Review of preclinical data of PF-07304814 and its active metabolite derivatives against SARS-CoV-2 infection
et al., Frontiers in Pharmacology, doi:10.3389/fphar.2022.1035969, NCT05011812, Nov 2022
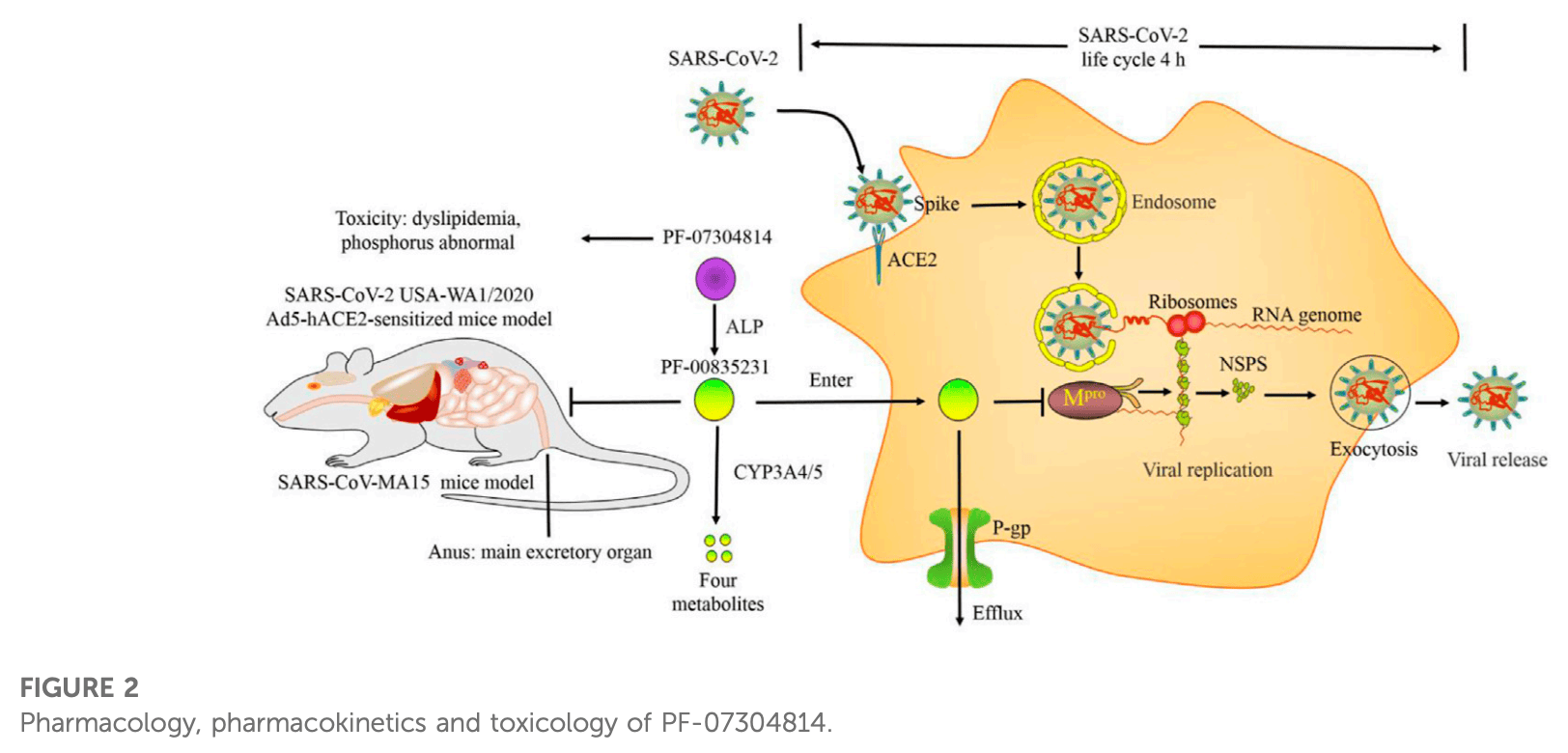
Review of preclinical data for lufotrelvir and active metabolites for SARS-CoV-2. Lufotrelvir (PF-07304814) is a phosphate ester prodrug that is rapidly metabolized by alkaline phosphatase into the active compound PF-0835231, which suppresses SARS-CoV-2 replication by inhibiting the main protease (Mpro). Authors note that PF-07304814 increased the bioavailability of PF-00835231 by enhancing plasma protein binding. P-glycoprotein inhibitors and cytochrome P450 3A inhibitors increased the efficacy of PF-00835231 by suppressing its efflux from target cells and metabolism, respectively. PF-00835231 inhibited SARS-CoV-2 infection in various cell lines, human respiratory epithelial organ models, and animal models. PF-07304814 exhibited a short terminal elimination half-life and was cleared primarily through renal elimination, with no significant adverse effects observed in rats.
