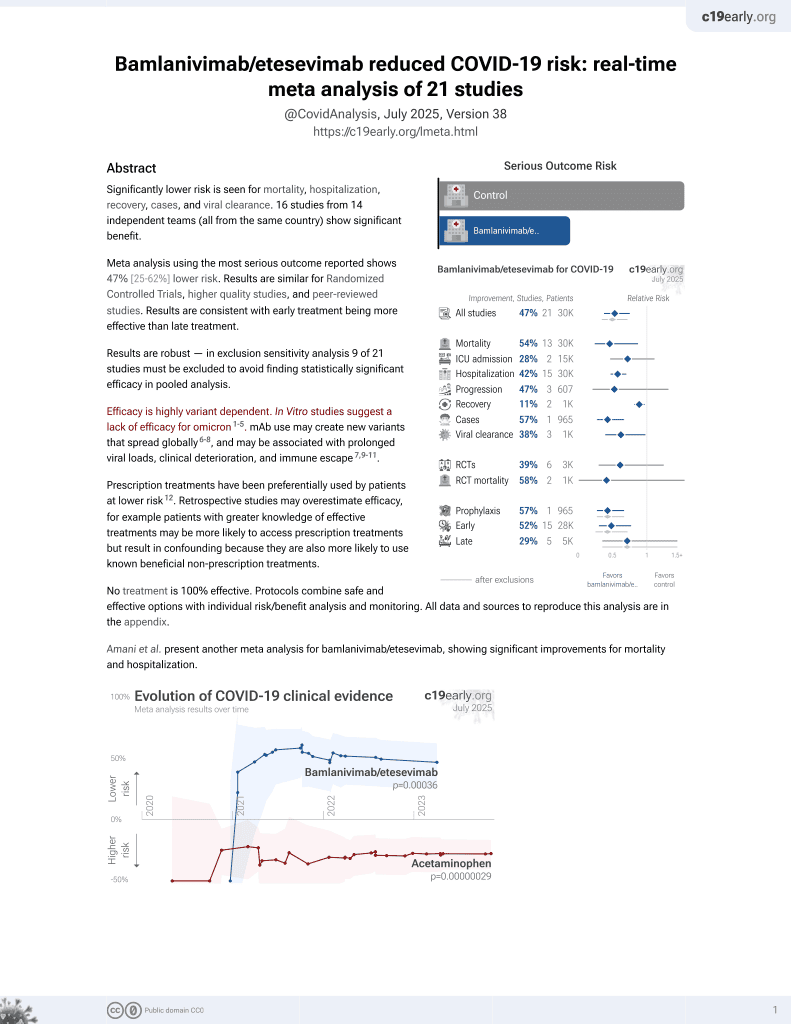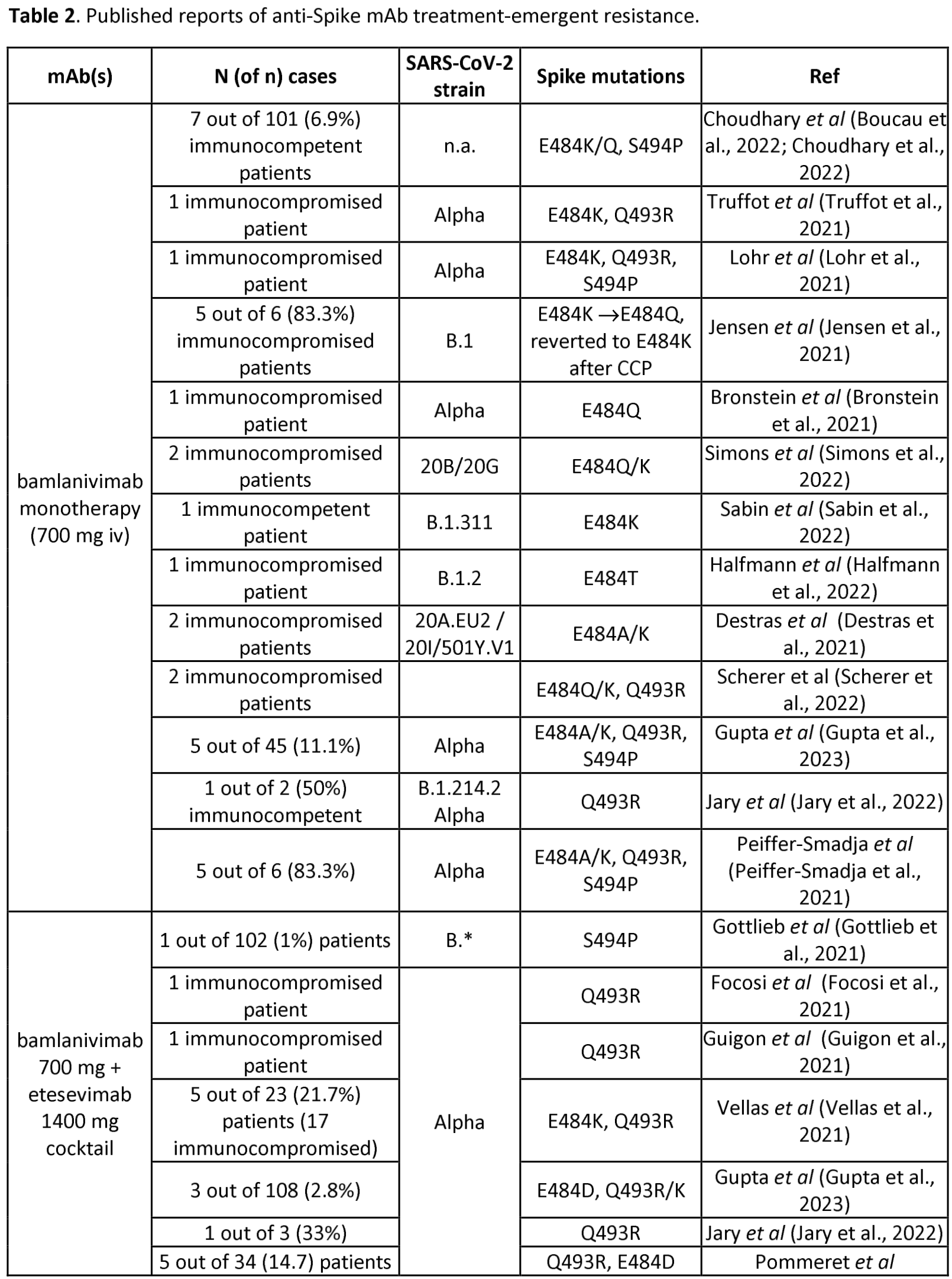
Analysis of SARS-CoV-2 mutations associated with resistance to therapeutic monoclonal antibodies that emerge after treatment
et al., Drug Resistance Updates, doi:10.1016/j.drup.2023.100991, Aug 2023
24th treatment shown to reduce risk in
May 2021, now with p = 0.00049 from 22 studies, recognized in 11 countries.
Efficacy is variant dependent.
No treatment is 100% effective. Protocols
combine treatments.
6,600+ studies for
220+ treatments. c19early.org
|
Review of reports of treatment-emergent resistance to COVID-19 monoclonal antibodies (mAbs), showing that some post-mAb treatment mutations appeared to spread globally soon after the mAb was introduced, raising concerns about transmission from treated patients. Treatment-emergent resistance was common, occurring in 10-50% of patients, with most events in immunocompromised patients.
Study covers bamlanivimab/etesevimab, casirivimab/imdevimab, tixagevimab/cilgavimab, sotrovimab, and bebtelovimab.
1.
Focosi et al., The Emergence of Escape Mutations in COVID-19 Following Anti-Spike Monoclonal Antibody Treatment: How Do We Tackle It?, Infection and Drug Resistance, doi:10.2147/IDR.S540928.
Focosi et al., 10 Aug 2023, Italy, peer-reviewed, 4 authors.
Contact: daniele.focosi@gmail.com.
