SARS-CoV-2 3CLpro mutations T21I and E166A confer differential resistance to simnotrelvir, bofutrelvir, and ensitrelvir
et al., Journal of Virology, doi:10.1128/jvi.02223-25, Apr 2026
In vitro and mouse study showing differential resistance of SARS-CoV-2 3CLpro T21I/E166A mutations to simnotrelvir, bofutrelvir, nirmatrelvir, and ensitrelvir.
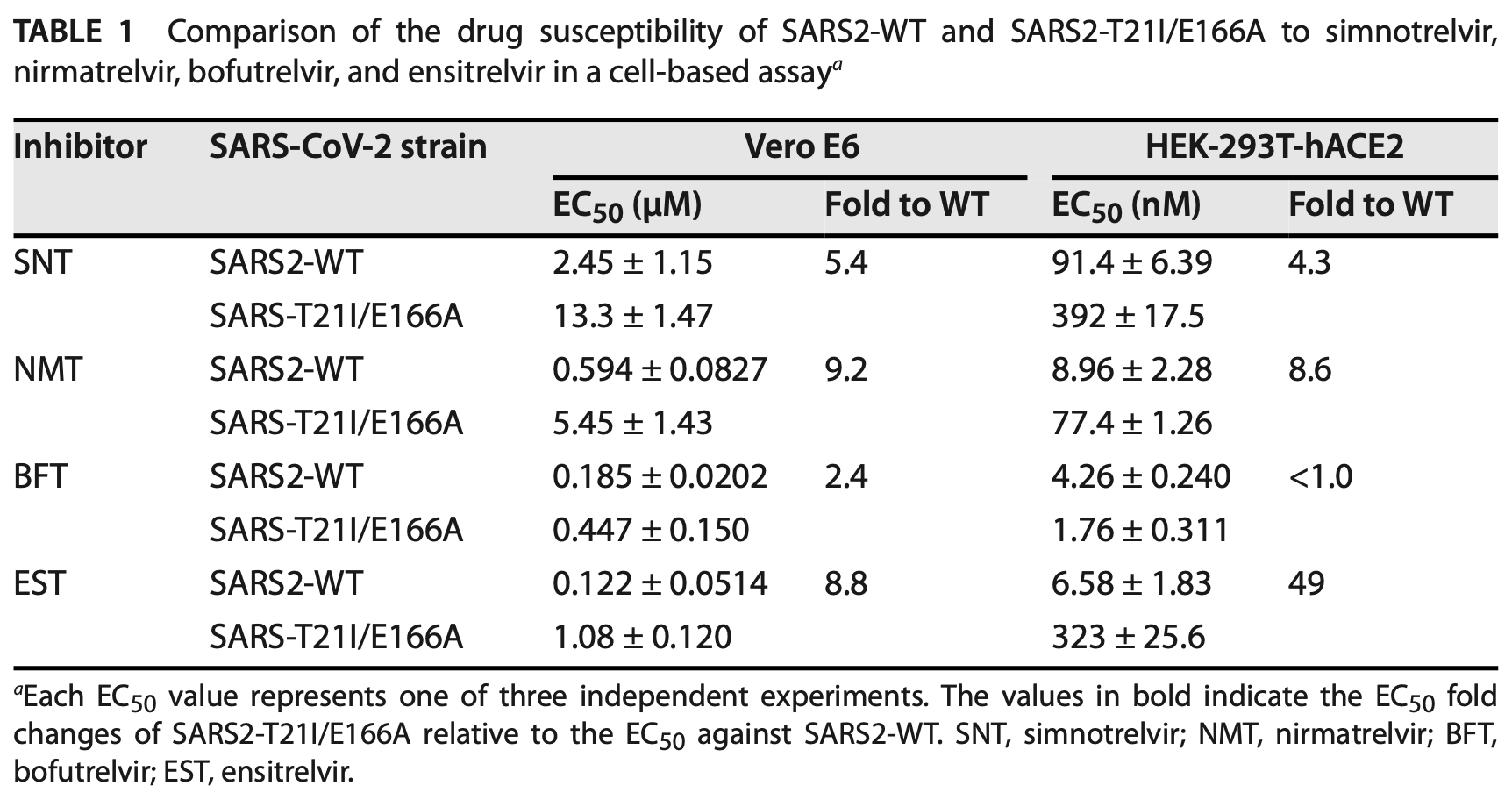
The resistant isolate SARS2-T21I/E166A showed 4.3-9.2-fold resistance to simnotrelvir and nirmatrelvir, and 8.8-49-fold resistance to ensitrelvir in cell-based assays, but maintained sensitivity to bofutrelvir (≤2.4-fold EC50 increase).
