Multimodal SARS-CoV-2 interactome sketches the virus-host spatial organization
et al., Communications Biology, doi:10.1038/s42003-025-07933-z, Mar 2025
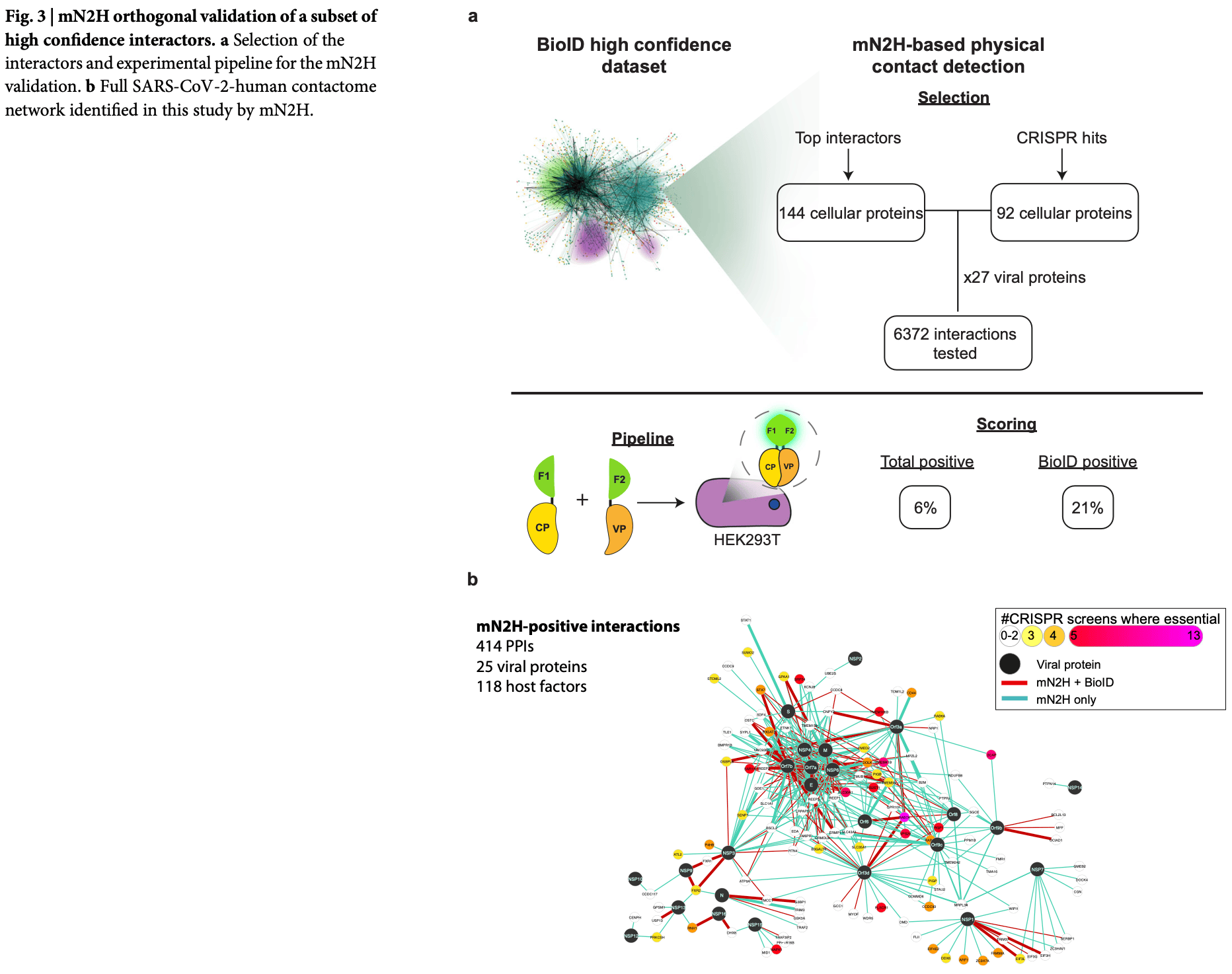
SARS-CoV-2-human interactome analysis in 3D space using Proximity-Dependent Biotin Identification (BioID) data. By analyzing 10,125 interactions between 27 viral proteins and 2,593 human proteins, authors created a spatially accurate model of the virus-host protein network, revealing how SARS-CoV-2 exploits cellular machinery during infection. Authors identified 414 high-confidence direct protein-protein interactions (PPIs) involving 25 viral proteins and 118 host factors, indicating a large number of potential therapeutic targets for SARS-CoV-2.
Dugied et al., 26 Mar 2025, multiple countries, peer-reviewed, 24 authors.
Contact: etienne.coyaud@inserm.fr, caroline.demeret@pasteur.fr.