Analysis of SARS-CoV-2 mutations associated with resistance to therapeutic monoclonal antibodies that emerge after treatment
doi:10.1101/2023.03.02.23286677
The mutation rate of the Omicron sublineage has led to baseline resistance against all previously authorized anti-Spike monoclonal antibodies (mAbs). Nevertheless, in case more antiviral mAbs will be authorized in the future, it is relevant to understand how frequently treatment-emergent resistance has emerged so far, under different combinations and in different patient subgroups. We report the results of a systematic review of the medical literature for case reports and case series for treatment-emergent immune escape, which is defined as emergence of a resistance-driving mutation in at least 20% of sequences in a given host at a given timepoint. We identified 31 publications detailing 201 cases that included different variants of concern (VOC) and found that the incidence of treatment emergentresistance ranged from 10% to 50%. Most of the treatment-emergent resistance events occurred in immunocompromised patients. Interestingly, resistance also emerged against cocktails of two mAbs, albeit at lower frequencies. The heterogenous therapeutic management of those cases doesn't allow
References
>438 (baum, None, FDA
Ai, Wang, He, Zhao, Zhang et al., Antibody evasion of SARS-CoV-2 Omicron BA.1, BA.1.1, BA.2, and BA.3 sub-lineages, Cell host & microbe
Anderson, O'donoghue, Mechanic, Dechen, Stevens, Administration of Anti-SARS-CoV-2 Monoclonal Antibodies After US Food and Drug Administration Deauthorization, JAMA Network Open
Andrés, González-Sánchez, Jiménez, Márquez-Algaba, Piñana et al., Emergence of Delta and Omicron variants carrying resistance-associated mutations in immunocompromised patients undergoing sotrovimab treatment with long-term viral excretion, Clinical Microbiology and Infection
Annavajhala, Mohri, Wang, Nair, Zucker et al., Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York, Nature
Ashano, Myers, Rokadiya, Hopkins, Brown et al., Comparison of the risk of hospitalisation among BA.1 and BA.2 COVID-19 cases treated with Sotrovimab in the community in England
Baum, Fulton, Wloga, Copin, Pascal et al., Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies
Birnie, Biemond, Appelman, De Bree, Jonges et al., Development of Resistance-Associated Mutations After Sotrovimab Administration in High-risk Individuals Infected With the SARS-CoV-2 Omicron Variant, Jama
Boucau, Chew, Choudhary, Deo, Regan et al., Evolution of spike mutations following antibody treatment in two immunocompromised patients with persistent COVID-19 infection
Cao, Jian, Wang, Yu, Song et al., Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution, Nature
Casadevall, Focosi, SARS-CoV-2 variants resistant to monoclonal antibodies in immunocompromised patients is a public health concern, The Journal of clinical investigation
Cathcart, Havenar-Daughton, Lempp, Ma, Schmid et al., The dual function monoclonal antibodies VIR-7831 and VIR-7832 demonstrate potent in vitro and in vivo activity against SARS-CoV-2
Chen, Nadeau, Yared, Voinov, Xie et al., CoV-Spectrum: analysis of globally shared SARS-CoV-2 data to identify and characterize new variants, Bioinformatics
Chen, Zhang, Case, Winkler, Liu et al., None
Chik, -T. Yuen, Yoon, To, Chen et al., Striking Antibody Evasion Manifested by the Omicron Variant of SARS-CoV
Choudhary, Chew, Deo, Flynn, Regan et al., for the, Emergence of SARS-CoV-2 escape mutations during Bamlanivimab therapy in a phase II randomized clinical trial, Nature Microbiology
Clark, Clark, Pan, Coscia, Mckay et al., SARS-CoV-2 evolution in an immunocompromised host reveals shared neutralization escape mechanisms, Cell
Copin, Baum, Wloga, Pascal, Giordano et al., The monoclonal antibody combination REGEN-COV protects against SARS-CoV-2 mutational escape in preclinical and human studies, Cell
Destras, Assaad, Bal, Bouscambert-Duchamp, Avrillon et al., Bamlanivimab as monotherapy in two immunocompromised patients with COVID-19, The Lancet Microbe
Destras, Bal, Simon, Lina, Josset, Sotrovimab drives SARS-CoV-2 Omicron variant evolution in immunocompromised patients, The Lancet. Microbe
Du, Hurdiss, Drabek, Mykytyn, Kaiser et al., An ACE2-blocking antibody confers broad neutralization and protection against Omicron and other SARS-CoV-2 variants
Ema, Lilly, Company Limited use of bamlanivimab and etesevimab for the treatment of COVID-19
Fabeni, Gruber, Tucci, Mazzotta, Rueca et al., Treatment-emergent cilgavimab resistance in Omicron
Fda, FDA, FDA authorizes revisions to fact sheets to address SARS-CoV-2 variants for monoclonal antibody products under emergency use authorization
Fda, Fact sheet for healthcare providers: emergency use authorization for Evusheld™ (tixagevimab copackaged with cilgavimab
Focosi, A web tool to estimate baaseline anti-Spike monoclonal antibody efficacy based on regional genomic surveillance
Focosi, Mcconnell, Casadevall, Cappello, Valdiserra et al., Monoclonal antibody therapies against SARS-CoV-2
Focosi, Novazzi, Baj, Ferrante, Boutahar et al., Sotrovimab-emergent resistance in SARS-CoV-2 Omicron: A series of three cases, J Clin Virol Plus
Focosi, Novazzi, Genoni, Dentali, Dalla et al., None
Focosi, Quiroga, Mcconnell, Johnson, Casadevall, Convergent evolution in SARS-CoV-2 Spike creates a variant soup from which new COVID-19 waves emerge, International Journal of Molecular Sciences
Focosi, Tuccori, Prescription of Anti-Spike Monoclonal Antibodies in COVID-19 Patients with Resistant SARS-CoV-2 Variants in Italy, Pathogens
Gliga, Luebke, Killer, Gruell, Walker et al., Rapid selection of sotrovimab escape variants in SARS-CoV-2 Omicron infected immunocompromised patients, Clinical Infectious Diseases
Gottlieb, Nirula, Chen, Boscia, Heller et al., Effect of Bamlanivimab as Monotherapy or in Combination With Etesevimab on Viral Load in Patients With Mild to Moderate COVID-19: A Randomized Clinical Trial
Greaney, Loes, Crawford, Starr, Malone et al., Comprehensive mapping of mutations to the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human serum antibodies, bioRxiv
Guigon, Faure, Lemaire, Chopin, Tinez et al., Emergence of Q493R mutation in SARS-CoV-2 spike protein during bamlanivimab/etesevimab treatment and resistance to viral clearance, J Infect S
Gupta, Konnova, Smet, Berkell, Savoldi et al., Host immunological responses facilitate development of SARS-CoV-2 2 mutations in patients receiving monoclonal antibody treatments, The Journal of clinical investigation
Halfmann, Minor, Haddock, Maddox, Moreno et al., Evolution of a globally unique SARS-CoV-2 Spike E484T monoclonal antibody escape mutation in a persistently infected
Harman, Nash, Webster, Groves, Hardstaff et al., None
Hoffmann, Zhang, Krüger, Graichen, Kleine-Weber et al., SARS-CoV-2 mutations acquired in mink reduce antibody-mediated neutralization
House, Broge, Suscovich, Snow, Tomic et al., Evaluation of strategies to modify Anti-SARS-CoV-2 monoclonal antibodies for optimal functionality as therapeutics, PloS one
Huygens, Munnink, Gharbharan, Koopmans, Rijnders, Sotrovimab resistance and viral persistence after treatment of immunocompromised patients infected with the SARS-CoV-2 Omicron variant, Clinical Infectious Diseases