Chen et al., 11 Nov 2022, placebo-controlled, China, peer-reviewed, 6 authors, trial NCT05011812 (history).
Contact: 18253199502@163.com, xdm_tsinghua@163.com.
Review of preclinical data of PF-07304814 and its active metabolite derivatives against SARS-CoV-2 infection
Frontiers in Pharmacology, doi:10.3389/fphar.2022.1035969
Main protease (M pro ) is a superior target for anti-SARS-COV-2 drugs. PF-07304814 is a phosphate ester prodrug of PF-00835231 that is rapidly metabolized into the active metabolite PF-00835231 by alkaline phosphatase (ALP) and then suppresses SARS-CoV-2 replication by inhibiting M pro . PF-07304814 increased the bioavailability of PF-00835231 by enhancing plasma protein binding (PPB). P-glycoprotein (P-gp) inhibitors and cytochrome P450 3A (CYP3A) inhibitors increased the efficacy of PF-00835231 by suppressing its efflux from target cells and metabolism, respectively. The life cycle of SARS-CoV-2 is approximately 4 h. The mechanisms and efficacy outcomes of PF-00835231 occur simultaneously. PF-00835231 can inhibit not only cell infection (such as Vero E6, 293T, Huh-7.5, HeLa +angiotensin- converting enzyme 2 (ACE2) , A549 +ACE2 , and MRC-5) but also the human respiratory epithelial organ model and animal model infection. PF-07304814 exhibits a short terminal elimination half-life and is cleared primarily through renal elimination. There were no significant adverse effects of PF-07304814 administration in rats. Therefore, PF-07304814 exhibits good tolerability, pharmacology, pharmacodynamics, pharmacokinetics, and safety in preclinical trials. However, the Phase 1 data of PF-07304814 were not released. The Phase 2/3 trial of PF-07304814 was also suspended. Interestingly, the antiviral activities of PF-00835231 derivatives (compounds 5-22) are higher than, similar to, or slightly weaker than those of PF-00835231. In particular, compound 22 exhibited the highest potency and had good safety and stability. However, the low solubility of compound 22 limits its clinical application. Prodrugs, nanotechnology and salt form drugs may solve this problem. In this review, we focus on the preclinical data of PF-07304814 and its active metabolite derivatives to hopefully provide knowledge for researchers to study SARS-CoV-2 infection.
Author contributions Conception and design: DX and JX. Manuscript writing: WC and YS. Acquisition and assembly of data: XP and BL. Final discussions and approval of the manuscript: All authors contributed to the article and approved the submitted version.
Conflict of interest The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's note All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fphar. 2022.1035969/full#supplementary-material
References
Agost-Beltrán, Bou-Iserte, Rodríguez, Fernández-De-La-Pradilla, González et al., Advances in the development of SARS-CoV-2 Mpro inhibitors, Molecules, doi:10.3390/molecules27082523
Ahmad, Batool, Ain, Kim, Choi, Exploring the binding mechanism of PF-07321332 SARS-CoV-2 protease inhibitor through molecular dynamics and binding free energy simulations, Int. J. Mol. Sci, doi:10.3390/ijms22179124
Anirudhan, Cheng, Cooper, Rong, Targeting SARS-CoV-2 viral proteases as a therapeutic strategy to treat COVID-19, J. Med. Virol, doi:10.1002/jmv.26814
Baig, Sharma, Ahmad, Abohashrh, Alam, Unveiling the effect of low pH on the SARS-CoV-2 main protease by molecular dynamics simulations, Polym. (Basel), doi:10.3390/polym13213823
Boras, Jones, Anson, Arenson, Aschenbrenner et al., Preclinical characterization of an intravenous coronavirus 3CL protease inhibitor for the potential treatment of COVID19, Nat. Commun, doi:10.1038/s41467-021-26239-2
Charlier, Coglianese, De Rosa, De Grazia, Operto et al., The effect of plasma protein binding on the therapeutic monitoring of antiseizure medications, Pharmaceutics, doi:10.3390/pharmaceutics13081208