Multimodal SARS-CoV-2 interactome sketches the virus-host spatial organization
Communications Biology, doi:10.1038/s42003-025-07933-z
An accurate spatial representation of protein-protein interaction networks is needed to achieve a realistic and biologically relevant representation of interactomes. Here, we leveraged the spatial information included in Proximity-Dependent Biotin Identification (BioID) interactomes of SARS-CoV-2 proteins to calculate weighted distances and model the organization of the SARS-CoV-2-human interactome in three dimensions (3D) within a cell-like volume. Cell regions with viral occupancy were highlighted, along with the coordination of viral proteins exploiting the cellular machinery. Profiling physical intra-virus and virus-host contacts enabled us to demonstrate both the accuracy and the predictive value of our 3D map for direct interactions, meaning that proteins in closer proximity tend to interact physically. Several functionally important virus-host complexes were detected, and robust structural models were obtained, opening the way to structure-directed drug discovery screens. This PPI discovery pipeline approach brings us closer to a realistic spatial representation of interactomes, which, when applied to viruses or other pathogens, can provide significant information for infection. Thus, it represents a promising tool for coping with emerging infectious diseases. Proteins are the main agents involved in the regulation of biological functions and don't act alone to mediate and regulate these functions; instead, they act in concert with other proteins through their interactions 1-5 . These protein-protein interactions (PPIs) are involved in the development of human diseases, either genetic, physiologic or infectious [6] [7] [8] . Therefore, mapping PPIs is essential to understanding their role in the development of diseases at the cellular level. Representation of networks of interactions is a critical component of the analysis, allowing the identification of proteins involved in similar complexes or pathways. However, PPI networks are often incomplete and hard to exploit, primarily due to the intrinsic nature of PPI detection methods and the two-dimensional network representations. Additionally, most interaction networks only represent a single study, performed with a single PPI detection method, while a complete network would require the aggregation of multiple studies. Moreover, distances within a protein interaction network are often empirical and without biological meaning, precluding the representation of actual volumes and regions. Contrasting with the huge increase in PPI detection methods sensitivity and deepness, little if any progress has been made toward the spatial representation of PPI networks. The tools developed to extrapolate distances from PPI networks dating back more than 10 years 9,10 . The inference of protein locations from interaction data to generate a spatial representation of interaction networks has so far remained out of reach. This step is yet a prerequisite towards an accurate three-dimensional representation..
Author contributions
Competing interests The authors declare no competing interests.
Additional information Supplementary information The online version contains supplementary material available at https://doi.org/10.1038/s42003-025-07933-z . Correspondence and requests for materials should be addressed to Yorgos Sofianatos, Etienne Coyaud or Caroline Demeret. Peer review information Communications Biology thanks Xinyu Cui and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Xiaohui Frank Zhang and Tobias Goris. Reprints and permissions information is available at http://www.nature.com/reprints Publisher's note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the..
References
Agapito, Guzzi, Cannataro, Visualization of protein interaction networks: problems and solutions, BMC Bioinformatics
Baggen, TMEM106B is a receptor mediating ACE2independent SARS-CoV-2 cell entry, Cell
Barnes, Hut, A hierarchical O(N log N) force-calculation algorithm, Nature
Basu, Wallner, DockQ: A Quality Measure for Protein-Protein Docking Models, PLoS One
Bessa, The intrinsically disordered SARS-CoV-2 nucleoprotein in dynamic complex with its viral partner nsp3a, Sci. Adv
Bryant, Pozzati, Elofsson, Improved prediction of proteinprotein interactions using AlphaFold2, Nat. Commun
Cheng, Comprehensive characterization of protein-protein interactions perturbed by disease mutations, Nat. Genet
Cho, Endogenous tagging for the cartography of human cellular organization, Science
Choi, Maximizing binary interactome mapping with a minimal number of assays, Nat. Commun
Ciccosanti, Proteomic analysis identifies the RNA helicase DDX3X as a host target against SARS-CoV-2 infection, Antivir. Res
Coutant, Gram-scale synthesis of luciferins derived from coelenterazine and original insights into their bioluminescence properties, Org. Biomol. Chem
Coyaud, Global Interactomics Uncovers Extensive Organellar Targeting by Zika Virus, Mol. Cell. Proteom
Dixon, NanoLuc Complementation Reporter Optimized for Accurate Measurement of Protein Interactions in Cells, ACS Chem. Biol
Drewes, Bantscheff, Chemical Proteomics: Methods and Protocols
Evans, Protein complex prediction with AlphaFold-Multimer, bioRxiv
Fruchterman, Reingold, Graph drawing by forcedirected placement, Softw. Pract. Exp
Gagnot, Core-Modified Coelenterazine Luciferin Analogues: Synthesis and Chemiluminescence Properties, Chemistry
Garvanska, The NSP3 protein of SARS-CoV-2 binds fragile X mental retardation proteins to disrupt UBAP2L interactions, EMBO Rep