Chen et al., 27 Apr 2026, multiple countries, peer-reviewed, 11 authors.
Contact: liuwen@ctgu.edu.cn, ycxu@simm.ac.cn, ymzhang@wh.iov.cn.
Abstract:
| Molecular and Cellular Biology | Full-Length Text
SARS-CoV-2 3CLpro mutations T21I and E166A confer differential resistance to simnotrelvir, bofutrelvir, and ensitrelvir
Lu Chen, 1,2 Haixia Su, 2,3 Weijuan Shang, 1 Tianqing Nie, 3 Wenhua Kuang, 1 Chenchen Wang, 2,3,4 Qiang Shao, 2,3 Leike Zhang, 1,2,5 Wen Liu, 6 Yechun Xu, 2,3,7 Yumin Zhang 1
AUTHOR AFFILIATIONS See affiliation list on p. 18.
ABSTRACT Inhibiting the catalytic activity of 3CLpro is a mainstream strategy to block coronavirus replication. However, the appearance of SARS-CoV-2 3CLpro resistance to protease inhibitors raises concerns for effective therapies. In this work, we first investiga ted the resistance profile of simnotrelvir, an approved anti-SARS-CoV-2 drug that targets 3CLpro. We found that the T21I/E166A mutations in 3CLpro equally emerged when SARS-CoV-2 was passaged in the HEK293T-hACE2 cells with increasing concentrations of simnotrelvir. The SARS-CoV-2 isolate carrying 3CLpro T21I/E166A (SARS2-T21I/E166A) showed cross-resistance to simnotrelvir, nirmatrelvir, and ensitrelvir, but not significant resist ance to bofutrelvir. Biochemical and cellular assays confirmed that 3CLpro T21I/E166A was associated with the differential resistance to these protease inhibitors. Crystallographic structural analysis indicated that the alanine substitution disrupted hydrogen bonding interactions surrounding the γ-lactam rings (P1) of the inhibitors, which is similar to the model rebuilding observed with the previously reported E166V mutation. However, in contrast to the valine substitution, the alanine substitution resulted in a more spacious S2 subsite, thereby causing stronger interaction between the P1 and residues F140 and Ser1 of protomer B. Further computational simulations demonstrated that the covalent binding of bofutrelvir preserves strong binding affinity despite modifications in the S2 subsite caused by the E166A mutation, suggesting that inhibitors containing an aldehyde warhead may partially overcome resistance. Notably, both simnotrelvir and bofutrelvir exhibited therapeutic efficacy against the SARS2-T21I/E166A variant in K18-hACE2 mice. These findings advance our understanding of the resistance profiles and mechanistic underpinnings of SARS-CoV-2 3CLpro and underscore the necessity for diversified antiviral therapeutic strategies.
IMPORTANCE Considering that the nirmatrelvir-resistant SARS-CoV-2 has emerged in immunocompromised patients who received long-term Paxlovid therapy, it is essential to investigate the response of resistance 3CLpro mutants to various protease inhibi tors. Simnotrelvir, a novel inhibitor targeting SARS-CoV-2 3CLpro, has been authorized for the treatment of mild-to-moderate COVID-19 in China and has treated over 1 million patients. However, the resistance profile of simnotrelvir to SARS-CoV-2 remains unknown. Here, we identified that 3CLpro with T21I/E166A mutations confers resistance to simnotrelvir and showed cross-resistance to nirmatrelvir and ensitrelvir, but not bofutrelvir. More importantly, we further revealed that E166A showed a novel resistance mechanism to both the covalent inhibitors consisting of a γ-lactam ring and non-cova lent inhibitors like ensitrelvir, which is different from that of E166V previously reported. In contrast, bofutrelvir maintains high..