Iketani, Liu, Guo, Liu, Huang et al., Antibody Evasion Properties of SARS-CoV-2 Omicron Sublineages, Nature
Jary, Marot, Faycal, Leon, Sayon et al., Spike Gene Evolution and Immune Escape Mutations in Patients with Mild or Moderate Forms of COVID-19 and Treated with Monoclonal Antibodies Therapies, Viruses
Jensen, Luebke, Feldt, Keitel, Brandenburger et al., Emergence of the E484K mutation in SARS-COV-2-infected immunocompromised patients treated with bamlanivimab in Germany, The Lancet Regional Health -Europe
Liu, Iketani, Guo, -W. Chan, Wang et al., None
Lohr, Niemann, Verheyen, Bamlanivimab treatment leads to rapid selection of immune escape variant carrying E484K mutation in a B.1.1.7 infected and immunosuppressed patient, Clinical Infectious Diseases
Nabel, Clark, Shankar, Pan, Clark et al., Structural basis for continued antibody evasion by the SARS-CoV-2 receptor binding domain
Nabel, EMA
Patel, Levick, Boult, Gibbons, Drysdale et al., Characteristics and Outcomes of COVID-19 Patients Presumed to be Treated with Sotrovimab in NHS, Hospitals in England
Peiffer-Smadja, Bridier-Nahmias, Ferré, Charpentier, Garé et al., Emergence of E484K Mutation Following Bamlanivimab Monotherapy among High-Risk Patients Infected with the Alpha Variant of SARS-CoV-2, Viruses
Pommeret, Colomba, Bigenwald, Laparra, Bockel et al., Bamlanivimab+ etesevimab therapy induces SARS-CoV-2 immune escape mutations and secondary clinical deterioration in COVID-19 patients with B-cell malignancies, Annals of Oncology
Ragonnet-Cronin, Nutalai, Huo, Dijokaite-Guraliuc, Das et al., Genome-first detection of emerging resistance to novel therapeutic agents for SARS-CoV-2, bioRxiv
Rockett, Basile, Maddocks, Fong, Agius et al., Resistance mutations in SARS-CoV-2 Delta variant after sotrovimab use, N Engl J Med
Ryu, Woo, Kang, Noh, Kim et al., The in vitro and in vivo potency of CT-P59 against Delta and its associated variants of SARS-CoV-2
Sabin, Richmond, Kenny, Emergence and onward transmission of a SARS-CoV-2 E484K variant among household contacts of a bamlanivimab-treated patient, Diagn Microbiol Infect Dis
Scherer, Babiker, Adelman, Allman, Key et al., SARS-CoV-2 Evolution and Immune Escape in Immunocompromised Patients
Simons, Ozer, Gambut, Dean, Zhang et al., De novo emergence of SARS-CoV-2 spike mutations in immunosuppressed patients, Transplant infectious disease : an official journal of the Transplantation Society
Stadler, Burgess, Schlub, Chai, Mcquilten et al., Monoclonal antibody levels and protection from COVID-19
Stadler, Chai, Schlub, Cromer, Polizzotto et al., Determinants of passive antibody efficacy in SARS-CoV-2 infection
Starr, Greaney, Addetia, Hannon, Choudhary et al., Prospective mapping of viral mutations that escape antibodies used to treat COVID-19, Science
Suryadevara, Gilchuk, Zost, Tahan, Droit et al., Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies, Nat Med
Tada, Dcosta, Samanovic, Herati, Cornelius et al., Convalescent-Phase Sera and Vaccine-Elicited Antibodies Largely Maintain Neutralizing Titer against Global SARS-CoV-2 Variant Spikes
Tada, Zhou, Dcosta, Samanovic, Chivukula et al., Increased resistance of SARS-CoV-2 Omicron Variant to Neutralization by Vaccine-Elicited and Therapeutic Antibodies, EBioMedicine
Tada, Zhou, Dcosta, Samanovic, Mulligan et al., Partial resistance of SARS-CoV-2 Delta variants to vaccine-elicited antibodies and convalescent sera, iScience
Tada, Zhou, Samanovic, Dcosta, Cornelius et al., Neutralization of SARS-CoV-2 Variants by mRNA and Adenoviral Vector Vaccine-Elicited Antibodies, Frontiers in immunology
Thomson, Rosen, Shepherd, Spreafico, Da Silva Filipe et al., The circulating SARS-CoV
Truffot, Andreani, Le Marechal, Caporossi, Epaulard et al., SARS-CoV-2 Variants in Immunocompromised Patient Given Antibody Monotherapy, Emerging infectious diseases
Tzou, Tao, Nouhin, Rhee, Hu et al., Coronavirus Antiviral Research Database (CoV-RDB): An Online Database Designed to Facilitate Comparisons between Candidate Anti-Coronavirus Compounds
Vellas, Del, Bello, Alexa, Steinmeyer et al., Influence of neutralizing monoclonal antibodies on the SARS-CoV-2 quasispecies in patients with COVID-19
Vellas, Kamar, Izopet, Resistance mutations in SARS-CoV-2 Omicron variant after tixagevimab-cilgavimab treatment, Journal of Infection
Vellas, Trémeaux, Del, Bello, Latour et al., Resistance mutations in SARS-CoV-2 Omicron variant in patients treated with sotrovimab, Clinical Microbiology and Infection
Wang, Guo, Iketani, Li, Mohri et al., Antibody evasion by SARS-CoV-2 Omicron subvariants BA.2.12.1, BA.4 and BA, Nature
Wang, Iketani, Li, Guo, Yeh et al., Antigenic characterization of the SARS-CoV-2 Omicron subvariant BA
Wang, Nair, Lihong, Iketani, Luo et al., Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7, Nature
Wang, Schmidt, Weisblum, Muecksch, Barnes et al., mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants, bioRxiv
Wang, Zhang, Ge, Ren, Zhang et al., Analysis of SARS-CoV-2 variant mutations reveals neutralization escape mechanisms and the ability to use ACE2 receptors from additional species
Wang, Zhang, Liu, Wang, Zhan et al., Combating the SARS-CoV-2 Omicron variant with non-Omicron neutralizing antibodies
Wang, Zhou, Zhang, Yang, Schramm et al., None
Westendorf, Žentelis, Foster, Vaillancourt, Wiggin et al., None, Cell Rep
Wu, Carr, Harvey, Mears, Kjaer et al., WHO's Therapeutics and COVID-19 Living Guideline on mAbs needs to be reassessed, Lancet
Yao, Ma, Wang, Tang, Du et al., Effect of SARS-CoV-2 spike mutations on animal ACE2 usage and in vitro neutralization sensitivity
Yi, Novel Coronavirus Is Undergoing Active Recombination, Clinical Infectious Diseases
Yi, Sun, Lin, Gu, Ding et al., Comprehensive mapping of binding hot spots of SARS-CoV-2 RBD-specific neutralizing antibodies for tracking immune escape variants, Genome Medicine
Yuan, Huang, Lee, Wu, Jackson et al., Structural and functional ramifications of antigenic drift in recent SARS-CoV-2 variants
Zhou, Dcosta, Landau, Tada, Resistance of SARS-CoV-2 Omicron BA.1 and BA.2 Variants to Vaccine-Elicited Sera and Therapeutic Monoclonal Antibodies, Viruses
Zhou, Wang, Misasi, Pegu, Zhang et al., Structural basis for potent antibody neutralization of SARS-CoV
DOI record:
{
"DOI": "10.1016/j.drup.2023.100991",
"ISSN": [
"1368-7646"
],
"URL": "http://dx.doi.org/10.1016/j.drup.2023.100991",
"alternative-id": [
"S1368764623000742"
],
"article-number": "100991",
"assertion": [
{
"label": "This article is maintained by",
"name": "publisher",
"value": "Elsevier"
},
{
"label": "Article Title",
"name": "articletitle",
"value": "Analysis of SARS-CoV-2 mutations associated with resistance to therapeutic monoclonal antibodies that emerge after treatment"
},
{
"label": "Journal Title",
"name": "journaltitle",
"value": "Drug Resistance Updates"
},
{
"label": "CrossRef DOI link to publisher maintained version",
"name": "articlelink",
"value": "https://doi.org/10.1016/j.drup.2023.100991"
},
{
"label": "Content Type",
"name": "content_type",
"value": "article"
},
{
"label": "Copyright",
"name": "copyright",
"value": "© 2023 Elsevier Ltd. All rights reserved."