Gonzalez, Kann, Chapter 4: Protein interactions and disease, PLoS Comput. Biol
Gordon, A SARS-CoV-2 protein interaction map reveals targets for drug repurposing, Nature
Guo, SARS-CoV-2 non-structural protein 13 (nsp13) hijacks host deubiquitinase USP13 and counteracts host antiviral immune response, Signal Transduct. Target Ther
Hagberg, Swart, Chult, Exploring Network Structure, Dynamics, and Function Using NetworkX
Halehalli, Nagarajaram, Molecular principles of human virus protein-protein interactions, Bioinformatics
Harris, Array programming with NumPy, Nature
Hoffmann, Functional interrogation of a SARS-CoV-2 host protein interactome identifies unique and shared coronavirus host factors, Cell Host Microbe
Hu, Efficient, high-quality force-directed graph drawing, Mathematica J
Inc, Collaborative data science
Jones, Thornton, Principles of protein-protein interactions, Proc. Natl Acad. Sci
Kim, A Comprehensive, Flexible Collection of SARS-CoV-2 Coding Regions, G
Kim, A proteome-scale map of the SARS-CoV-2-human contactome, Nat. Biotechnol, doi:10.1038/s41587-022-01475-z
Kim, Probing nuclear pore complex architecture with proximity-dependent biotinylation, Proc. Natl Acad. Sci. USA
Laurent, Global BioID-based SARS-CoV-2 proteins proximal interactome unveils novel ties between viral polypeptides and host factors involved in multiple COVID19-associated mechanisms, bioRxiv, doi:10.1101/2020.08.28.272955
Li, Atlas of interactions between SARS-CoV-2 macromolecules and host proteins, Cell Insight
Li, Liu, 3DProIN: Protein-Protein Interaction Networks and Structure Visualization, Am. J. Bioinform. Comput. Biol
Lin, Crystal structure of SARS-CoV-2 nsp10/nsp16 2'-Omethylase and its implication on antiviral drug design, Sig. Transduct. Target Ther
Link, Targeted Identification of Protein Interactions in Eukaryotic mRNA Translation, Proteomics
Liu, SARS-CoV-2-host proteome interactions for antiviral drug discovery, Mol. Syst. Biol
Mariano, Farthing, Lale-Farjat, Bergeron, Structural Characterization of SARS-CoV-2: Where We Are, and Where We Need to Be, Front Mol. Biosci
May, A BioID-Derived Proximity Interactome for SARS-CoV-2 Proteins, Viruses
Perez-Riverol, The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences, Nucleic Acids Res
Reback, None, Pandas-Dev/pandas: Pandas, doi:10.5281/zenodo.6408044
Samavarchi-Tehrani, A SARS-CoV-2host proximity interactome, doi:10.1101/2020.09.03.282103
Sevimoglu, Arga, The role of protein interaction networks in systems biomedicine, Comput. Struct. Biotechnol. J
Sheng, Integrated interactome and transcriptome analysis reveals key host factors critical for SARS-CoV-2 infection, Virol. Sin
Sowmya, Breen, Ranganathan, Linking structural features of protein complexes and biological function, Protein Sci
St-Germain, A SARS-CoV-2 BioID-based virus-host membrane protein interactome and virus peptide compendium: new proteomics resources for COVID-19 research, bioRxiv, doi:10.1101/2020.08.28.269175
Van Rossum, Python tutorial
Varadi, AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models, Nucleic Acids Res
Vincentelli, Quantifying domain-ligand affinities and specificities by high-throughput holdup assay, Nat. Methods
Walia, SARS-CoV-2 virulence factor ORF3a blocks lysosome function by modulating TBC1D5-dependent Rab7 GTPase cycle, Nat. Commun
Walshaw, A Multilevel Algorithm for Force-Directed Graph Drawing, doi:10.1007/3-540-44541-2_17
Zhou, A comprehensive SARS-CoV-2-human protein-protein interactome reveals COVID-19 pathobiology and potential host therapeutic targets, Nat. Biotechnol
DOI record:
{
"DOI": "10.1038/s42003-025-07933-z",
"ISSN": [
"2399-3642"
],
"URL": "http://dx.doi.org/10.1038/s42003-025-07933-z",
"abstract": "<jats:title>Abstract</jats:title>\n <jats:p>An accurate spatial representation of protein-protein interaction networks is needed to achieve a realistic and biologically relevant representation of interactomes. Here, we leveraged the spatial information included in Proximity-Dependent Biotin Identification (BioID) interactomes of SARS-CoV-2 proteins to calculate weighted distances and model the organization of the SARS-CoV-2-human interactome in three dimensions (3D) within a cell-like volume. Cell regions with viral occupancy were highlighted, along with the coordination of viral proteins exploiting the cellular machinery. Profiling physical intra-virus and virus-host contacts enabled us to demonstrate both the accuracy and the predictive value of our 3D map for direct interactions, meaning that proteins in closer proximity tend to interact physically. Several functionally important virus-host complexes were detected, and robust structural models were obtained, opening the way to structure-directed drug discovery screens. This PPI discovery pipeline approach brings us closer to a realistic spatial representation of interactomes, which, when applied to viruses or other pathogens, can provide significant information for infection. Thus, it represents a promising tool for coping with emerging infectious diseases.</jats:p>",
"alternative-id": [
"7933"
],
"article-number": "501",
"assertion": [
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Received",
"name": "received",
"order": 1,
"value": "18 September 2024"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "12 March 2025"
},
{
"group": {
"label": "Article History",
"name": "ArticleHistory"
},
"label": "First Online",
"name": "first_online",
"order": 3,
"value": "26 March 2025"
},
{
"group": {
"label": "Competing interests",
"name": "EthicsHeading"
},
"name": "Ethics",
"order": 1,
"value": "The authors declare no competing interests."