Chen, Feng, Recent progress of glutathione (GSH) specific fluorescent probes: Molecular design, photophysical property, recognition mechanism and bioimaging, Crit. Rev. Anal. Chem, doi:10.1080/10408347.2020.1819193
Dai, Lee, Nathanson, Leonelli, Petros et al., Viral kinetics of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron infection in mRNA-vaccinated individuals treated and not treated with nirmatrelvir-ritonavir, medRxiv, doi:10.1101/2022.08.04.22278378
De Freitas Santoro, Câmara, De Freitas, Oliveira, SARS-COV-2 and ocular surface: From physiology to pathology, a route to understand transmission and disease, Front. Physiol, doi:10.3389/fphys.2021.612319
De Vries, Mohamed, Prescott, Valero-Jimenez, Desvignes et al., A comparative analysis of SARS-CoV-2 antivirals characterizes 3CL(pro) inhibitor PF-00835231 as a potential new treatment for COVID-19, J. Virol, doi:10.1128/JVI.01819-20
Fernandes, Merhi, Mestiri, Taib, Moustafa et al., Emerging COVID-19 variants and their impact on SARS-CoV-2 diagnosis, therapeutics and vaccines, Ann. Med, doi:10.1080/07853890.2022.2031274
Fulmali, Bharate, Phosphate moiety in FDA-approved pharmaceutical salts and prodrugs, Drug Dev. Res, doi:10.1002/ddr.21953
Garcia-Lledo, Gomez-Pavon, Gonzalez Del Castillo, Hernandez-Sampelayo, Martin-Delgado et al., Pharmacological treatment of COVID-19: An opinion paper, Rev. Esp. Quimioter, doi:10.37201/req/158.2021
Giacomorossetti, Barriot, Tropia, Dionellis, Gorgulla et al., Identification of low micromolar SARS-CoV-2 Mpro inhibitors from hits identified by in silico screens, bioRxiv, doi:10.1101/2020.12.03.409441
Hoffman, Kania, Brothers, Davies, Ferre et al., Discovery of ketone-based covalent inhibitors of coronavirus 3CL proteases for the potential therapeutic treatment of COVID-19, J. Med. Chem, doi:10.1021/acs.jmedchem.0c01063
Hoffman, Kania, Owen, Pettersson, Sammonsjamison et al., Method of treating Covid-19
Hoffman, Kaniajames, Niemansimon, Plankengeorge, Smith, Anticoronviral compounds and compositions, their pharmaceutical uses and materials for their synthesis
Hu, Su, Shao, Zhao, Chen et al., Identification of C270 as a novel site for allosteric modulators of SARS-CoV-2 papain-like protease, bioRxiv, doi:10.1101/2022.03.30.486313
Hulda, Jonsdottir, Julien, Padey, Bouveret et al., Molnupiravir combined with different repurposed drugs further inhibits SARS-CoV-2 infection in human nasal epithelium in vitro, bioRxiv, doi:10.1101/2022.01.10.475377
Ja, Pham-The, Pérez-Doñate, Torrens, Pérez-Giménez et al., A review of computational approaches targeting SARS-CoV-2 main protease to the discovery of new potential antiviral compounds, Curr. Top. Med. Chem, doi:10.2174/2667387816666220426133555
Johansen-Leete, Ullrich, Fry, Frkic, Bedding et al., Antiviral cyclic peptides targeting the main protease of SARS-CoV-2, Chem. Sci, doi:10.1039/d1sc06750h
Kim, Lee, Yang, Kim, Kim et al., The architecture of SARS-CoV-2 transcriptome, Cell, doi:10.1016/j.cell.2020.04.011
Kodan, Kimura, Kioka, Nakatsu, Kato et al., ABCB1/MDR1/P-gp employs an ATP-dependent twist-and-squeeze mechanism to export hydrophobic drugs, FEBS Lett, doi:10.1002/1873-3468.14018
Li, Zhou, Zhong, Zeng, Mccormick et al., Structural basis of main proteases of coronavirus bound to drug candidate PF-07304814, J. Mol. Biol, doi:10.1016/j.jmb.2022.167706
Liu, Boland, Scholle, Bardiot, Marchand et al., Dual inhibition of SARS-CoV-2 and human rhinovirus with protease inhibitors in clinical development, Antivir. Res, doi:10.1016/j.antiviral.2021.105020
Liu, Rong, Conditionally reprogrammed human normal airway epithelial cells at ali: A physiological model for emerging viruses, Virol. Sin, doi:10.1007/s12250-020-00244-z