}
],
"author": [
{
"ORCID": "http://orcid.org/0000-0001-8811-195X",
"affiliation": [],
"authenticated-orcid": false,
"family": "Focosi",
"given": "Daniele",
"sequence": "first"
},
{
"affiliation": [],
"family": "McConnell",
"given": "Scott",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Sullivan",
"given": "David J.",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Casadevall",
"given": "Arturo",
"sequence": "additional"
}
],
"container-title": "Drug Resistance Updates",
"container-title-short": "Drug Resistance Updates",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"clinicalkey.fr",
"clinicalkey.jp",
"clinicalkey.es",
"clinicalkey.com.au",
"clinicalkey.com",
"elsevier.com",
"sciencedirect.com"
]
},
"created": {
"date-parts": [
[
2023,
7,
31
]
],
"date-time": "2023-07-31T15:48:21Z",
"timestamp": 1690818501000
},
"deposited": {
"date-parts": [
[
2023,
8,
14
]
],
"date-time": "2023-08-14T16:34:22Z",
"timestamp": 1692030862000
},
"indexed": {
"date-parts": [
[
2023,
8,
15
]
],
"date-time": "2023-08-15T04:24:16Z",
"timestamp": 1692073456065
},
"is-referenced-by-count": 0,
"issued": {
"date-parts": [
[
2023,
11
]
]
},
"language": "en",
"license": [
{
"URL": "https://www.elsevier.com/tdm/userlicense/1.0/",
"content-version": "tdm",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
},
{
"URL": "https://doi.org/10.15223/policy-017",
"content-version": "stm-asf",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
},
{
"URL": "https://doi.org/10.15223/policy-037",
"content-version": "stm-asf",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
},
{
"URL": "https://doi.org/10.15223/policy-012",
"content-version": "stm-asf",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
},
{
"URL": "https://doi.org/10.15223/policy-029",
"content-version": "stm-asf",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
},
{
"URL": "https://doi.org/10.15223/policy-004",
"content-version": "stm-asf",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2023,
11,
1
]
],
"date-time": "2023-11-01T00:00:00Z",
"timestamp": 1698796800000
}
}
],
"link": [
{
"URL": "https://api.elsevier.com/content/article/PII:S1368764623000742?httpAccept=text/xml",
"content-type": "text/xml",
"content-version": "vor",
"intended-application": "text-mining"
},
{
"URL": "https://api.elsevier.com/content/article/PII:S1368764623000742?httpAccept=text/plain",
"content-type": "text/plain",
"content-version": "vor",
"intended-application": "text-mining"
}
],
"member": "78",
"original-title": [],
"page": "100991",
"prefix": "10.1016",
"published": {
"date-parts": [
[
2023,
11
]
]
},
"published-print": {
"date-parts": [
[
2023,
11
]
]
},
"publisher": "Elsevier BV",
"reference": [
{
"DOI": "10.1101/2021.03.09.434607",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib1",
"unstructured": "A.L. Cathcart, C. Havenar-Daughton, F.A. Lempp, D. Ma, M. Schmid, M.L. Agostini, B. Guarino, J. Di iulio, L. Rosen, H. Tucker, J. Dillen, S. Subramanian, B. Sloan, S. Bianchi, J. Wojcechowskyj, J. Zhou, H. Kaiser, A. Chase, M. Montiel-Ruiz, N. Czudnochowski, E. Cameroni, S. Ledoux, C. Colas, L. Soriaga, A. Telenti, S. Hwang, G. Snell, H.W. Virgin, D. Corti and C.M. Hebner, The dual function monoclonal antibodies VIR-7831 and VIR-7832 demonstrate potent in vitro and in vivo activity against SARS-CoV-2, (2021), p. 2021.2003.2009.434607."
},
{
"DOI": "10.1016/j.chom.2022.05.001",
"article-title": "Antibody evasion of SARS-CoV-2 Omicron BA.1, BA.1.1, BA.2, and BA.3 sub-lineages",
"author": "Ai",
"doi-asserted-by": "crossref",
"first-page": "1077",
"journal-title": "Cell Host Microbe",
"key": "10.1016/j.drup.2023.100991_bib2",
"volume": "30",
"year": "2022"
},
{
"DOI": "10.1001/jamanetworkopen.2022.28997",
"article-title": "Administration of anti–SARS-CoV-2 monoclonal antibodies after US food and drug administration deauthorization",
"author": "Anderson",
"doi-asserted-by": "crossref",
"journal-title": "JAMA Netw. Open",
"key": "10.1016/j.drup.2023.100991_bib3",
"volume": "5",
"year": "2022"
},
{
"DOI": "10.1016/j.cmi.2022.08.021",
"article-title": "Emergence of Delta and Omicron variants carrying resistance-associated mutations in immunocompromised patients undergoing sotrovimab treatment with long-term viral excretion",
"author": "Andrés",
"doi-asserted-by": "crossref",
"first-page": "240",
"journal-title": "Clin. Microbiol. Infect.",
"key": "10.1016/j.drup.2023.100991_bib4",
"volume": "29",
"year": "2023"
},
{
"DOI": "10.1038/s41586-021-03908-2",
"article-title": "Emergence and expansion of SARS-CoV-2 B.1.526 after identification in New York",
"author": "Annavajhala",
"doi-asserted-by": "crossref",
"first-page": "703",
"journal-title": "Nature",
"key": "10.1016/j.drup.2023.100991_bib5",
"volume": "597",
"year": "2021"
},
{
"article-title": "Antibody cocktail to SARS-CoV-2 spike protein prevents rapid mutational escape seen with individual antibodies",
"author": "Baum",
"first-page": "1014",
"key": "10.1016/j.drup.2023.100991_bib6",
"volume": "369",
"year": "2020"
},
{
"DOI": "10.1001/jama.2022.13854",
"article-title": "Development of resistance-associated mutations after sotrovimab administration in high-risk individuals infected with the SARS-CoV-2 omicron variant",
"author": "Birnie",
"doi-asserted-by": "crossref",
"first-page": "1104",
"journal-title": "JAMA",
"key": "10.1016/j.drup.2023.100991_bib7",
"volume": "328",
"year": "2022"
},
{
"article-title": "Monoclonal antibody treatment drives rapid culture conversion in SARS-CoV-2 infection",
"author": "Boucau",
"journal-title": "Cell Rep. Med.",
"key": "10.1016/j.drup.2023.100991_bib8",
"volume": "3",
"year": "2022"
},
{
"article-title": "Evolution of spike mutations following antibody treatment in two immunocompromised patients with persistent COVID-19 infection",
"author": "Bronstein",
"journal-title": "J. Med. Virol.",
"key": "10.1016/j.drup.2023.100991_bib9",
"year": "2021"
},
{
"DOI": "10.1038/s41586-022-05644-7",
"article-title": "Imprinted SARS-CoV-2 humoral immunity induces convergent Omicron RBD evolution",
"author": "Cao",
"doi-asserted-by": "crossref",
"journal-title": "Nature",
"key": "10.1016/j.drup.2023.100991_bib10",
"year": "2022"
},
{
"DOI": "10.1172/JCI168603",
"article-title": "SARS-CoV-2 variants resistant to monoclonal antibodies in immunocompromised patients is a public health concern",
"author": "Casadevall",
"doi-asserted-by": "crossref",
"issue": "6",
"journal-title": "J. Clin. Investig.",
"key": "10.1016/j.drup.2023.100991_bib11",
"volume": "133",
"year": "2023"
},
{
"DOI": "10.1093/bioinformatics/btab856",
"article-title": "CoV-Spectrum: analysis of globally shared SARS-CoV-2 data to identify and characterize new variants",
"author": "Chen",
"doi-asserted-by": "crossref",
"first-page": "1735",
"journal-title": "Bioinformatics",
"key": "10.1016/j.drup.2023.100991_bib12",
"volume": "38",
"year": "2021"
},
{
"DOI": "10.1038/s41591-021-01294-w",
"article-title": "Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies",
"author": "Chen",
"doi-asserted-by": "crossref",
"first-page": "717",
"journal-title": "Nat. Med.",
"key": "10.1016/j.drup.2023.100991_bib13",
"volume": "27",
"year": "2021"
},
{
"DOI": "10.1056/NEJMc2031364",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib14",
"unstructured": "B. Choi, M.C. Choudhary, J. Regan, J.A. Sparks, R.F. Padera, X. Qiu, I.H. Solomon, H.-H. Kuo, J. Boucau, K. Bowman, U.D. Adhikari, M.L. Winkler, A.A. Mueller, T.Y.-T. Hsu, M. Desjardins, L.R. Baden, B.T. Chan, B.D. Walker, M. Lichterfeld, M. Brigl, D.S. Kwon, S. Kanjilal, E.T. Richardson, A.H. Jonsson, G. Alter, A.K. Barczak, W.P. Hanage, X.G. Yu, G.D. Gaiha, M.S. Seaman, M. Cernadas and J.Z. Li, Persistence and Evolution of SARS-CoV-2 in an Immunocompromised Host, 383 (2020), pp. 2291–2293."