}
],
"author": [
{
"affiliation": [],
"family": "Dugied",
"given": "Guillaume",
"sequence": "first"
},
{
"affiliation": [],
"family": "Laurent",
"given": "Estelle MN",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Attia",
"given": "Mikaël",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Gimeno",
"given": "Jean-Pascal",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Bachiri",
"given": "Kamel",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Samavarchi-Tehrani",
"given": "Payman",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0003-2862-9751",
"affiliation": [],
"authenticated-orcid": false,
"family": "Donati",
"given": "Flora",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Rahou",
"given": "Yannis",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Munier",
"given": "Sandie",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Amara",
"given": "Faustine",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Dos Santos",
"given": "Mélanie",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Soler",
"given": "Nicolas",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Volant",
"given": "Stevenn",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Pietrosemoli",
"given": "Natalia",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-6090-4437",
"affiliation": [],
"authenticated-orcid": false,
"family": "Gingras",
"given": "Anne-Claude",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-4577-8276",
"affiliation": [],
"authenticated-orcid": false,
"family": "Pavlopoulos",
"given": "Georgios A.",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-1148-4456",
"affiliation": [],
"authenticated-orcid": false,
"family": "van der Werf",
"given": "Sylvie",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0003-2012-6746",
"affiliation": [],
"authenticated-orcid": false,
"family": "Falter-Braun",
"given": "Pascal",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Aloy",
"given": "Patrick",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Jacob",
"given": "Yves",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Komarova",
"given": "Anastassia",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0009-0001-1109-0211",
"affiliation": [],
"authenticated-orcid": false,
"family": "Sofianatos",
"given": "Yorgos",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-5893-4557",
"affiliation": [],
"authenticated-orcid": false,
"family": "Coyaud",
"given": "Etienne",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-1103-6805",
"affiliation": [],
"authenticated-orcid": false,
"family": "Demeret",
"given": "Caroline",
"sequence": "additional"
}
],
"container-title": "Communications Biology",
"container-title-short": "Commun Biol",
"content-domain": {
"crossmark-restriction": false,
"domain": [
"link.springer.com"
]
},
"created": {
"date-parts": [
[
2025,
3,
26
]
],
"date-time": "2025-03-26T19:51:51Z",
"timestamp": 1743018711000
},
"deposited": {
"date-parts": [
[
2025,
3,
26
]
],
"date-time": "2025-03-26T19:54:55Z",
"timestamp": 1743018895000
},
"funder": [
{
"DOI": "10.13039/100010661",
"award": [
"101003589"
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/100010661",
"id-type": "DOI"
}
],
"name": "EC | Horizon 2020 Framework Programme"
},
{
"DOI": "10.13039/100000009",
"award": [
"R21AI153738"
],
"doi-asserted-by": "publisher",
"id": [
{
"asserted-by": "publisher",
"id": "10.13039/100000009",
"id-type": "DOI"
}
],
"name": "Foundation for the National Institutes of Health"
}
],
"indexed": {
"date-parts": [
[
2025,
3,
26
]
],
"date-time": "2025-03-26T20:10:28Z",
"timestamp": 1743019828491,
"version": "3.40.3"
},
"is-referenced-by-count": 0,
"issue": "1",
"issued": {
"date-parts": [
[
2025,
3,
26
]
]
},
"journal-issue": {
"issue": "1",
"published-online": {
"date-parts": [
[
2025,
12
]
]
}
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2025,
3,
26
]
],
"date-time": "2025-03-26T00:00:00Z",
"timestamp": 1742947200000
}
},
{
"URL": "https://creativecommons.org/licenses/by/4.0",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2025,
3,
26
]
],
"date-time": "2025-03-26T00:00:00Z",
"timestamp": 1742947200000
}
}
],
"link": [
{
"URL": "https://www.nature.com/articles/s42003-025-07933-z.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.nature.com/articles/s42003-025-07933-z",
"content-type": "text/html",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://www.nature.com/articles/s42003-025-07933-z.pdf",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "297",
"original-title": [],
"prefix": "10.1038",
"published": {
"date-parts": [
[
2025,
3,
26
]
]
},
"published-online": {
"date-parts": [
[
2025,
3,
26
]
]
},
"publisher": "Springer Science and Business Media LLC",
"reference": [
{
"DOI": "10.1371/journal.pcbi.1002819",
"author": "M.W. Gonzalez",
"doi-asserted-by": "crossref",
"first-page": "e1002819",
"journal-title": "PLoS Comput. Biol.",
"key": "7933_CR1",
"unstructured": "Gonzalez, M. W. & Kann, M. G. Chapter 4: Protein interactions and disease. PLoS Comput. Biol. 8, e1002819 (2012).",
"volume": "8",
"year": "2012"
},
{
"DOI": "10.1073/pnas.93.1.13",
"author": "S. Jones",
"doi-asserted-by": "crossref",
"first-page": "13",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "7933_CR2",
"unstructured": "Jones, S. & Thornton, J.M. Principles of protein-protein interactions. Proc. Natl Acad. Sci. USA 93, 13–20 (1996).",
"volume": "93",
"year": "1996"
},
{
"DOI": "10.1002/pro.2736",
"author": "G. Sowmya",
"doi-asserted-by": "crossref",
"first-page": "1486",
"journal-title": "Protein Sci.",
"key": "7933_CR3",
"unstructured": "Sowmya, G., Breen, E. J. & Ranganathan, S. Linking structural features of protein complexes and biological function. Protein Sci. 24, 1486–1494 (2015).",
"volume": "24",
"year": "2015"
},
{
"DOI": "10.1007/978-1-61779-364-6",
"doi-asserted-by": "crossref",
"key": "7933_CR4",
"unstructured": "Drewes, G. & Bantscheff, M. Chemical Proteomics: Methods and Protocols (Humana Press, 2011)."