Ltd, Pfizer reports fourth-quarter and full-year 2021 results
Ltd, Pfizer reports second-quarter 2021 results
Ltd, Pfizer reports third-quarter 2021 results
Ramos-Guzman, Ruiz-Pernia, Tunon, Inhibition mechanism of SARS-CoV-2 main protease with ketone-based inhibitors unveiled by multiscale simulations: Insights for improved designs, Angew. Chem. Int. Ed. Engl, doi:10.1002/anie.202110027
Sacco, Hu, Gongora, Meilleur, Kemp et al., The P132H mutation in the main protease of Omicron SARS-CoV-2 decreases thermal stability without compromising catalysis or smallmolecule drug inhibition, Cell Res, doi:10.1038/s41422-022-00640-y
Sanches Bma, Ferreira, Is prodrug design an approach to increase water solubility?, Int. J. Pharm, doi:10.1016/j.ijpharm.2019.118498
Simsek Yavuz, Komsuoglu Celikyurt, An update of anti-viral treatment of COVID-19, Turk. J. Med. Sci, doi:10.3906/sag-2106-250
Singh, Toussi, Hackman, Chan, Rao et al., Innovative randomized Phase 1 study and dosing regimen selection to accelerate and inform pivotal COVID-19 trial of nirmatrelvir, medRxiv, doi:10.1101/2022.02.08.22270649
Sokullu, Gauthier, Coulombe, Analysis of the SARS-CoV-2-host protein interaction network reveals new biology and drug candidates: Focus on the spike surface glycoprotein and RNA polymerase, Expert Opin. Drug Discov, doi:10.1080/17460441.2021.1909566
Steven, Joseph Mitchell, Nieman, Anticoronaviral compounds and compositions, their pharmaceutical uses and materials for their synthesis, PCT Pat. Appl
Taskar, Neuhoff, Patel, Yoshida, Paine, Clinical relevance of hepatic and renal P-gp/BCRP inhibition of drugs: An International Frontiers in Pharmacology frontiersin
Tripathi, Goshisht, COVID-19: Inflammatory responses, structure-based drug design and potential therapeutics, Mol. Divers, doi:10.1007/s11030-020-10176-1
Vandyck, Abdelnabi, Gupta, Jochmans, Jekle et al., ALG-097111, a potent and selective SARS-CoV-2 3chymotrypsin-like cysteine protease inhibitor exhibits in vivo efficacy in a Syrian Hamster model, Biochem. Biophys. Res. Commun, doi:10.1016/j.bbrc.2021.03.096
Xi He, Xu, Rodriguez, Goh, Wei et al., Generation of a SARS-CoV-2 replicon as a model system to dissect virus replication and antiviral inhibition, bioRxiv, doi:10.1101/2020.12.12.422532
Yao, Song, Chen, Wu, Xu et al., Molecular architecture of the SARS-CoV-2 virus, Cell, doi:10.1016/j.cell.2020.09.018
Zhang, Sun, Curth, Drosten, Sauerhering et al., Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors, Frontiers in Pharmacology frontiersin, doi:10.1126/science.abb3405
DOI record:
{
"DOI": "10.3389/fphar.2022.1035969",
"ISSN": [
"1663-9812"
],
"URL": "http://dx.doi.org/10.3389/fphar.2022.1035969",
"abstract": "<jats:p>Main protease (M<jats:sup>pro</jats:sup>) is a superior target for anti-SARS-COV-2 drugs. PF-07304814 is a phosphate ester prodrug of PF-00835231 that is rapidly metabolized into the active metabolite PF-00835231 by alkaline phosphatase (ALP) and then suppresses SARS-CoV-2 replication by inhibiting M<jats:sup>pro</jats:sup>. PF-07304814 increased the bioavailability of PF-00835231 by enhancing plasma protein binding (PPB). P-glycoprotein (P-gp) inhibitors and cytochrome P450 3A (CYP3A) inhibitors increased the efficacy of PF-00835231 by suppressing its efflux from target cells and metabolism, respectively. The life cycle of SARS-CoV-2 is approximately 4 h. The mechanisms and efficacy outcomes of PF-00835231 occur simultaneously. PF-00835231 can inhibit not only cell infection (such as Vero E6, 293T, Huh-7.5, HeLa<jats:sup>+angiotensin-converting enzyme 2 (ACE2)</jats:sup>, A549<jats:sup>+ACE2</jats:sup>, and MRC-5) but also the human respiratory epithelial organ model and animal model infection. PF-07304814 exhibits