},
{
"DOI": "10.1038/s41564-022-01254-1",
"article-title": "Emergence of SARS-CoV-2 escape mutations during Bamlanivimab therapy in a phase II randomized clinical trial",
"author": "Choudhary",
"doi-asserted-by": "crossref",
"first-page": "1906",
"journal-title": "Nat. Microbiol.",
"key": "10.1016/j.drup.2023.100991_bib15",
"volume": "7",
"year": "2022"
},
{
"DOI": "10.1016/j.cell.2021.03.027",
"article-title": "SARS-CoV-2 evolution in an immunocompromised host reveals shared neutralization escape mechanisms",
"author": "Clark",
"doi-asserted-by": "crossref",
"first-page": "2605",
"journal-title": "Cell",
"key": "10.1016/j.drup.2023.100991_bib16",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.1016/j.cell.2021.06.002",
"article-title": "The monoclonal antibody combination REGEN-COV protects against SARS-CoV-2 mutational escape in preclinical and human studies",
"author": "Copin",
"doi-asserted-by": "crossref",
"first-page": "3949",
"journal-title": "Cell",
"key": "10.1016/j.drup.2023.100991_bib17",
"volume": "184",
"year": "2021"
},
{
"DOI": "10.1016/S2666-5247(21)00189-0",
"article-title": "Bamlanivimab as monotherapy in two immunocompromised patients with COVID-19",
"author": "Destras",
"doi-asserted-by": "crossref",
"journal-title": "Lancet Microbe",
"key": "10.1016/j.drup.2023.100991_bib18",
"volume": "2",
"year": "2021"
},
{
"DOI": "10.1016/S2666-5247(22)00120-3",
"article-title": "Sotrovimab drives SARS-CoV-2 Omicron variant evolution in immunocompromised patients",
"author": "Destras",
"doi-asserted-by": "crossref",
"journal-title": "Lancet Microbe",
"key": "10.1016/j.drup.2023.100991_bib19",
"volume": "3",
"year": "2022"
},
{
"article-title": "Real-world effectiveness of sotrovimab for the treatment of SARS-CoV-2 infection during Omicron BA.2 subvariant predominance: a systematic literature review",
"author": "Drysdale",
"journal-title": "medRxiv",
"key": "10.1016/j.drup.2023.100991_bib20",
"year": "2023"
},
{
"DOI": "10.1101/2022.02.17.480751",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib21",
"unstructured": "W. Du, D.L. Hurdiss, D. Drabek, A.Z. Mykytyn, F. Kaiser, M. Gonzalez-Hernandez, D. Munoz-Santos, M.M. Lamers, R. van Haperen, W. Li, I. Drulyte, C. Wang, I. Sola, F. Armando, G. Beythien, M. Ciurkiewicz, W. Baumgartner, K. Guilfoyle, T. Smits, J. van der Lee, F.J.M. van Kuppeveld, G. van Amerongen, B.L. Haagmans, L. Enjuanes, A.D. Osterhaus, F. Grosveld and B.J. Bosch, An ACE2-blocking antibody confers broad neutralization and protection against Omicron and other SARS-CoV-2 variants, (2022), p. 2022.2002.2017.480751."
},
{
"DOI": "10.1101/2020.11.04.355842",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib22",
"unstructured": "E.C. Thomson, L.E. Rosen, J.G. Shepherd, R. Spreafico, A. da Silva Filipe, J.A. Wojcechowskyj, C. Davis, L. Piccoli, D.J. Pascall, J. Dillen, S. Lytras, N. Czudnochowski, R. Shah, M. Meury, N. Jesudason, A. De Marco, K. Li, J. Bassi, A. O’Toole, D. Pinto, R.M. Colquhoun, K. Culap, B. Jackson, F. Zatta, A. Rambaut, S. Jaconi, V.B. Sreenu, J. Nix, R.F. Jarrett, M. Beltramello, K. Nomikou, M. Pizzuto, L. Tong, E. Cameroni, N. Johnson, A. Wickenhagen, A. Ceschi, D. Mair, P. Ferrari, K. Smollett, F. Sallusto, S. Carmichael, C. Garzoni, J. Nichols, M. Galli, J. Hughes, A. Riva, A. Ho, M.G. Semple, P.J.M. Openshaw, J.K. Baillie, S.J. Rihn, S.J. Lycett, H.W. Virgin, A. Telenti, D. Corti, D.L. Robertson and G. Snell, The circulating SARS-CoV-2 spike variant N439K maintains fitness while evading antibody-mediated immunity, (2020), p. 2020.2011.2004.355842."
},
{
"key": "10.1016/j.drup.2023.100991_bib23",
"unstructured": "EMA, Eli Lilly and Company Limited use of bamlanivimab and etesevimab for the treatment of COVID-19, (2021)."
},
{
"DOI": "10.7326/M22-3428",
"article-title": "Safety and efficacy of combination SARS-CoV-2 neutralizing monoclonal antibodies amubarvimab plus romlusevimab in nonhospitalized patients with COVID-19",
"author": "Evering",
"doi-asserted-by": "crossref",
"first-page": "658",
"journal-title": "Ann. Intern Med",
"key": "10.1016/j.drup.2023.100991_bib24",
"volume": "176",
"year": "2023"
},
{
"key": "10.1016/j.drup.2023.100991_bib25",
"unstructured": "L. Fabeni, C.E. Gruber, F. Tucci, V. Mazzotta, M. Rueca, G. Gramigna, A. Vergori, E. Giombini, O. Butera, D. Focosi, E. Nicastri, E. Girardi, F. Vaia, A. Antinori and F. Maggi, Treatment-emergent cilgavimab resistance in Omicron BA.4/5, submitted (2023)."
},
{
"key": "10.1016/j.drup.2023.100991_bib26",
"unstructured": "FDA, Fact sheet for healthcare providers: emergency use authorization for Evusheld™ (tixagevimab co-packaged with cilgavimab). Accessed online at https://www.fda.gov/media/154701/download on February 3, 2023, (2021)."
},
{
"key": "10.1016/j.drup.2023.100991_bib27",
"unstructured": "FDA, FDA authorizes revisions to Evusheld dosing. Accessed online on April 29, 2022 at https://www.fda.gov/drugs/drug-safety-and-availability/fda-authorizes-revisions-evusheld-dosing, (2022)."
},
{
"DOI": "10.3390/v15051048",
"article-title": "A web tool to estimate baseline anti-Spike monoclonal antibody efficacy based on regional genomic surveillance",
"author": "Focosi",
"doi-asserted-by": "crossref",
"first-page": "1048",
"issue": "5",
"journal-title": "Viruses",
"key": "10.1016/j.drup.2023.100991_bib28",
"volume": "15",
"year": "2023"
},
{
"article-title": "Prescription of anti-spike monoclonal antibodies in COVID-19 patients with resistant SARS-CoV-2 variants in Italy",
"author": "Focosi",
"journal-title": "Pathogenes",
"key": "10.1016/j.drup.2023.100991_bib29",
"volume": "11",
"year": "2022"
},
{
"key": "10.1016/j.drup.2023.100991_bib30",
"unstructured": "D. Focosi, F. Novazzi, A. Genoni, F. Dentali, D. Dalla Gasperina, A. Baj and F. Maggi, Daniele Focosi, Federica Novazzi, Angelo Genoni, Francesco Dentali, Daniela Dalla gasperina, Andreina Baj, Fabrizio Maggi, (2021)."
},
{
"DOI": "10.1016/S1473-3099(22)00311-5",
"article-title": "Monoclonal antibody therapies against SARS-CoV-2",
"author": "Focosi",
"doi-asserted-by": "crossref",
"first-page": "00311",
"journal-title": "Lancet Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib31",
"volume": "22",
"year": "2022"
},
{
"article-title": "Sotrovimab-emergent resistance in SARS-CoV-2 Omicron: a series of three cases",
"author": "Focosi",
"journal-title": "J. Clin. Virol.",
"key": "10.1016/j.drup.2023.100991_bib32",
"volume": "2",
"year": "2022"
},
{
"DOI": "10.3390/ijms24032264",
"article-title": "Convergent evolution in SARS-CoV-2 Spike creates a variant soup from which new COVID-19 waves emerge",
"author": "Focosi",
"doi-asserted-by": "crossref",
"first-page": "2264",
"journal-title": "Int. J. Mol. Sci.",
"key": "10.1016/j.drup.2023.100991_bib33",
"volume": "24",
"year": "2023"
},
{
"article-title": "The SARS-CoV-2 monoclonal antibody AZD3152 potently neutralises historical and currently circulating variants",
"author": "Francica",
"journal-title": "ECCMID",
"key": "10.1016/j.drup.2023.100991_bib34",
"year": "2023"
},
{
"DOI": "10.1093/cid/ciac802",
"article-title": "Rapid selection of sotrovimab escape variants in SARS-CoV-2 Omicron infected immunocompromised patients",
"author": "Gliga",
"doi-asserted-by": "crossref",
"first-page": "408",
"journal-title": "Clin. Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib35",
"volume": "76",
"year": "2023"
},
{
"DOI": "10.1038/s41467-023-38867-x",
"article-title": "Sequential intrahost evolution and onward transmission of SARS-CoV-2 variants",
"author": "Gonzalez-Reiche",
"doi-asserted-by": "crossref",
"journal-title": "Nature Comm",
"key": "10.1016/j.drup.2023.100991_bib36",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.1001/jama.2021.0202",
"article-title": "Effect of bamlanivimab as monotherapy or in combination with etesevimab on viral load in patients with mild to moderate COVID-19: a randomized clinical trial",
"author": "Gottlieb",
"doi-asserted-by": "crossref",
"first-page": "632",
"journal-title": "JAMA",
"key": "10.1016/j.drup.2023.100991_bib37",
"volume": "325",
"year": "2021"
},
{
"DOI": "10.1101/2020.12.31.425021",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib38",
"unstructured": "A.J. Greaney, A.N. Loes, K.H. Crawford, T.N. Starr, K.D. Malone, H.Y. Chu and J.D. Bloom, Comprehensive mapping of mutations to the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human serum antibodies, bioRxiv [Preprint] (2021)."