},
{
"author": "A.J. Link",
"journal-title": "Proteomics",
"key": "7933_CR5",
"unstructured": "Link, A. J. et al. Targeted Identification of Protein Interactions in Eukaryotic mRNA Translation. Proteomics 20, e1900177 (2020).",
"volume": "20",
"year": "2020"
},
{
"DOI": "10.1038/s41588-020-00774-y",
"author": "F. Cheng",
"doi-asserted-by": "crossref",
"first-page": "342",
"journal-title": "Nat. Genet.",
"key": "7933_CR6",
"unstructured": "Cheng, F. et al. Comprehensive characterization of protein–protein interactions perturbed by disease mutations. Nat. Genet. 53, 342–353 (2021).",
"volume": "53",
"year": "2021"
},
{
"DOI": "10.1093/bioinformatics/btu763",
"author": "R.R. Halehalli",
"doi-asserted-by": "crossref",
"first-page": "1025",
"journal-title": "Bioinformatics",
"key": "7933_CR7",
"unstructured": "Halehalli, R. R. & Nagarajaram, H. A. Molecular principles of human virus protein-protein interactions. Bioinformatics 31, 1025–1033 (2015).",
"volume": "31",
"year": "2015"
},
{
"DOI": "10.1016/j.csbj.2014.08.008",
"author": "T. Sevimoglu",
"doi-asserted-by": "crossref",
"first-page": "22",
"journal-title": "Comput. Struct. Biotechnol. J.",
"key": "7933_CR8",
"unstructured": "Sevimoglu, T. & Arga, K. Y. The role of protein interaction networks in systems biomedicine. Comput. Struct. Biotechnol. J. 11, 22–27 (2014).",
"volume": "11",
"year": "2014"
},
{
"author": "H. Li",
"first-page": "32",
"journal-title": "Am. J. Bioinform. Comput. Biol.",
"key": "7933_CR9",
"unstructured": "Li, H. & Liu, C. 3DProIN: Protein-Protein Interaction Networks and Structure Visualization. Am. J. Bioinform. Comput. Biol. 2, 32–37 (2014).",
"volume": "2",
"year": "2014"
},
{
"DOI": "10.1186/1471-2105-14-S1-S1",
"doi-asserted-by": "crossref",
"key": "7933_CR10",
"unstructured": "Agapito, G., Guzzi, P. H. & Cannataro, M. Visualization of protein interaction networks: problems and solutions. BMC Bioinformatics 14, S1 (2013)."
},
{
"DOI": "10.1038/324446a0",
"author": "J. Barnes",
"doi-asserted-by": "crossref",
"first-page": "446",
"journal-title": "Nature",
"key": "7933_CR11",
"unstructured": "Barnes, J. & Hut, P. A hierarchical O(N log N) force-calculation algorithm. Nature 324, 446–449 (1986).",
"volume": "324",
"year": "1986"
},
{
"DOI": "10.1002/spe.4380211102",
"author": "T.M.J. Fruchterman",
"doi-asserted-by": "crossref",
"first-page": "1129",
"journal-title": "Softw. Pract. Exp.",
"key": "7933_CR12",
"unstructured": "Fruchterman, T. M. J. & Reingold, E. M. Graph drawing by force-directed placement. Softw. Pract. Exp. 21, 1129–1164 (1991).",
"volume": "21",
"year": "1991"
},
{
"DOI": "10.1007/3-540-44541-2_17",
"doi-asserted-by": "publisher",
"key": "7933_CR13",
"unstructured": "Walshaw, C. A Multilevel Algorithm for Force-Directed Graph Drawing. In: Graph Drawing 171–182 (Springer Berlin Heidelberg, 2001). https://doi.org/10.1007/3-540-44541-2_17."
},
{
"DOI": "10.1101/2020.08.28.269175",
"doi-asserted-by": "publisher",
"key": "7933_CR14",
"unstructured": "St-Germain, J. R. et al. A SARS-CoV-2 BioID-based virus-host membrane protein interactome and virus peptide compendium: new proteomics resources for COVID-19 research. bioRxiv, https://doi.org/10.1101/2020.08.28.269175 (2020)."
},
{
"DOI": "10.1038/s41587-022-01475-z",
"doi-asserted-by": "publisher",
"key": "7933_CR15",
"unstructured": "Kim, D.-K. et al. A proteome-scale map of the SARS-CoV-2–human contactome. Nat. Biotechnol. 1–10, https://doi.org/10.1038/s41587-022-01475-z (2022)."