DOI record:
{
"DOI": "10.1128/jvi.02223-25",
"ISSN": [
"0022-538X",
"1098-5514"
],
"URL": "http://dx.doi.org/10.1128/jvi.02223-25",
"abstract": "<jats:title>ABSTRACT</jats:title>\n <jats:sec>\n <jats:title/>\n <jats:p>\n Inhibiting the catalytic activity of 3CLpro is a mainstream strategy to block coronavirus replication. However, the appearance of SARS-CoV-2 3CLpro resistance to protease inhibitors raises concerns for effective therapies. In this work, we first investigated the resistance profile of simnotrelvir, an approved anti-SARS-CoV-2 drug that targets 3CLpro. We found that the T21I/E166A mutations in 3CLpro equally emerged when SARS-CoV-2 was passaged in the HEK293T-hACE2 cells with increasing concentrations of simnotrelvir. The SARS-CoV-2 isolate carrying 3CLpro\n <jats:sup>T21I/E166A</jats:sup>\n (SARS2-T21I/E166A) showed cross-resistance to simnotrelvir, nirmatrelvir, and ensitrelvir, but not significant resistance to bofutrelvir. Biochemical and cellular assays confirmed that 3CLpro\n <jats:sup>T21I/E166A</jats:sup>\n was associated with the differential resistance to these protease inhibitors. Crystallographic structural analysis indicated that the alanine substitution disrupted hydrogen bonding interactions surrounding the γ-lactam rings (P1) of the inhibitors, which is similar to the model rebuilding observed with the previously reported E166V mutation. However, in contrast to the valine substitution, the alanine substitution resulted in a more spacious S2 subsite, thereby causing stronger interaction between the P1 and residues F140 and Ser1 of protomer B. Further computational simulations demonstrated that the covalent binding of bofutrelvir preserves strong binding affinity despite modifications in the S2 subsite caused by the E166A mutation, suggesting that inhibitors containing an aldehyde warhead may partially overcome resistance. Notably, both simnotrelvir and bofutrelvir exhibited therapeutic efficacy against the SARS2-T21I/E166A variant in K18-hACE2 mice. These findings advance our understanding of the resistance profiles and mechanistic underpinnings of SARS-CoV-2 3CLpro and underscore the necessity for diversified antiviral therapeutic strategies.\n </jats:p>\n </jats:sec>\n <jats:sec>\n <jats:title>IMPORTANCE</jats:title>\n <jats:p>Considering that the nirmatrelvir-resistant SARS-CoV-2 has emerged in immunocompromised patients who received long-term Paxlovid therapy, it is essential to investigate the response of resistance 3CLpro mutants to various protease inhibitors. Simnotrelvir, a novel inhibitor targeting SARS-CoV-2 3CLpro, has been authorized for the treatment of mild-to-moderate COVID-19 in China and has treated over 1 million patients. However, the resistance profile of simnotrelvir to SARS-CoV-2 remains unknown. Here, we identified that 3CLpro with T21I/E166A mutations confers resistance to simnotrelvir and showed cross-resistance to nirmatrelvir and ensitrelvir, but not bofutrelvir. More importantly, we further revealed that E166A showed a novel resistance mechanism to both the covalent inhibitors consisting of a γ-lactam ring and non-covalent inhibitors like ensitrelvir, which is different from that of E166V previously reported. In contrast, bofutrelvir maintains high affinity to T21I/E166A, suggesting that inhibitors with aldehyde warhead can partly neutralize the resistance.</jats:p>\n </jats:sec>",
"alternative-id": [
"10.1128/jvi.02223-25"
],
"article-number": "e02223-25",
"assertion": [
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Received",
"name": "received",
"order": 0,
"value": "2025-12-30"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Accepted",
"name": "accepted",
"order": 2,
"value": "2026-04-03"
},
{
"group": {
"label": "Publication History",
"name": "publication_history"
},
"label": "Published",
"name": "published",
"order": 3,
"value": "2026-04-27"
}
],
"author": [
{
"ORCID": "https://orcid.org/0009-0002-1540-4625",
"affiliation": [
{
"name": "Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences",
"place": [
"Wuhan, China"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
}
],
"authenticated-orcid": false,
"family": "Chen",
"given": "Lu",
"sequence": "first"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
},
{
"name": "State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences",
"place": [
"Shanghai, China"
]
}
],
"family": "Su",
"given": "Haixia",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences",
"place": [
"Wuhan, China"
]
}
],
"family": "Shang",
"given": "Weijuan",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences",
"place": [
"Shanghai, China"
]
}
],
"family": "Nie",
"given": "Tianqing",