},
{
"article-title": "Alidjinou, Emergence of Q493R mutation in SARS-CoV-2 spike protein during bamlanivimab/etesevimab treatment and resistance to viral clearance",
"author": "Guigon",
"first-page": "00435",
"journal-title": "J. Infect. S0163-",
"key": "10.1016/j.drup.2023.100991_bib39",
"volume": "4453",
"year": "2021"
},
{
"DOI": "10.1172/JCI166032",
"article-title": "Host immunological responses facilitate development of SARS-CoV-2 2 mutations in patients receiving monoclonal antibody treatments",
"author": "Gupta",
"doi-asserted-by": "crossref",
"journal-title": "J. Clin. Investig.",
"key": "10.1016/j.drup.2023.100991_bib40",
"year": "2023"
},
{
"DOI": "10.1093/ve/veac104",
"article-title": "Evolution of a globally unique SARS-CoV-2 Spike E484T monoclonal antibody escape mutation in a persistently infected, immunocompromised individual",
"author": "Halfmann",
"doi-asserted-by": "crossref",
"journal-title": "Virus Evol.",
"key": "10.1016/j.drup.2023.100991_bib41",
"year": "2022"
},
{
"DOI": "10.1101/2022.10.21.22281171",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib42",
"unstructured": "K. Harman, S.G. Nash, H.H. Webster, N. Groves, J. Hardstaff, J. Bridgen, P.B. Blomquist, R. Hope, E. Ashano, R. Myers, S. Rokadiya, S. Hopkins, C.S. Brown, M. Chand, G. Dabrera and S. Thelwall, Comparison of the risk of hospitalisation among BA.1 and BA.2 COVID-19 cases treated with Sotrovimab in the community in England, (2022), p. 2022.2010.2021.22281171."
},
{
"DOI": "10.1101/2021.02.12.430998",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib43",
"unstructured": "M. Hoffmann, L. Zhang, N. Krüger, L. Graichen, H. Kleine-Weber, H. Hofmann-Winkler, A. Kempf, S. Nessler, J. Riggert, M.S. Winkler, S. Schulz, H.-M. Jäck and S. Pöhlmann, SARS-CoV-2 mutations acquired in mink reduce antibody-mediated neutralization, (2021), p. 2021.2002.2012.430998."
},
{
"DOI": "10.1371/journal.pone.0267796",
"article-title": "Evaluation of strategies to modify Anti-SARS-CoV-2 monoclonal antibodies for optimal functionality as therapeutics",
"author": "House",
"doi-asserted-by": "crossref",
"journal-title": "PLoS One",
"key": "10.1016/j.drup.2023.100991_bib44",
"volume": "17",
"year": "2022"
},
{
"DOI": "10.1093/cid/ciac601",
"article-title": "Sotrovimab resistance and viral persistence after treatment of immunocompromised patients infected with the SARS-CoV-2 Omicron variant",
"author": "Huygens",
"doi-asserted-by": "crossref",
"first-page": "e507",
"journal-title": "Clin. Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib45",
"volume": "76",
"year": "2023"
},
{
"DOI": "10.1038/s41586-022-04594-4",
"article-title": "Antibody evasion properties of SARS-CoV-2 omicron sublineages",
"author": "Iketani",
"doi-asserted-by": "crossref",
"first-page": "553",
"journal-title": "Nature",
"key": "10.1016/j.drup.2023.100991_bib46",
"volume": "604",
"year": "2022"
},
{
"DOI": "10.3390/v14020226",
"article-title": "Spike gene evolution and immune escape mutations in patients with mild or moderate forms of COVID-19 and treated with monoclonal antibodies therapies",
"author": "Jary",
"doi-asserted-by": "crossref",
"journal-title": "Viruses",
"key": "10.1016/j.drup.2023.100991_bib47",
"volume": "14",
"year": "2022"
},
{
"article-title": "Emergence of the E484K mutation in SARS-COV-2-infected immunocompromised patients treated with bamlanivimab in Germany",
"author": "Jensen",
"journal-title": "Lancet Reg. Health – Eur.",
"key": "10.1016/j.drup.2023.100991_bib48",
"volume": "8",
"year": "2021"
},
{
"DOI": "10.1101/2021.12.14.472719",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib49",
"unstructured": "L. Liu, S. Iketani, Y. Guo, J.F.-W. Chan, M. Wang, L. Liu, Y. Luo, H. Chu, Y. Huang, M.S. Nair, J. Yu, K.K.-H. Chik, T.T.-T. Yuen, C. Yoon, K.K.-W. To, H. Chen, M.T. Yin, M.E. Sobieszczyk, Y. Huang, H.H. Wang, Z. Sheng, K.-Y. Yuen and D.D. Ho, Striking Antibody Evasion Manifested by the Omicron Variant of SARS-CoV-2, (2021), p. 2021.2012.2014.472719."
},
{
"DOI": "10.1093/cid/ciab392",
"article-title": "Bamlanivimab treatment leads to rapid selection of immune escape variant carrying E484K mutation in a B.1.1.7 infected and immunosuppressed patient",
"author": "Lohr",
"doi-asserted-by": "crossref",
"journal-title": "Clin. Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib50",
"year": "2021"
},
{
"article-title": "SARS-CoV-2 evolution during persistent infection in a CAR-T recipient shows an escape to both sotrovimab and T-cell responses",
"author": "Mazzetti",
"journal-title": "J. Clin. Virol.",
"key": "10.1016/j.drup.2023.100991_bib51",
"volume": "3",
"year": "2023"
},
{
"DOI": "10.1126/science.abl6251",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib52",
"unstructured": "K.G. Nabel, S.A. Clark, S. Shankar, J. Pan, L.E. Clark, P. Yang, A. Coscia, L.G.A. McKay, H.H. Varnum, V. Brusic, N.V. Tolan, G. Zhou, M. Desjardins, S.E. Turbett, S. Kanjilal, A.C. Sherman, A. Dighe, R.C. LaRocque, E.T. Ryan, C. Tylek, J.F. Cohen-Solal, A.T. Darcy, D. Tavella, A. Clabbers, Y. Fan, A. Griffiths, I.R. Correia, J. Seagal, L.R. Baden, R.C. Charles and J. Abraham, Structural basis for continued antibody evasion by the SARS-CoV-2 receptor binding domain, 375 (2022), p. eabl6251."
},
{
"DOI": "10.1128/aac.00266-23",
"article-title": "Frequent emergence of resistance mutations following complex intra-host genomic dynamics in SARS-CoV-2 patients receiving Sotrovimab",
"author": "Palomino-Cabrera",
"doi-asserted-by": "crossref",
"journal-title": "Antimicrob. Agents Chemother.",
"key": "10.1016/j.drup.2023.100991_bib53",
"year": "2023"
},
{
"DOI": "10.1101/2023.02.08.23285654",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib54",
"unstructured": "V. Patel, B. Levick, S. Boult, D.C. Gibbons, M. Drysdale, E.J. Lloyd, M. Singh and H.J. Birch, Characteristics and Outcomes of COVID-19 Patients Presumed to be Treated with Sotrovimab in NHS Hospitals in England, (2023), p. 2023.2002.2008.23285654."