},
{
"DOI": "10.1038/s41586-020-2286-9",
"author": "D.E. Gordon",
"doi-asserted-by": "crossref",
"first-page": "459",
"journal-title": "Nature",
"key": "7933_CR16",
"unstructured": "Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).",
"volume": "583",
"year": "2020"
},
{
"DOI": "10.1038/s41587-022-01474-0",
"author": "Y. Zhou",
"doi-asserted-by": "crossref",
"first-page": "128",
"journal-title": "Nat. Biotechnol.",
"key": "7933_CR17",
"unstructured": "Zhou, Y. et al. A comprehensive SARS-CoV-2-human protein-protein interactome reveals COVID-19 pathobiology and potential host therapeutic targets. Nat. Biotechnol. 41, 128–139 (2023).",
"volume": "41",
"year": "2023"
},
{
"DOI": "10.1101/2020.09.03.282103",
"author": "P. Samavarchi-Tehrani",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "7933_CR18",
"unstructured": "Samavarchi-Tehrani, P. et al. A SARS-CoV-2 – host proximity interactome. bioRxiv https://doi.org/10.1101/2020.09.03.282103 (2020).",
"year": "2020"
},
{
"DOI": "10.3390/v14030611",
"doi-asserted-by": "crossref",
"key": "7933_CR19",
"unstructured": "May, D. et al. A BioID-Derived Proximity Interactome for SARS-CoV-2 Proteins. Viruses 14, 611 (2022)."
},
{
"author": "X. Liu",
"journal-title": "Mol. Syst. Biol.",
"key": "7933_CR20",
"unstructured": "Liu, X. et al. SARS-CoV-2-host proteome interactions for antiviral drug discovery. Mol. Syst. Biol. 17, e10396 (2021).",
"volume": "17",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2023.06.005",
"author": "J. Baggen",
"doi-asserted-by": "crossref",
"first-page": "3427",
"journal-title": "Cell",
"key": "7933_CR21",
"unstructured": "Baggen, J. et al. TMEM106B is a receptor mediating ACE2-independent SARS-CoV-2 cell entry. Cell 186, 3427–3442.e22 (2023).",
"volume": "186",
"year": "2023"
},
{
"DOI": "10.1016/j.antiviral.2021.105064",
"author": "F. Ciccosanti",
"doi-asserted-by": "crossref",
"first-page": "105064",
"journal-title": "Antivir. Res.",
"key": "7933_CR22",
"unstructured": "Ciccosanti, F. et al. Proteomic analysis identifies the RNA helicase DDX3X as a host target against SARS-CoV-2 infection. Antivir. Res. 190, 105064 (2021).",
"volume": "190",
"year": "2021"
},
{
"DOI": "10.1038/s44319-023-00043-z",
"author": "D.H. Garvanska",
"doi-asserted-by": "crossref",
"first-page": "902",
"journal-title": "EMBO Rep.",
"key": "7933_CR23",
"unstructured": "Garvanska, D. H. et al. The NSP3 protein of SARS-CoV-2 binds fragile X mental retardation proteins to disrupt UBAP2L interactions. EMBO Rep. 25, 902–926 (2024).",
"volume": "25",
"year": "2024"
},
{
"DOI": "10.1038/s41467-024-46417-2",
"author": "K. Walia",
"doi-asserted-by": "crossref",
"journal-title": "Nat. Commun.",
"key": "7933_CR24",
"unstructured": "Walia, K. et al. SARS-CoV-2 virulence factor ORF3a blocks lysosome function by modulating TBC1D5-dependent Rab7 GTPase cycle. Nat. Commun. 15 2053 (2024).",
"volume": "15",
"year": "2024"
},
{
"DOI": "10.1073/pnas.1315147111",
"author": "D.I. Kim",
"doi-asserted-by": "crossref",
"first-page": "E2453",
"journal-title": "Proc. Natl Acad. Sci. USA",
"key": "7933_CR25",
"unstructured": "Kim, D. I. et al. Probing nuclear pore complex architecture with proximity-dependent biotinylation. Proc. Natl Acad. Sci. USA 111, E2453–E2461 (2014).",
"volume": "111",
"year": "2014"
},
{
"DOI": "10.1101/2020.08.28.272955",
"doi-asserted-by": "publisher",
"key": "7933_CR26",
"unstructured": "Laurent, E. M. N. et al. Global BioID-based SARS-CoV-2 proteins proximal interactome unveils novel ties between viral polypeptides and host factors involved in multiple COVID19-associated mechanisms. bioRxiv, https://doi.org/10.1101/2020.08.28.272955 (2020)."