a short terminal elimination half-life and is cleared primarily through renal elimination. There were no significant adverse effects of PF-07304814 administration in rats. Therefore, PF-07304814 exhibits good tolerability, pharmacology, pharmacodynamics, pharmacokinetics, and safety in preclinical trials. However, the Phase 1 data of PF-07304814 were not released. The Phase 2/3 trial of PF-07304814 was also suspended. Interestingly, the antiviral activities of PF-00835231 derivatives (compounds 5–22) are higher than, similar to, or slightly weaker than those of PF-00835231. In particular, compound 22 exhibited the highest potency and had good safety and stability. However, the low solubility of compound 22 limits its clinical application. Prodrugs, nanotechnology and salt form drugs may solve this problem. In this review, we focus on the preclinical data of PF-07304814 and its active metabolite derivatives to hopefully provide knowledge for researchers to study SARS-CoV-2 infection.</jats:p>",
"alternative-id": [
"10.3389/fphar.2022.1035969"
],
"author": [
{
"affiliation": [],
"family": "Chen",
"given": "Wujun",
"sequence": "first"
},
{
"affiliation": [],
"family": "Shao",
"given": "Yingchun",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Peng",
"given": "Xiaojin",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Liang",
"given": "Bing",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Xu",
"given": "Jiazhen",
"sequence": "additional"
},
{
"affiliation": [],
"family": "Xing",
"given": "Dongming",
"sequence": "additional"
}
],
"container-title": "Frontiers in Pharmacology",
"container-title-short": "Front. Pharmacol.",
"content-domain": {
"crossmark-restriction": true,
"domain": [
"frontiersin.org"
]
},
"created": {
"date-parts": [
[
2022,
11,
11
]
],
"date-time": "2022-11-11T05:59:29Z",
"timestamp": 1668146369000
},
"deposited": {
"date-parts": [
[
2022,
11,
11
]
],
"date-time": "2022-11-11T05:59:33Z",
"timestamp": 1668146373000
},
"funder": [
{
"DOI": "10.13039/501100007129",
"doi-asserted-by": "publisher",
"name": "Natural Science Foundation of Shandong Province"
}
],
"indexed": {
"date-parts": [
[
2024,
1,
12
]
],
"date-time": "2024-01-12T14:04:25Z",
"timestamp": 1705068265745
},
"is-referenced-by-count": 4,
"issued": {
"date-parts": [
[
2022,
11,
11
]
]
},
"license": [
{
"URL": "https://creativecommons.org/licenses/by/4.0/",
"content-version": "vor",
"delay-in-days": 0,
"start": {
"date-parts": [
[
2022,
11,
11
]
],
"date-time": "2022-11-11T00:00:00Z",
"timestamp": 1668124800000
}
}
],
"link": [
{
"URL": "https://www.frontiersin.org/articles/10.3389/fphar.2022.1035969/full",
"content-type": "unspecified",
"content-version": "vor",
"intended-application": "similarity-checking"
}
],
"member": "1965",
"original-title": [],
"prefix": "10.3389",
"published": {
"date-parts": [
[
2022,
11,
11
]
]
},
"published-online": {
"date-parts": [
[
2022,
11,
11
]
]
},
"publisher": "Frontiers Media SA",
"reference": [
{
"DOI": "10.3390/molecules27082523",
"article-title": "Advances in the development of SARS-CoV-2 Mpro inhibitors",
"author": "Agost-Beltrán",
"doi-asserted-by": "publisher",
"first-page": "2523",
"journal-title": "Molecules",
"key": "B1",
"volume": "27",
"year": "2022"
},
{
"DOI": "10.3390/ijms22179124",
"article-title": "Exploring the binding mechanism of PF-07321332 SARS-CoV-2 protease inhibitor through molecular dynamics and binding free energy simulations",
"author": "Ahmad",
"doi-asserted-by": "publisher",
"first-page": "9124",
"journal-title": "Int. J. Mol. Sci.",
"key": "B2",
"volume": "22",
"year": "2021"
},
{
"DOI": "10.1002/jmv.26814",
"article-title": "Targeting SARS‐CoV‐2 viral proteases as a therapeutic strategy to treat COVID‐19",
"author": "Anirudhan",
"doi-asserted-by": "publisher",