},
{
"DOI": "10.3390/v13081642",
"article-title": "Emergence of E484K mutation following bamlanivimab monotherapy among high-risk patients infected with the alpha variant of SARS-CoV-2",
"author": "Peiffer-Smadja",
"doi-asserted-by": "crossref",
"journal-title": "Viruses",
"key": "10.1016/j.drup.2023.100991_bib55",
"volume": "13",
"year": "2021"
},
{
"DOI": "10.1016/j.annonc.2021.07.015",
"article-title": "Bamlanivimab+ etesevimab therapy induces SARS-CoV-2 immune escape mutations and secondary clinical deterioration in COVID-19 patients with B-cell malignancies",
"author": "Pommeret",
"doi-asserted-by": "crossref",
"journal-title": "Ann. Oncol.",
"key": "10.1016/j.drup.2023.100991_bib56",
"year": "2021"
},
{
"DOI": "10.1038/s41467-023-37826-w",
"article-title": "Generation of SARS-CoV-2 escape mutations by monoclonal antibody therapy",
"author": "Ragonnet-Cronin",
"doi-asserted-by": "crossref",
"first-page": "3334",
"journal-title": "Nat. Comm.",
"key": "10.1016/j.drup.2023.100991_bib57",
"volume": "14",
"year": "2023"
},
{
"DOI": "10.1056/NEJMc2120219",
"article-title": "Resistance mutations in SARS-CoV-2 Delta variant after sotrovimab use",
"author": "Rockett",
"doi-asserted-by": "crossref",
"first-page": "1477",
"journal-title": "New Engl. J. Med.",
"key": "10.1016/j.drup.2023.100991_bib58",
"volume": "386",
"year": "2021"
},
{
"DOI": "10.1101/2021.07.23.453472",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib59",
"unstructured": "D.-K. Ryu, H.-M. Woo, B. Kang, H. Noh, J.-I. Kim, J.-M. Seo, C. Kim, M. Kim, J.-W. Kim, N. Kim, P. Jeon, H. Lee, J.-S. Yang, K.-C. Kim, J.-Y. Lee, M.-H. Lee, S.-S. Oh, H.-Y. Chung, K.-S. Kwon and S.-Y. Lee, The in vitro and in vivo potency of CT-P59 against Delta and its associated variants of SARS-CoV-2, (2021), p. 2021.2007.2023.453472."
},
{
"DOI": "10.1016/j.diagmicrobio.2022.115656",
"article-title": "Emergence and onward transmission of a SARS-CoV-2 E484K variant among household contacts of a bamlanivimab-treated patient",
"author": "Sabin",
"doi-asserted-by": "crossref",
"journal-title": "Diagn. Microbiol Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib60",
"volume": "103",
"year": "2022"
},
{
"article-title": "SARS-CoV-2 evolution and immune escape in immunocompromised patients",
"author": "Scherer",
"first-page": "2436",
"key": "10.1016/j.drup.2023.100991_bib61",
"volume": "386",
"year": "2022"
},
{
"DOI": "10.1111/tid.13914",
"article-title": "De novo emergence of SARS-CoV-2 spike mutations in immunosuppressed patients",
"author": "Simons",
"doi-asserted-by": "crossref",
"journal-title": "Transpl. Infect. Dis.: Off. J. Transplant. Soc.",
"key": "10.1016/j.drup.2023.100991_bib62",
"volume": "24",
"year": "2022"
},
{
"DOI": "10.1101/2022.03.21.22272672",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib63",
"unstructured": "E. Stadler, K.L. Chai, T.E. Schlub, D. Cromer, M.N. Polizzotto, S.J. Kent, C. Beecher, H. White, T. Turner, N. Skoetz, L. Estcourt, Z.K. McQuilten, E.M. Wood, D.S. Khoury and M.P. Davenport, Determinants of passive antibody efficacy in SARS-CoV-2 infection, (2022b), p. 2022.2003.2021.22272672."
},
{
"DOI": "10.1101/2022.11.22.22282199",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib64",
"unstructured": "E. Stadler, M.T. Burgess, T.E. Schlub, K.L. Chai, Z.K. McQuilten, E.M. Wood, M.N. Polizzotto, S.J. Kent, D. Cromer, M.P. Davenport and D.S. Khoury, Monoclonal antibody levels and protection from COVID-19, (2022a), p. 2022.2011.2022.22282199."
},
{
"article-title": "Prospective mapping of viral mutations that escape antibodies used to treat COVID-19",
"author": "Starr",
"first-page": "eabf9302",
"journal-title": "Science",
"key": "10.1016/j.drup.2023.100991_bib65",
"year": "2021"
},
{
"DOI": "10.1128/mBio.00696-21",
"article-title": "Convalescent-phase sera and vaccine-elicited antibodies largely maintain neutralizing titer against global SARS-CoV-2 variant spikes",
"author": "Tada",
"doi-asserted-by": "crossref",
"journal-title": "mBio",
"key": "10.1016/j.drup.2023.100991_bib66",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1016/j.isci.2021.103341",
"article-title": "Partial resistance of SARS-CoV-2 Delta variants to vaccine-elicited antibodies and convalescent sera",
"author": "Tada",
"doi-asserted-by": "crossref",
"journal-title": "iScience",
"key": "10.1016/j.drup.2023.100991_bib67",
"year": "2021"
},
{
"DOI": "10.1016/j.ebiom.2022.103944",
"article-title": "Increased resistance of SARS-CoV-2 omicron variant to neutralization by vaccine-elicited and therapeutic antibodies",
"author": "Tada",
"doi-asserted-by": "crossref",
"journal-title": "EBioMedicine",
"key": "10.1016/j.drup.2023.100991_bib68",
"year": "2022"
},
{
"DOI": "10.3389/fimmu.2022.797589",
"article-title": "Neutralization of SARS-CoV-2 variants by mRNA and adenoviral vector vaccine-elicited antibodies",
"author": "Tada",
"doi-asserted-by": "crossref",
"journal-title": "Front. Immunol.",
"key": "10.1016/j.drup.2023.100991_bib69",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.3201/eid2710.211509",
"article-title": "SARS-CoV-2 variants in immunocompromised patient given antibody monotherapy",
"author": "Truffot",
"doi-asserted-by": "crossref",
"journal-title": "Emerg. Infect. Dis.",
"key": "10.1016/j.drup.2023.100991_bib70",
"volume": "27",
"year": "2021"
},
{
"article-title": "Coronavirus antiviral research database (CoV-RDB): an online database designed to facilitate comparisons between candidate anti-coronavirus compounds",
"author": "Tzou",
"first-page": "1006",
"key": "10.1016/j.drup.2023.100991_bib71",
"volume": "12",
"year": "2020"
},
{
"article-title": "Influence of neutralizing monoclonal antibodies on the SARS-CoV-2 quasispecies in patients with COVID-19",
"author": "Vellas",
"journal-title": "Clin. Micro Infect.",
"key": "10.1016/j.drup.2023.100991_bib72",
"year": "2021"
},
{
"DOI": "10.1016/j.jinf.2022.07.014",
"article-title": "Resistance mutations in SARS-CoV-2 Omicron variant after tixagevimab-cilgavimab treatment",
"author": "Vellas",
"doi-asserted-by": "crossref",
"first-page": "e162",
"journal-title": "J. Infect.",
"key": "10.1016/j.drup.2023.100991_bib73",
"volume": "85",
"year": "2022"
},
{
"DOI": "10.1016/j.cmi.2022.05.002",
"article-title": "Resistance mutations in SARS-CoV-2 Omicron variant in patients treated with sotrovimab",
"author": "Vellas",
"doi-asserted-by": "crossref",
"first-page": "1297",
"journal-title": "Clin. Microbiol. Infect.",
"key": "10.1016/j.drup.2023.100991_bib74",
"volume": "28",
"year": "2022"
},
{
"DOI": "10.1126/science.abh1766",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib75",
"unstructured": "L. Wang, T. Zhou, Y. Zhang, E.S. Yang, C.A. Schramm, W. Shi, A. Pegu, O.K. Oloniniyi, A.R. Henry, S. Darko, S.R. Narpala, C. Hatcher, D.R. Martinez, Y. Tsybovsky, E. Phung, O.M. Abiona, A. Antia, E.M. Cale, L.A. Chang, M. Choe, K.S. Corbett, R.L. Davis, A.T. DiPiazza, I.J. Gordon, S.H. Hait, T. Hermanus, P. Kgagudi, F. Laboune, K. Leung, T. Liu, R.D. Mason, A.F. Nazzari, L. Novik, S. O’Connell, S. O’Dell, A.S. Olia, S.D. Schmidt, T. Stephens, C.D. Stringham, C.A. Talana, I.-T. Teng, D.A. Wagner, A.T. Widge, B. Zhang, M. Roederer, J.E. Ledgerwood, T.J. Ruckwardt, M.R. Gaudinski, P.L. Moore, N.A. Doria-Rose, R.S. Baric, B.S. Graham, A.B. McDermott, D.C. Douek, P.D. Kwong, J.R. Mascola, N.J. Sullivan and J. Misasi, Ultrapotent antibodies against diverse and highly transmissible SARS-CoV-2 variants, 373 (2021a), p. eabh1766."