},
{
"DOI": "10.1016/j.chom.2020.12.009",
"author": "H.-H. Hoffmann",
"doi-asserted-by": "crossref",
"first-page": "267",
"journal-title": "Cell Host Microbe",
"key": "7933_CR27",
"unstructured": "Hoffmann, H.-H. et al. Functional interrogation of a SARS-CoV-2 host protein interactome identifies unique and shared coronavirus host factors. Cell Host Microbe 29, 267–280.e5 (2021).",
"volume": "29",
"year": "2021"
},
{
"author": "G. Li",
"journal-title": "Cell Insight",
"key": "7933_CR28",
"unstructured": "Li, G. et al. Atlas of interactions between SARS-CoV-2 macromolecules and host proteins. Cell Insight 2, 100068 (2023).",
"volume": "2",
"year": "2023"
},
{
"DOI": "10.1016/j.virs.2023.05.004",
"author": "J. Sheng",
"doi-asserted-by": "crossref",
"first-page": "508",
"journal-title": "Virol. Sin.",
"key": "7933_CR29",
"unstructured": "Sheng, J. et al. Integrated interactome and transcriptome analysis reveals key host factors critical for SARS-CoV-2 infection. Virol. Sin. 38, 508–519 (2023).",
"volume": "38",
"year": "2023"
},
{
"DOI": "10.3389/fmolb.2020.605236",
"author": "G. Mariano",
"doi-asserted-by": "crossref",
"first-page": "605236",
"journal-title": "Front Mol. Biosci.",
"key": "7933_CR30",
"unstructured": "Mariano, G., Farthing, R. J., Lale-Farjat, S. L. M. & Bergeron, J. R. C. Structural Characterization of SARS-CoV-2: Where We Are, and Where We Need to Be. Front Mol. Biosci. 7, 605236 (2020).",
"volume": "7",
"year": "2020"
},
{
"author": "S.G. Choi",
"journal-title": "Nat. Commun.",
"key": "7933_CR31",
"unstructured": "Choi, S. G. et al. Maximizing binary interactome mapping with a minimal number of assays. Nat. Commun. 10, 3907 (2019).",
"volume": "10",
"year": "2019"
},
{
"DOI": "10.1038/nmeth.3438",
"author": "R. Vincentelli",
"doi-asserted-by": "crossref",
"first-page": "787",
"journal-title": "Nat. Methods",
"key": "7933_CR32",
"unstructured": "Vincentelli, R. et al. Quantifying domain-ligand affinities and specificities by high-throughput holdup assay. Nat. Methods 12, 787–793 (2015).",
"volume": "12",
"year": "2015"
},
{
"DOI": "10.1126/sciadv.abm4034",
"author": "L.M. Bessa",
"doi-asserted-by": "crossref",
"journal-title": "Sci. Adv.",
"key": "7933_CR33",
"unstructured": "Bessa, L. M. et al. The intrinsically disordered SARS-CoV-2 nucleoprotein in dynamic complex with its viral partner nsp3a. Sci. Adv. 8, eabm4034 (2022).",
"volume": "8",
"year": "2022"
},
{
"DOI": "10.1038/s41392-020-00241-4",
"author": "S. Lin",
"doi-asserted-by": "crossref",
"first-page": "131",
"journal-title": "Sig. Transduct. Target Ther.",
"key": "7933_CR34",
"unstructured": "Lin, S. et al. Crystal structure of SARS-CoV-2 nsp10/nsp16 2’-O-methylase and its implication on antiviral drug design. Sig. Transduct. Target Ther. 5, 131 (2020).",
"volume": "5",
"year": "2020"
},
{
"author": "R. Evans",
"first-page": "463034",
"journal-title": "bioRxiv 2021",
"key": "7933_CR35",
"unstructured": "Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. bioRxiv 2021 10, 463034 (2022).",
"volume": "10",
"year": "2022"
},
{
"DOI": "10.1371/journal.pone.0161879",
"author": "S. Basu",
"doi-asserted-by": "crossref",
"first-page": "e0161879",
"journal-title": "PLoS One",
"key": "7933_CR36",
"unstructured": "Basu, S. & Wallner, B. DockQ: A Quality Measure for Protein-Protein Docking Models. PLoS One 11, e0161879 (2016).",
"volume": "11",
"year": "2016"
},
{
"DOI": "10.1093/nar/gkab1061",
"author": "M. Varadi",
"doi-asserted-by": "crossref",
"first-page": "D439",
"journal-title": "Nucleic Acids Res.",
"key": "7933_CR37",
"unstructured": "Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).",
"volume": "50",
"year": "2022"
},
{
"DOI": "10.1038/s41392-021-00509-3",
"author": "G. Guo",
"doi-asserted-by": "crossref",
"first-page": "119",
"journal-title": "Signal Transduct. Target Ther.",
"key": "7933_CR38",
"unstructured": "Guo, G. et al. SARS-CoV-2 non-structural protein 13 (nsp13) hijacks host deubiquitinase USP13 and counteracts host antiviral immune response. Signal Transduct. Target Ther. 6, 119 (2021).",
"volume": "6",
"year": "2021"
},
{
"DOI": "10.1126/science.abi6983",
"author": "N.H. Cho",
"doi-asserted-by": "crossref",
"first-page": "eabi6983",
"journal-title": "Science",
"key": "7933_CR39",
