",
"first-page": "3061",
"journal-title": "Nat. Commun.",
"key": "D3SC03345G/cit46/1",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1038/s41591-020-0968-3",
"author": "Gupta",
"doi-asserted-by": "crossref",
"first-page": "1017",
"journal-title": "Nat. Med.",
"key": "D3SC03345G/cit47/1",
"volume": "26",
"year": "2020"
},
{
"DOI": "10.1002/rmv.2262",
"author": "Noori",
"doi-asserted-by": "crossref",
"first-page": "e2262",
"journal-title": "Rev. Med. Virol.",
"key": "D3SC03345G/cit48/1",
"volume": "32",
"year": "2022"
},
{
"DOI": "10.1615/CritRevImmunol.v32.i1.30",
"author": "Iyer",
"doi-asserted-by": "crossref",
"first-page": "23",
"journal-title": "Crit. Rev. Immunol.",
"key": "D3SC03345G/cit49/1",
"volume": "32",
"year": "2012"
},
{
"DOI": "10.1017/S0029665122002646",
"author": "Rayman",
"doi-asserted-by": "crossref",
"first-page": "1",
"journal-title": "Proc. Nutr. Soc.",
"key": "D3SC03345G/cit50/1",
"volume": "82",
"year": "2023"
},
{
"DOI": "10.1016/j.immuni.2021.07.004",
"author": "Huang",
"doi-asserted-by": "crossref",
"first-page": "1728",
"journal-title": "Immunity",
"key": "D3SC03345G/cit51/1",
"volume": "54",
"year": "2021"
},
{
"DOI": "10.3389/fnut.2020.00164",
"author": "Hiffler",
"doi-asserted-by": "crossref",
"first-page": "164",
"journal-title": "Front. Nutr.",
"key": "D3SC03345G/cit52/1",
"volume": "7",
"year": "2020"
},
{
"DOI": "10.1016/j.fct.2021.112286",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "112286",
"journal-title": "Food Chem. Toxicol.",
"key": "D3SC03345G/cit53/1",
"volume": "153",
"year": "2021"
}
],
"reference-count": 53,
"references-count": 53,
"relation": {},
"resource": {
"primary": {
"URL": "http://xlink.rsc.org/?DOI=D3SC03345G"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"General Chemistry"
],
"subtitle": [],
"title": "Metal-coding assisted serological multi-omics profiling deciphers the role of selenium in COVID-19 immunity",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.1039/rsc_crossmark_policy",
"volume": "14"
}