"sequence": "additional"
},
{
"affiliation": [
{
"name": "Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences",
"place": [
"Wuhan, China"
]
}
],
"family": "Kuang",
"given": "Wenhua",
"sequence": "additional"
},
{
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
},
{
"name": "State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences",
"place": [
"Shanghai, China"
]
},
{
"name": "Microbiome Research Group, Research Centre for Life Science and Healthcare, Nottingham Ningbo China Beacons of Excellence Research and Innovation Institute (CBI), University of Nottingham Ningbo China",
"place": [
"Ningbo, China"
]
}
],
"family": "Wang",
"given": "Chenchen",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0001-6460-3095",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
},
{
"name": "State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences",
"place": [
"Shanghai, China"
]
}
],
"authenticated-orcid": true,
"family": "Shao",
"given": "Qiang",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-2593-2571",
"affiliation": [
{
"name": "Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences",
"place": [
"Wuhan, China"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
},
{
"name": "Hubei Jiangxia Laboratory",
"place": [
"Wuhan, China"
]
}
],
"authenticated-orcid": true,
"family": "Zhang",
"given": "Leike",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0003-2448-2107",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/0419nfc77",
"id-type": "ROR"
}
],
"name": "College of Biological and Pharmaceutical Sciences, China Three Gorges University",
"place": [
"Yichang, China"
]
}
],
"authenticated-orcid": false,
"family": "Liu",
"given": "Wen",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0002-1581-6155",
"affiliation": [
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/05qbk4x57",
"id-type": "ROR"
}
],
"name": "University of Chinese Academy of Sciences",
"place": [
"Beijing, China"
]
},
{
"name": "State Key Laboratory of Drug Research, Shanghai Institute of Materia Medica, Chinese Academy of Sciences",
"place": [
"Shanghai, China"
]
},
{
"id": [
{
"asserted-by": "publisher",
"id": "https://ror.org/00f809463",
"id-type": "ROR"
}
],
"name": "School of Pharmaceutical Science and Technology, Hangzhou Institute for Advanced Study, University of Chinese Academy of Sciences",
"place": [
"Hangzhou, China"
]
}
],
"authenticated-orcid": false,
"family": "Xu",
"given": "Yechun",
"sequence": "additional"
},
{
"ORCID": "https://orcid.org/0000-0003-0767-1221",
"affiliation": [
{
"name": "Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences",
"place": [
"Wuhan, China"
]
}
],
"authenticated-orcid": true,
"family": "Zhang",
"given": "Yumin",
"sequence": "additional"
}
],
"container-title": "Journal of Virology",
"container-title-short": "J Virol",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"journals.asm.org"
]
},
"created": {
"date-parts": [
[
2026,
4,
27
]
],
"date-time": "2026-04-27T09:03:08Z",
"timestamp": 1777280588000
},
"deposited": {
"date-parts": [
[
2026,
4,
27
]
],
"date-time": "2026-04-27T09:03:29Z",
"timestamp": 1777280609000
},
"editor": [
{
"affiliation": [],
"family": "Gallagher",
"given": "Tom",
"sequence": "additional"
}
],
"indexed": {
"date-parts": [
[
2026,
4,
27
]
],
"date-time": "2026-04-27T09:46:53Z",
"timestamp": 1777283213113,
"version": "3.51.4"
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2026,
4,
27
]
]
},
"language": "en",
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
4,
27
]
],
"date-time": "2026-04-27T00:00:00Z",
"timestamp": 1777248000000
}
},
{
"URL": "https://journals.asm.org/non-commercial-tdm-license",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2026,
4,
27
]
],
"date-time": "2026-04-27T00:00:00Z",
"timestamp": 1777248000000
}
}
],
"link": [
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/jvi.02223-25",
"content-type": "application/pdf",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://journals.asm.org/doi/pdf/10.1128/jvi.02223-25",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "235",
"original-title": [],
"prefix": "10.1128",
"published": {
"date-parts": [
[
2026,
4,
27
]
]
},
"published-online": {
"date-parts": [
[
2026,
4,
27
]
]
},
"publisher": "American Society for Microbiology",
"reference": [
{
"DOI": "10.1038/s41579-022-00841-7",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_2_2"
},
{
"DOI": "10.1056/NEJMoa2204919",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_3_2"
},
{
"DOI": "10.1126/scitranslmed.abq4064",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_4_2"
},
{
"DOI": "10.1038/s41467-023-42102-yB1",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_5_2"
},
{