"unstructured": "Cho, N. H. et al. OpenCell: Endogenous tagging for the cartography of human cellular organization. Science 375, eabi6983 (2022).",
"volume": "375",
"year": "2022"
},
{
"DOI": "10.1074/mcp.TIR118.000800",
"author": "E. Coyaud",
"doi-asserted-by": "crossref",
"first-page": "2242",
"journal-title": "Mol. Cell. Proteom.",
"key": "7933_CR40",
"unstructured": "Coyaud, E. et al. Global Interactomics Uncovers Extensive Organellar Targeting by Zika Virus. Mol. Cell. Proteom. 17, 2242–2255 (2018).",
"volume": "17",
"year": "2018"
},
{
"DOI": "10.1534/g3.120.401554",
"author": "D.-K. Kim",
"doi-asserted-by": "crossref",
"first-page": "3399",
"journal-title": "G3",
"key": "7933_CR41",
"unstructured": "Kim, D.-K. et al. A Comprehensive, Flexible Collection of SARS-CoV-2 Coding Regions. G3 10, 3399–3402 (2020).",
"volume": "10",
"year": "2020"
},
{
"DOI": "10.1093/nar/gkab1038",
"author": "Y. Perez-Riverol",
"doi-asserted-by": "crossref",
"first-page": "D543",
"journal-title": "Nucleic Acids Res.",
"key": "7933_CR42",
"unstructured": "Perez-Riverol, Y. et al. The PRIDE database resources in 2022: a hub for mass spectrometry-based proteomics evidences. Nucleic Acids Res. 50, D543–D552 (2022).",
"volume": "50",
"year": "2022"
},
{
"author": "Y. Hu",
"first-page": "37",
"journal-title": "Mathematica J.",
"key": "7933_CR43",
"unstructured": "Hu, Y. Efficient, high-quality force-directed graph drawing. Mathematica J. 10, 37–71 (2005).",
"volume": "10",
"year": "2005"
},
{
"key": "7933_CR44",
"unstructured": "Van Rossum, G. Python tutorial, Technical Report CS-R9526, Centrum voor Wiskunde en Informatica (CWI), Amsterdam, May 1995."
},
{
"DOI": "10.25080/TCWV9851",
"doi-asserted-by": "crossref",
"key": "7933_CR45",
"unstructured": "Hagberg, A., Swart, P. & Chult, D. S. Exploring Network Structure, Dynamics, and Function Using NetworkX. https://www.osti.gov/biblio/960616 (2008)."
},
{
"DOI": "10.1038/s41586-020-2649-2",
"author": "C.R. Harris",
"doi-asserted-by": "crossref",
"first-page": "357",
"journal-title": "Nature",
"key": "7933_CR46",
"unstructured": "Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).",
"volume": "585",
"year": "2020"
},
{
"DOI": "10.5281/zenodo.6408044",
"doi-asserted-by": "publisher",
"key": "7933_CR47",
"unstructured": "Reback, J. et al. Pandas-Dev/pandas: Pandas 1 2, https://doi.org/10.5281/zenodo.6408044 (2022)."
},
{
"key": "7933_CR48",
"unstructured": "Inc, P. T. Collaborative data science (Montreal: Plotly Technologies Inc Montral, 2015)."
},
{
"DOI": "10.1002/chem.202004311",
"author": "G. Gagnot",
"doi-asserted-by": "crossref",
"first-page": "2112",
"journal-title": "Chemistry",
"key": "7933_CR49",
"unstructured": "Gagnot, G. et al. Core-Modified Coelenterazine Luciferin Analogues: Synthesis and Chemiluminescence Properties. Chemistry 27, 2112–2123 (2021).",
"volume": "27",
"year": "2021"
},
{
"DOI": "10.1039/C9OB00459A",
"author": "E.P. Coutant",
"doi-asserted-by": "crossref",
"first-page": "3709",
"journal-title": "Org. Biomol. Chem.",
"key": "7933_CR50",
"unstructured": "Coutant, E. P. et al. Gram-scale synthesis of luciferins derived from coelenterazine and original insights into their bioluminescence properties. Org. Biomol. Chem. 17, 3709–3713 (2019).",
"volume": "17",
"year": "2019"
},
{
"author": "P. Bryant",
"journal-title": "Nat. Commun.",
"key": "7933_CR51",
"unstructured": "Bryant, P., Pozzati, G. & Elofsson, A. Improved prediction of protein-protein interactions using AlphaFold2. Nat. Commun. 13, 1265 (2022).",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.1021/acschembio.5b00753",
"author": "A.S. Dixon",
"doi-asserted-by": "crossref",
"first-page": "400",
"journal-title": "ACS Chem. Biol.",
"key": "7933_CR52",
"unstructured": "Dixon, A. S. et al. NanoLuc Complementation Reporter Optimized for Accurate Measurement of Protein Interactions in Cells. ACS Chem. Biol. 11, 400–408 (2016).",
"volume": "11",
"year": "2016"
}
],
"reference-count": 52,
"references-count": 52,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.nature.com/articles/s42003-025-07933-z"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "Multimodal SARS-CoV-2 interactome sketches the virus-host spatial organization",
"type": "journal-article",
"update-policy": "https://doi.org/10.1007/springer_crossmark_policy",
"volume": "8"
}