},
{
"DOI": "10.1038/s41586-021-03398-2",
"article-title": "Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "130",
"journal-title": "Nature",
"key": "10.1016/j.drup.2023.100991_bib76",
"volume": "593",
"year": "2021"
},
{
"DOI": "10.1101/2022.07.31.502235",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib77",
"unstructured": "Q. Wang, S. Iketani, Z. Li, Y. Guo, A.Y. Yeh, M. Liu, J. Yu, Z. Sheng, Y. Huang, L. Liu and D.D. Ho, Antigenic characterization of the SARS-CoV-2 Omicron subvariant BA.2.75, (2022b), p. 2022.2007.2031.502235."
},
{
"DOI": "10.1038/s41586-022-05053-w",
"article-title": "Antibody evasion by SARS-CoV-2 Omicron subvariants BA.2.12.1, BA.4 and BA.5",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "603",
"journal-title": "Nature",
"key": "10.1016/j.drup.2023.100991_bib78",
"volume": "608",
"year": "2022"
},
{
"DOI": "10.1016/j.immuni.2021.06.003",
"article-title": "Analysis of SARS-CoV-2 variant mutations reveals neutralization escape mechanisms and the ability to use ACE2 receptors from additional species",
"author": "Wang",
"doi-asserted-by": "crossref",
"first-page": "1611",
"journal-title": "Immunity",
"key": "10.1016/j.drup.2023.100991_bib79",
"volume": "54",
"year": "2021"
},
{
"DOI": "10.1101/2022.01.30.478305",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib80",
"unstructured": "Y. Wang, X. Zhang, J. Liu, Y. Wang, W. Zhan, M. Liu, M. Zhang, Q. Wang, Q. Liu, T. Zhu, Y. Wen, Z. Chen, J. Zhao, F. Wu, L. Sun and J. Huang, Combating the SARS-CoV-2 Omicron variant with non-Omicron neutralizing antibodies, (2022c), p. 2022.2001.2030.478305."
},
{
"article-title": "mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants",
"author": "Wang",
"journal-title": "bioRxiv",
"key": "10.1016/j.drup.2023.100991_bib81",
"year": "2021"
},
{
"key": "10.1016/j.drup.2023.100991_bib82",
"unstructured": "C. Webber, R. Beavon, S. Thomas, L.A. Chang, T. Cohen and J. Perez, Trial in progress: a Phase I/III, randomised, modified double-blind, placebo- and active-controlled pre-exposure prophylaxis study of the SARS-CoV-2–neutralising antibody AZD3152 (SUPERNOVA). ECCMID, Copenhagen (2023)."
},
{
"DOI": "10.1101/2021.04.30.442182",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib83",
"unstructured": "K. Westendorf, S. Žentelis, D. Foster, P. Vaillancourt, M. Wiggin, E. Lovett, J. Hendle, A. Pustilnik, J.M. Sauder, L. Kraft, Y. Hwang, R.W. Siegel, J. Chen, B.A. Heinz, R.E. Higgs, N. Kalleward, K. Jepson, R. Goya, M.A. Smith, D.W. Collins, D. Pellacani, P. Xiang, V. de Puyraimond, M. Ricicova, L. Devorkin, C. Pritchard, A. O'Neill, C. Cohen, J. Dye, K.I. Huie, C.V. Badger, D. Kobasa, J. Audet, J.J. Freitas, S. Hassanali, I. Hughes, L. Munoz, H.C. Palma, B. Ramamurthy, R.W. Cross, T.W. Geisbert, I. Lanz, L. Anderson, P. Sipahimalani, K.S. Corbett, L. Wang, E.S. Yang, Y. Zhang, W. Shi, B.S. Graham, J.R. Mascola, T.L. Fernandez, C.L. Hansen, E. Falconer, B.E. Jones and B.C. Barnhart, LY-CoV1404 (bebtelovimab) potently neutralizes SARS-CoV-2 variants, Cell Rep (2022), p. 2021.2004.2030.442182."
},
{
"DOI": "10.1016/S0140-6736(22)01938-9",
"article-title": "WHO's therapeutics and COVID-19 living guideline on mAbs needs to be reassessed",
"author": "Wu",
"doi-asserted-by": "crossref",
"first-page": "2193",
"journal-title": "Lancet",
"key": "10.1016/j.drup.2023.100991_bib84",
"volume": "400",
"year": "2022"
},
{
"DOI": "10.1101/2021.01.27.428353",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib85",
"unstructured": "W. Yao, D. Ma, H. Wang, X. Tang, C. Du, H. Pan, C. Li, H. Lin, M. Farzan, J. Zhao, Y. Li and G. Zhong, Effect of SARS-CoV-2 spike mutations on animal ACE2 usage and in vitro neutralization sensitivity, (2021), p. 2021.2001.2027.428353."
},
{
"DOI": "10.1186/s13073-021-00985-w",
"article-title": "Comprehensive mapping of binding hot spots of SARS-CoV-2 RBD-specific neutralizing antibodies for tracking immune escape variants",
"author": "Yi",
"doi-asserted-by": "crossref",
"first-page": "164",
"journal-title": "Genome Med.",
"key": "10.1016/j.drup.2023.100991_bib86",
"volume": "13",
"year": "2021"
},
{
"DOI": "10.1101/2021.02.16.430500",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib87",
"unstructured": "M. Yuan, D. Huang, C.-C.D. Lee, N.C. Wu, A.M. Jackson, X. Zhu, H. Liu, L. Peng, M.J. van Gils, R.W. Sanders, D.R. Burton, S.M. Reincke, H. Prüss, J. Kreye, D. Nemazee, A.B. Ward and I.A. Wilson, Structural and functional ramifications of antigenic drift in recent SARS-CoV-2 variants, (2021), p. 2021.2002.2016.430500."
},
{
"DOI": "10.3390/v14061334",
"article-title": "Resistance of SARS-CoV-2 Omicron BA.1 and BA.2 variants to vaccine-elicited sera and therapeutic monoclonal antibodies",
"author": "Zhou",
"doi-asserted-by": "crossref",
"journal-title": "Viruses",
"key": "10.1016/j.drup.2023.100991_bib88",
"volume": "14",
"year": "2022"
},
{
"DOI": "10.1101/2021.12.27.474307",
"doi-asserted-by": "crossref",
"key": "10.1016/j.drup.2023.100991_bib89",
"unstructured": "T. Zhou, L. Wang, J. Misasi, A. Pegu, Y. Zhang, D.R. Harris, A.S. Olia, C.A. Talana, E.S. Yang, M. Chen, M. Choe, W. Shi, I.-T. Teng, A. Creanga, C. Jenkins, K. Leung, T. Liu, E.-S.D. Stancofski, T. Stephens, B. Zhang, Y. Tsybovsky, B.S. Graham, J.R. Mascola, N.J. Sullivan and P.D. Kwong, Structural basis for potent antibody neutralization of SARS-CoV-2 variants including B.1.1.529, (2021), p. 2021.2012.2027.474307."
}
],
"reference-count": 89,
"references-count": 89,
"relation": {},
"resource": {
"primary": {
"URL": "https://linkinghub.elsevier.com/retrieve/pii/S1368764623000742"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"Pharmacology (medical)",
"Infectious Diseases",
"Cancer Research",
"Pharmacology",
"Oncology"
],
"subtitle": [],
"title": "Analysis of SARS-CoV-2 mutations associated with resistance to therapeutic monoclonal antibodies that emerge after treatment",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.1016/elsevier_cm_policy",
"volume": "71"
}