"first-page": "2722",
"journal-title": "J. Med. Virol.",
"key": "B3",
"volume": "93",
"year": "2021"
},
{
"DOI": "10.3390/molecules26061678",
"article-title": "Is PF-00835231 a pan-SARS-CoV-2 Mpro inhibitor? A comparative study",
"author": "Baig",
"doi-asserted-by": "publisher",
"first-page": "1678",
"journal-title": "Molecules",
"key": "B4",
"volume": "26",
"year": "2021"
},
{
"DOI": "10.3390/polym13213823",
"article-title": "Unveiling the effect of low pH on the SARS-CoV-2 main protease by molecular dynamics simulations",
"author": "Barazorda-Ccahuana",
"doi-asserted-by": "publisher",
"first-page": "3823",
"journal-title": "Polym. (Basel)",
"key": "B5",
"volume": "13",
"year": "2021"
},
{
"DOI": "10.1038/s41467-021-26239-2",
"article-title": "Preclinical characterization of an intravenous coronavirus 3CL protease inhibitor for the potential treatment of COVID19",
"author": "Boras",
"doi-asserted-by": "publisher",
"first-page": "6055",
"journal-title": "Nat. Commun.",
"key": "B6",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.2174/2667387816666220426133555",
"article-title": "A review of computational approaches targeting SARS-CoV-2 main protease to the discovery of new potential antiviral compounds",
"author": "Castillo-Garit Ja",
"doi-asserted-by": "publisher",
"journal-title": "Curr. Top. Med. Chem.",
"key": "B7",
"volume": "22",
"year": "2022"
},
{
"DOI": "10.3390/pharmaceutics13081208",
"article-title": "The effect of plasma protein binding on the therapeutic monitoring of antiseizure medications",
"author": "Charlier",
"doi-asserted-by": "publisher",
"first-page": "1208",
"journal-title": "Pharmaceutics",
"key": "B8",
"volume": "13",
"year": "2021"
},
{
"DOI": "10.1080/10408347.2020.1819193",
"article-title": "Recent progress of glutathione (GSH) specific fluorescent probes: Molecular design, photophysical property, recognition mechanism and bioimaging",
"author": "Chen",
"doi-asserted-by": "publisher",
"first-page": "649",
"journal-title": "Crit. Rev. Anal. Chem.",
"key": "B9",
"volume": "52",
"year": "2022"
},
{
"DOI": "10.1101/2022.08.04.22278378",
"article-title": "Viral kinetics of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) Omicron infection in mRNA-vaccinated individuals treated and not treated with nirmatrelvir-ritonavir",
"author": "Dai",
"doi-asserted-by": "publisher",
"first-page": "22278378",
"journal-title": "medRxiv.",
"key": "B10",
"volume": "2022",
"year": "2022"
},
{
"DOI": "10.3389/fphys.2021.612319",
"article-title": "SARS-COV-2 and ocular surface: From physiology to pathology, a route to understand transmission and disease",
"author": "de Freitas Santoro",
"doi-asserted-by": "publisher",
"first-page": "612319",
"journal-title": "Front. Physiol.",
"key": "B11",
"volume": "12",
"year": "2021"
},
{
"DOI": "10.1128/JVI.01819-20",
"article-title": "A comparative analysis of SARS-CoV-2 antivirals characterizes 3CL(pro) inhibitor PF-00835231 as a potential new treatment for COVID-19",
"author": "de Vries",
"doi-asserted-by": "publisher",
"first-page": "01819-20",
"journal-title": "J. Virol.",
"key": "B12",
"volume": "95",
"year": "2021"
},
{
"DOI": "10.1080/07853890.2022.2031274",
"article-title": "Emerging COVID-19 variants and their impact on SARS-CoV-2 diagnosis, therapeutics and vaccines",
"author": "Fernandes",
"doi-asserted-by": "publisher",
"first-page": "524",
"journal-title": "Ann. Med.",
"key": "B13",
"volume": "54",
"year": "2022"
},
{
"DOI": "10.1002/ddr.21953",
"article-title": "Phosphate moiety in FDA-approved pharmaceutical salts and prodrugs",
"author": "Fulmali A",
"doi-asserted-by": "publisher",
"first-page": "1059",
"journal-title": "Drug Dev. Res.",
"key": "B14",
"volume": "83",
"year": "2022"
},
{
"DOI": "10.37201/req/158.2021",