"DOI": "10.1038/s41573-023-00672-y",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_6_2"
},
{
"DOI": "10.1002/jmv.26670",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_7_2"
},
{
"DOI": "10.3389/fphar.2021.718763",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_8_2"
},
{
"DOI": "10.1016/j.coviro.2012.08.008",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_9_2"
},
{
"DOI": "10.1128/mbio.02815-22",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_10_2"
},
{
"DOI": "10.1126/sciadv.adl4013",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_11_2"
},
{
"DOI": "10.1371/journal.ppat.1011592",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_12_2"
},
{
"DOI": "10.1016/j.chom.2022.08.003",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_13_2"
},
{
"DOI": "10.1126/sciadv.add7197",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_14_2"
},
{
"DOI": "10.1038/s41586-023-06609-0",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_15_2"
},
{
"DOI": "10.1021/acscentsci.3c00538",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_16_2"
},
{
"DOI": "10.1128/mbio.00869-22",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_17_2"
},
{
"DOI": "10.1126/sciadv.ade8778",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_18_2"
},
{
"DOI": "10.1126/scitranslmed.abq7360",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_19_2"
},
{
"DOI": "10.1038/s44259-023-00009-0",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_20_2"
},
{
"DOI": "10.1056/NEJMoa2301425",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_21_2"
},
{
"DOI": "10.1016/j.medj.2023.08.001",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_22_2"
},
{
"DOI": "10.1038/s41586-022-05514-2",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_23_2"
},
{
"DOI": "10.1073/pnas.2404175121",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_24_2"
},
{
"DOI": "10.1038/s41467-024-51924-3",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_25_2"
},
{
"DOI": "10.1038/s41392-023-01587-1",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_26_2"
},
{
"DOI": "10.1038/s41467-023-39704-x",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_27_2"
},
{
"DOI": "10.1038/s41467-023-40018-1",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_28_2"
},
{
"DOI": "10.1016/j.ejps.2023.106598",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_29_2"
},
{
"DOI": "10.1021/acs.jmedchem.2c00404",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_30_2"
},
{
"DOI": "10.1107/S1600576724002188",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_31_2"
},
{
"DOI": "10.1107/S0907444909047337",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_32_2"
},
{
"DOI": "10.1107/S0907444913000061",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_33_2"
},
{
"DOI": "10.1107/S0907444994003112",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_34_2"
},
{
"DOI": "10.1038/s41401-020-0483-6",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_35_2"
},
{
"DOI": "10.1107/S0907444904019158",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_36_2"
},
{
"DOI": "10.1107/s0907444902016657",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_37_2"
},
{
"DOI": "10.1126/science.abl4784",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_38_2"
},
{
"DOI": "10.1126/science.abb4489",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_39_2"
},
{
"DOI": "10.1021/acs.jcim.3c01531",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_40_2"
},
{
"DOI": "10.1021/acs.jctc.5b00255",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_41_2"
},
{
"DOI": "10.1002/jcc.20035",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_42_2"
},
{
"DOI": "10.1021/j100142a004",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_43_2"
},
{
"DOI": "10.1063/1.464397",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_44_2"
},
{
"DOI": "10.1002/jcc.20857",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_45_2"
},
{
"DOI": "10.1021/jp070071l",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_46_2"
},
{
"DOI": "10.1016/s0959-440x(00)00194-9",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_47_2"
},
{
"DOI": "10.1002/jcc.540130812",
"doi-asserted-by": "publisher",
"key": "e_1_3_5_48_2"
}
],
"reference-count": 47,
"references-count": 47,
"relation": {},
"resource": {
"primary": {
"URL": "https://journals.asm.org/doi/10.1128/jvi.02223-25"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [],
"subtitle": [],
"title": "SARS-CoV-2 3CLpro mutations T21I and E166A confer differential resistance to simnotrelvir, bofutrelvir, and ensitrelvir",
"type": "journal-article",
"update-policy": "https://doi.org/10.1128/asmj-crossmark-policy-page"
}
chen51