"article-title": "Pharmacological treatment of COVID-19: An opinion paper",
"author": "Garcia-Lledo",
"doi-asserted-by": "publisher",
"first-page": "115",
"journal-title": "Rev. Esp. Quimioter.",
"key": "B15",
"volume": "35",
"year": "2021"
},
{
"DOI": "10.1101/2020.12.03.409441",
"article-title": "Identification of low micromolar SARS-CoV-2 Mpro inhibitors from hits identified by in silico screens",
"author": "GiacomoRossetti",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B16",
"year": "2020"
},
{
"DOI": "10.1101/2022.03.30.486313",
"article-title": "Identification of C270 as a novel site for allosteric modulators of SARS-CoV-2 papain-like protease",
"author": "Hangchen Hu",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B17",
"year": "2022"
},
{
"DOI": "10.1021/acs.jmedchem.0c01063",
"article-title": "Discovery of ketone-based covalent inhibitors of coronavirus 3CL proteases for the potential therapeutic treatment of COVID-19",
"author": "Hoffman",
"doi-asserted-by": "publisher",
"first-page": "12725",
"journal-title": "J. Med. Chem.",
"key": "B18",
"volume": "63",
"year": "2020"
},
{
"article-title": "Method of treating Covid-19",
"author": "Hoffman",
"key": "B19",
"year": "2020"
},
{
"article-title": "Anticoronviral compounds and compositions, their pharmaceutical uses and materials for their synthesis",
"author": "Hoffman",
"key": "B20",
"year": "2005"
},
{
"DOI": "10.1101/2022.01.10.475377",
"article-title": "Molnupiravir combined with different repurposed drugs further inhibits SARS-CoV-2 infection in human nasal epithelium in vitro",
"author": "Hulda",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B21",
"year": "2022"
},
{
"DOI": "10.1039/d1sc06750h",
"article-title": "Antiviral cyclic peptides targeting the main protease of SARS-CoV-2",
"author": "Johansen-Leete",
"doi-asserted-by": "publisher",
"first-page": "3826",
"journal-title": "Chem. Sci.",
"key": "B22",
"volume": "13",
"year": "2022"
},
{
"DOI": "10.1016/j.cell.2020.04.011",
"article-title": "The architecture of SARS-CoV-2 transcriptome",
"author": "Kim",
"doi-asserted-by": "publisher",
"first-page": "914",
"journal-title": "Cell",
"key": "B23",
"volume": "181",
"year": "2020"
},
{
"DOI": "10.1002/1873-3468.14018",
"article-title": "ABCB1/MDR1/P-gp employs an ATP-dependent twist-and-squeeze mechanism to export hydrophobic drugs",
"author": "Kodan A",
"doi-asserted-by": "publisher",
"first-page": "707",
"journal-title": "FEBS Lett.",
"key": "B24",
"volume": "595",
"year": "2021"
},
{
"DOI": "10.1016/j.jmb.2022.167706",
"article-title": "Structural basis of main proteases of coronavirus bound to drug candidate PF-07304814",
"author": "Li J",
"doi-asserted-by": "publisher",
"first-page": "167706",
"journal-title": "J. Mol. Biol.",
"key": "B25",
"volume": "434",
"year": "2022"
},
{
"DOI": "10.1016/j.antiviral.2021.105020",
"article-title": "Dual inhibition of SARS-CoV-2 and human rhinovirus with protease inhibitors in clinical development",
"author": "Liu",
"doi-asserted-by": "publisher",
"first-page": "105020",
"journal-title": "Antivir. Res.",
"key": "B26",
"volume": "187",
"year": "2021"
},
{
"DOI": "10.1007/s12250-020-00244-z",
"article-title": "Conditionally reprogrammed human normal airway epithelial cells at ali: A physiological model for emerging viruses",
"author": "Liu",
"doi-asserted-by": "publisher",
"first-page": "280",
"journal-title": "Virol. Sin.",
"key": "B27",
"volume": "35",
"year": "2020"
},
{
"key": "B28",
"unstructured": "Pfizer reports fourth-quarter and full-year 2021 results\n LtdP. P."
},
{
"key": "B29",
"unstructured": "Pfizer reports second-quarter 2021 results\n LtdP. P."
},
{
"DOI": "10.1016/j.fopow.2021.11.012",
"doi-asserted-by": "crossref",
"key": "B30",
"unstructured": "Pfizer reports third-quarter 2021 results\n LtdP. P."
},
{
"DOI": "10.1002/anie.202110027",
"article-title": "Inhibition mechanism of SARS-CoV-2 main protease with ketone-based inhibitors unveiled by multiscale simulations: Insights for improved designs",
"author": "Ramos-Guzman",
"doi-asserted-by": "publisher",
"first-page": "25933",
"journal-title": "Angew. Chem. Int. Ed. Engl.",
"key": "B31",
"volume": "60",
"year": "2021"
},
{
"article-title": "Anticoronaviral compounds and compositions, their pharmaceutical uses and materials for their synthesis",
"author": "Robert Steven",
"journal-title": "PCT Pat. Appl.",
"key": "B32",
"year": "2006"
},
{
"DOI": "10.1038/s41422-022-00640-y",
"article-title": "The P132H mutation in the main protease of Omicron SARS-CoV-2 decreases thermal stability without compromising catalysis or small-molecule drug inhibition",
"author": "Sacco",
"doi-asserted-by": "publisher",
"first-page": "498",
"journal-title": "Cell Res.",
"key": "B33",
"volume": "32",
"year": "2022"
},
{
"DOI": "10.1016/j.ijpharm.2019.118498",
"article-title": "Is prodrug design an approach to increase water solubility?",
"author": "Sanches Bma",
"doi-asserted-by": "publisher",
"first-page": "118498",
"journal-title": "Int. J. Pharm.",
"key": "B34",
"volume": "568",
"year": "2019"
},
{
"DOI": "10.3906/sag-2106-250",
"article-title": "An update of anti-viral treatment of COVID-19",
"author": "Simsek Yavuz",
"doi-asserted-by": "publisher",
"first-page": "3372",
"journal-title": "Turk. J. Med. Sci.",
"key": "B35",
"volume": "51",
"year": "2021"
},
{
"DOI": "10.1101/2022.02.08.22270649",
"article-title": "Innovative randomized Phase 1 study and dosing regimen selection to accelerate and inform pivotal COVID-19 trial of nirmatrelvir",
"author": "Singh",
"doi-asserted-by": "publisher",
"first-page": "1",
"journal-title": "medRxiv",
"key": "B36",
"year": "2022"
},
{
"DOI": "10.1080/17460441.2021.1909566",
"article-title": "Analysis of the SARS-CoV-2-host protein interaction network reveals new biology and drug candidates: Focus on the spike surface glycoprotein and RNA polymerase",
"author": "Sokullu",
"doi-asserted-by": "publisher",
"first-page": "881",
"journal-title": "Expert Opin. Drug Discov.",
"key": "B37",
"volume": "16",
"year": "2021"
},
{
"DOI": "10.1002/cpt.2670",
"article-title": "Clinical relevance of hepatic and renal P-gp/BCRP inhibition of drugs: An International Transporter Consortium perspective",
"author": "Taskar",
"doi-asserted-by": "publisher",
"first-page": "573",
"journal-title": "Clin. Pharmacol. Ther.",
"key": "B38",
"volume": "112",
"year": "2022"
},
{
"DOI": "10.1007/s11030-020-10176-1",
"article-title": "COVID-19: Inflammatory responses, structure-based drug design and potential therapeutics",
"author": "Tripathi N",
"doi-asserted-by": "publisher",
"first-page": "629",
"journal-title": "Mol. Divers.",
"key": "B39",
"volume": "26",
"year": "2022"
},
{
"DOI": "10.1016/j.bbrc.2021.03.096",
"article-title": "ALG-097111, a potent and selective SARS-CoV-2 3-chymotrypsin-like cysteine protease inhibitor exhibits in vivo efficacy in a Syrian Hamster model",
"author": "Vandyck",
"doi-asserted-by": "publisher",
"first-page": "134",
"journal-title": "Biochem. Biophys. Res. Commun.",
"key": "B40",
"volume": "555",
"year": "2021"
},
{
"DOI": "10.1101/2020.12.12.422532",
"article-title": "Generation of a SARS-CoV-2 replicon as a model system to dissect virus replication and antiviral inhibition",
"author": "Xi He",
"doi-asserted-by": "publisher",
"journal-title": "bioRxiv",
"key": "B41",
"year": "2020"
},
{
"DOI": "10.1016/j.cell.2020.09.018",
"article-title": "Molecular architecture of the SARS-CoV-2 virus",
"author": "Yao",
"doi-asserted-by": "publisher",
"first-page": "730",
"journal-title": "Cell",
"key": "B42",
"volume": "183",
"year": "2020"
},
{
"DOI": "10.1126/science.abb3405",
"article-title": "Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors",
"author": "Zhang",
"doi-asserted-by": "publisher",
"first-page": "409",
"journal-title": "Science",
"key": "B43",
"volume": "368",
"year": "2020"
}
],
"reference-count": 43,
"references-count": 43,
"relation": {},
"resource": {
"primary": {
"URL": "https://www.frontiersin.org/articles/10.3389/fphar.2022.1035969/full"
}
},
"score": 1,
"short-title": [],
"source": "Crossref",
"subject": [
"Pharmacology (medical)",
"Pharmacology"
],
"subtitle": [],
"title": "Review of preclinical data of PF-07304814 and its active metabolite derivatives against SARS-CoV-2 infection",
"type": "journal-article",
"update-policy": "http://dx.doi.org/10.3389/crossmark-policy",
"volume": "13"
}